DNA
DNA. Chapter 16 and 17 Biology II. Meselson and Stahl Experiment. http://www.sumanasinc.com/webcontent/anisamples/majorsbiology/meselson.html. Search for the Genetic Material. Scientists knew the hereditary material contained Protein—macromolecules, very diverse, specific

DNA
E N D
Presentation Transcript
DNA Chapter 16 and 17 Biology II
Meselson and Stahl Experiment • http://www.sumanasinc.com/webcontent/anisamples/majorsbiology/meselson.html
Search for the Genetic Material • Scientists knew the hereditary material contained • Protein—macromolecules, very diverse, specific • Nucleic Acids—little known
Experiments • Griffith--1928 • Studied Streptococcus pneumonia • Two strains of pneumococcus: • S-strain • Smooth, polysaccharide capsule; pathogenic • R-strain • Rough, no capsule, not pathogenic
Griffith’s Experiments • Four Sets of Experiments: • 1. Live S injected into mice • 2. Live R into mice • 3. Heat killed S • 4. Heat killed S+R
Griffith’s Experiment • Live S cells retrieved from Group 4 • R strains had acquired from dead S the ability to make polysaccharide coats Transformation • *protein is NOT the transforming agent b/c • heat denatures protein
Hersey and Chase Experiments--1952 • http://highered.mcgraw-hill.com/sites/0072437316/student_view0/chapter14/animations.html# • Bacteriophage • Virus that infects bacteria • Consists of : • Protein coat • Viral protein tagged with35S • With DNA • DNA tagged with 32P
Hersey and Chase Experiments--1952 • S was incorporated into bacteriophage protein • P was incorporated into bacteriophage DNA • Labeled T2 phages were allowed to infect separate samples of E.coli • Cultures were agitated to shake loose phages on outside
Hersey-Chase Experiments • What happened? • Radioactivity in pelletP in DNA of bacteria • What does it mean? • Radioactivity in supernatantS in protein coat of virus • PROTEIN is NOT the hereditary material • DNA IS the hereditary material
Additional evidence that DNA is the genetic material • Eukaryotic cell doubles DNA prior to mitosis • During mitosis DNA is divided between daughter cells • Diploid cells have 2x the DNA of haploid cells • Chargaff’s Rule (using paper chromatography) • A=T; G=C
Watson-Crick Double Helix • 1953—Three groups race to find the answer: • Linus Pauling Cal Tech • Maurice Wilkins and Rosalind Franklin Kings College London • James Watson and Francis Crick Cambridge University
Rosalind Franklin • *Watson went to Cambridge saw x-ray crystallography (Rosalind Franklin) • From photo, Watson concluded: DNA is uniform 2nm • Bases are .34 nm apart with 10 bases making a full rung at 3.4 nm
DNA Nitrogenous bases • Purines • Adenine and guanine • Pyrimidines • Thymine • Cytosine • A-T (2 hydrogen bonds) • G-C (3 hydrogen bonds)
DNA Structure • Nucleotides line up A-T; G-C • Enzymes link the nucleotides together with P-S groups • Each strand=one old and one newly created strand
Purines and Pyrimidines Purines Pyrimidines
DNA Replication • DNADNA • http://highered.mcgraw-hill.com/sites/0072437316/student_view0/chapter14/animations.html • Origin of replication • Where replication begins • Replication
Enzymes • DNA polymerase • Elongation on new DNA strand 5’3’ only • DNA ligase • Joins fragments into single strand by phosphodiester bonds
Enzymes • 3. DNA helicase • Unwind and unzip DNA • 4. RNA primase • Need to begin the process of replication b/c polymerase cannot begin adding nucleotides unless there is one there • Binding proteins—keep the strands apart
DNA • DNA strands are antiparellel • Sugar-phosphate backbone run in opposite directions • Leading strand--Phosphates are attached to #5 carbon 5’ Overall direction of replication
Leading Strand • Replicated continuously • 5’-3’ direction
Lagging Strand • Lagging strand—phosphates are attached to #3 carbon 3’ • Replicated in pieces in 5’-3’ direction • Pieces are called Okazaki fragments Lagging strand Leading strand
Replicate this! • TAC TTA AAA CTT CGA CTA TTT ATT • _________________________________ • http://www.stolaf.edu/people/giannini/flashanimat/molgenetics/dna-rna2.swf
Semiconservative Model of DNA • 1950 Meselson and Stahl • Two DNA strands separate • Each strandtemplate for complementary strand
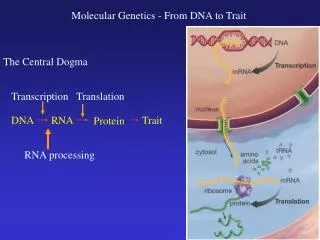
TranscriptionDNARNA • Two blue/purple strands • DNA • Red Blob • RNA polymerase • Green strand • mRNA • Where occur?
Transcription • http://www.stolaf.edu/people/giannini/flashanimat/molgenetics/transcription.swf • DNA base sequence below: • TAC TTA AAA CTT CGA CTA TTT ATT • Transcription • DNARNA • Transcribe top line • _______________________________
RNA • Single stranded • Nucleotide: 1. 5-C sugar • Ribose 2. Phosphate group 3. Nitrogenous bases • Uracil in place of Thymine (A-U)
Three types of RNA • mRNA • Carries the message to ribosome • tRNA • Specific for a particular amino acid • rRNA • Two subunits compose the ribosome
RNA • Function: • To synthesis proteins • Translation or protein synthesis • Structure: • Polymer of nucleotides • Single strand • 3 types
rRNA • rRNA—ribosomal RNA • Two subunits • Ribosome “reads” mRNA and produces a polypepide
3 Types of RNA • 1.mRNA • Messenger RNA • Single strand • Serves as a template (pattern for translation)
3 Types of RNA • 2. tRNA • Transfer RNA • 20+ types of tRNA • Cloverleaf shape • Each tRNA is specific for an amino acid
3 Types of RNA • 3. rRNA • Ribosomal RNA • Globular • 2 parts compose the ribosome • Where are they made?
mRNA--Codons • Codon • sequence of 3 nucleotide bases on mRNA
Transcription • Enzyme: RNA polymerase (3 kinds in eukaryotes) • “unzips” DNA and adds RNA nucleotides in the 5’ 3” direction
Transcription • Promotor • Site where the polymerase attaches • Termination site • Site where transcription ends • Transcription Unit • The stretch of DNA transcribed
Transcription • In eukaryotes, the mRNA is modified after transcription • A 5’ cap is added (guanine nuicleotide) • Poly A tail (adenine) • 50-250 nucleotides long
RNA splicing • “cut and paste” of original RNA molecule that was synthesized
RNA splicing • Introns • Noncoding regions of RNA • Interspersed between the coding regions • Exons • Coding regions that will exit the nucleus and be translated
Translation • RNA protein • Structure of a ribosome • Protein and rRNA • Most common form of RNA • Ribosomes are formed in the nucleolus
Translation • Three stages of translation • Initiation • Elongation • Termination
Initiation • Small ribosomal subunit binds to both the mRNA and the tRNA • Large ribosomal subunit attaches
Elongation • Codon recognition--mRNA and tRNA form hydrogen bonds at the “A” site of the ribosome • Peptide bond forms between amino acid at the “A” site and the growing polypeptide at the “P” site • Translocation • Ribosome moves the tRNA with polypeptide from the “A” to the “P” Exit site
Termination • Translation continues until “stop” codon on mRNA—UAA, UAG, or UGA • Polyribosomes • Multiple ribosomes translating the same rRNA (polysomes)
Genetic Code Tablecodons • Universal for almost all organisms • P. 308 in text • Use it to decode the base sequence on the next slide
Translate This • http://www.stolaf.edu/people/giannini/flashanimat/molgenetics/translation.swf • AUG AAU UUU GAA GCU GAU AAA UAA • __________________________________