Chapter 9 T-Cell Receptor
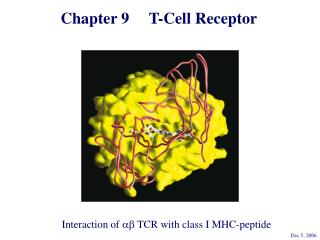
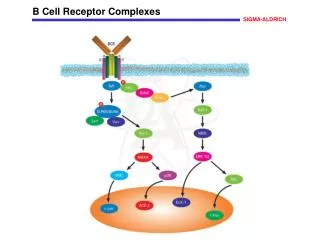
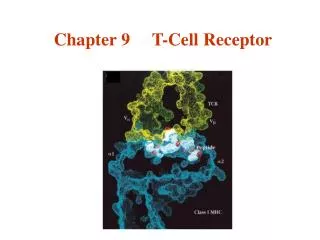
Chapter 9 T-Cell Receptor. Interaction of ab TCR with class I MHC-peptide. Dec 5, 2006. 你需要學習的課題: T 細胞受體是如何發現的 ? 有什麼特性 ? ab TCR 與 gd TCR 在辨認上有什麼不同 ? TCR 的基因及分子。 T 細胞辨識時,除了 TCR 之外還有哪些 coreceptors ?.

Chapter 9 T-Cell Receptor
E N D
Presentation Transcript
Chapter 9 T-Cell Receptor Interaction of ab TCR with class I MHC-peptide Dec 5, 2006
你需要學習的課題: • T 細胞受體是如何發現的? 有什麼特性? • ab TCR 與 gd TCR在辨認上有什麼不同? • TCR 的基因及分子。 • T 細胞辨識時,除了TCR 之外還有哪些 coreceptors ?
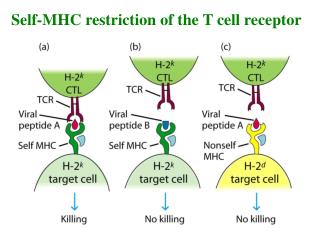
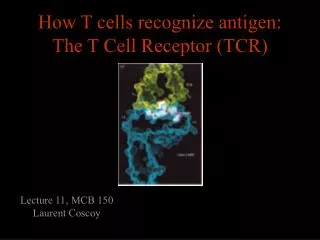
T Cells Recognize Ag Only When Presented on the Membrane of a Cell by a Self-MHC Molecule This work was published in 1974. Zinkernagel and Doherty were awarded the Nobel prize In 1996.
Identification of T-cell Receptors: • Clonotypic mAb • how? • 2. Gene cloning by subtractive hybridization
Identification and Cloning of the TCR Genes ~ Hedrick and Davis, 1984 ~ Well-thought-out assumptions of TCR Genes : 1. mRNAs are associated with membrane-bound polyribosomes rather than with free cytoplasmic ribosomes. 2. mRNAs are only expressed in T, but not in B cells. subtractive hybridization (98% of the genes expressed in T and B cells are the same) 3. DNA is rearranged in mature T cells.
Production and Identification of a cDNA Clone Encoding the TCR
subtractive hybridization ~ 2% of total cDNA: 3% x 2% = 0.06% each of the 6 T-cell clones showed different Southern blot patterns. , and clones 3, 4, 5, 6, 7, 8, 9, 10
The cDNA clone 1 identified by the Southern- blot analysis shown in the previous slide has all the hallmarks of a putative TCR gene: 1. It represents a gene sequence that rearranges. 2. It is expressed as a membrane-bound protein. 3. It is expressed only in T cells. The cDNA clone 1 is the b chain of the TCR. Later, cDNA clones were identified encoding theα chain, theγchain, and finally the δchain.
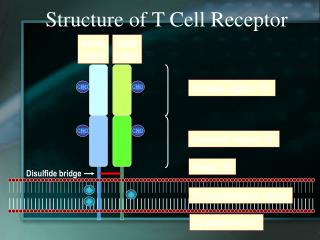
ab and gd T-cell Receptors: Structure and Roles
Structural Similarity between mIgM and TCR Fab or gd T-cell receptor • resembling an Fab fragment ?
TCRab TCRgd * * The δ-chain gene segments are located between the Vαand Jαsegments. a, g genes : V, J, C gene segments b, d genes : V, D, J, C gene segments
Comparison of the gd TCR and ab TCR % of CD3+1-10 % 90-99 % T cells elbow angle →Contribute to differences in signaling mechanism and in how the molecules interact with coreceptors.
gd T cells • In humans, the predominant receptor expressed • on circulating gd cells recognizes a microbial • phospholipid Ag,3-formyl-1-butyl pyrophosphate, • found on M. tuberculosis and other bacteria and • parasites (similat to pattern recognition receptor?) • This specificity for frequently encountered pathogens • led to speculation that gd cells may function as an • arm of the innate immune response, allowing rapid • reactivity to certain Ags without the need for a • processing step.
The specificity of circulating gd cells in the • mouse and of other species studied does not • parallel that of humans, suggesting that the gd • response may be directed against pathogens • commonly encountered by a given species. • Since gd cells can secrete a spectrum of • chemokines and cytokines, they may play a • regulatory role in recruiting other cells to the • site of invasion by pathogens.
Ligands Recognized by gd T Cells: - gd T cells appear to bind directly to Ags without requiring Ag processing and presentation by MHC. - Some gd T cells may uniquely respond to heat-shock proteinsand may have evolved to eliminate damaged cells as well as microbial invaders.
Organization and Rearrangement of TCR Genes
Germ-line Organization of the Mouse TCR a-, b-, g-, and d-chain Gene Segments d : a productive rearrangement of a-chain gene segments deletes Cd between Va and Ja
Gene Rearrangements That Yield a Functional Gene Encoding the ab TCR
The C region of TCR is much simpler than • the C region of Ig genes: • TCR is expressed only in a membrane-bound • form; thus, no differential RNA processing is • required to produce membrane and secreted form. • TCRa has only a single C gene segment and • TCRb has two C gene segments. • No known functional differences exist in C • regions.
Although B cells and T cells use very similar • mechanisms for V-region gene rearrangements, • the Ig genes are not rearranged in T cells and the • TCR genes are not rearranged in B cells. • The recombinase enzyme system is differently • regulated in B and T cell lineage, so that only • rearrangement of the correct receptor DNA occurs. • Chromatin is also uniquely re-configured in B • cells and T cells to allow the recombinase access • to Ig and TCR genes, respectively.
Comparison of Mechanisms for Generating Diversity in TCR Genes and Ig Genes
The Location of One-turn (12-bp) and Two-turn (23-bp) Recognition Signal Sequences (RSS) in TCR b- and d-chain DNA Differs from That in Ig H-chain DNA or V-D-D-D-J in humans generate considerable additional diversity in TCR genes.
1+41+42+43+44+45+46 =5461 N-addition occurs in all the TCR genes. • Although each junctional region in a TCR gene encodes only 10 to 20 a.a., • enormous diversity can be generated in these regions. • The combined effects of P- and N- addition plus joining flexibility are • estimated to be 1013 possible a.a. sequences in the TCR CDR3 region.
abTCR: 3.0 x 103 x 4.6 x 102 = 1.4 x 106
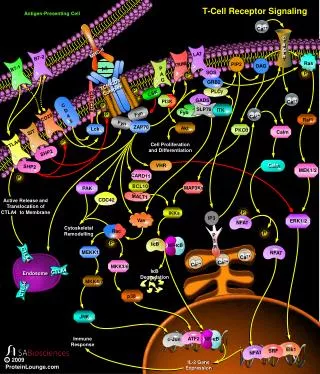
T-cell Receptor Complex: TCR-CD3
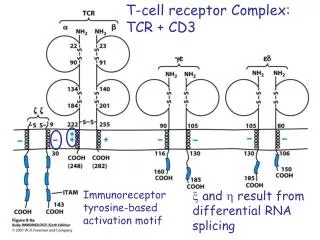
T-Cell Receptor Complex: TCR-CD3 CD3 : ge + de + zz (90%) or zh (10%) or immunoreceptor tyrosine-based activation motif
Accessory Molecules Which Strengthen the Interaction between T Cells and APC (T cell) (APC) (costimulatory)
CD4 and CD8 Coreceptors Bind to Conserved Regions of MHC Class II or I Molecules sometimes aa homodimer 55-kDa 30-38 kDa each chain CD4 CD8 class I class II
Interaction of CD8 Coreceptor with TCR and Class I MHC Molecule a1 a2
Interaction of CD4 Coreceptor with TCR and Class II MHC Molecule
Dissociation Constants (Kd) for Various Biological Systems (10-4 to 10-7) (10-6 to 10-10)
Interactions between TCR/Peptide-MHC and Accessory Molecules/Ligands
Two-point Contact - extracellular portion of CD4 : MHC intracellular CD4 : p56lck - z
Three-dimensional Structures of TCR-peptide-MHC Complexes
Interaction between TCR and HLA-A2 with Bound HTLV-I Tax Peptide HTLV-1 tax peptide HLA-A2
MHC Molecule Viewed from Above HV loops of TCR-Vb Peptide HV loops of TCR-Va HLA-A2
Ternary Complex of TCR Bound to H-2Kb and Peptide TCR CDR1 & CDR2 of Vb CDR3 of Vb peptide H-2Kb CDR1 & CDR2 of Va CDR3 of Va
Comparison of the Interactions between ab TCR and MHC-peptides • The angles at which the TCR molecule sits on the class I and class II MHC-peptide are different. • More number of contact residues between TCR and class II
Alloreactivity of T Cells - a puzzling finding T cell recognition 1. self-MHC + foreign peptides + 2. allo-MHC + foreign peptides – self-MHC restriction
Alloreactivity of T Cells - a puzzling finding T cell recognition 1. self-MHC + foreign peptides + 2. allo-MHC + foreign peptides – however, 3. allo-MHC ± allo-peptides +++ why? one explanation : cross-reactivity of 1 and 3 self-MHC restriction