The T Cell Antigen Receptor Complex
610 likes | 850 Views
The T Cell Antigen Receptor Complex. Generating Evidence for and isolating the TCR. Generation of cytotoxic T lymphocytes (CTLs) Zinkernagel and Doherty – 1974 demonstrated role of another molecule, MHC got Nobel Prize 1996

The T Cell Antigen Receptor Complex
E N D
Presentation Transcript
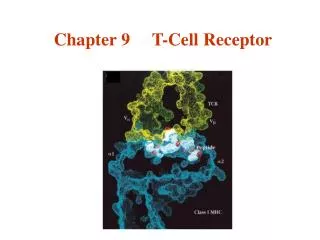
The T Cell Antigen Receptor Complex
Generating Evidence for and isolating the TCR • Generation of cytotoxic T lymphocytes (CTLs) • Zinkernagel and Doherty – 1974 demonstrated role of another molecule, MHC got Nobel Prize 1996 • Isolation of the TCR using monclonal antibodies that were clonotypici.e recognized single T cell clone • Identification of genes of the TCR
Grow and clone a single antigen-specific T cell in-vitro with antigen, IL-2 and antigen presenting cells Discovery of the T cell antigen receptor (TcR) Polyclonal T cells from an immunised strain A mouse Monoclonal (cloned) T cells In vitro “clonal selection” means each daughter cell has the same antigen specificity as the parent cell Most molecules present on the monoclonal T cells will be identical to the polyclonal T cells EXCEPTfor the antigen combining site of the T cell antigen receptor
T cell clone from a strain A mouse Naïve strain A mouse Make monoclonal antibodies by hybridisation of the spleen cells with a myeloma cell line Making anti- clonotypic TcR antibodies The strain A mouse will not make antibodies to the hundreds of different molecules associated with strain A T cells due to self tolerance BUT The naïve mouse has never raised T cells with the specificity of the T cell clone, SO the only antigen in the immunisation that the A strain mouse has never seen will be the antigen receptor of the monoclonal T cells
Y Monoclonal antibodies Clone used for immunisation T cell clones Making anti- clonotypic TcR antibodies Screen the supernatant of each cloned hybridoma against a panel of T cell clones of different specificity (i.e.cells with subtly different antigen-binding structures) Y Y Y Y Y Y Y Y Y Y Y Anti-TcR Abs that recognise only one clone of T cells areCLONOTYPIC Hypothesise that anti-clonotype Abs recognise the antigen receptor
Y Y Y Y Y Y Y Y Y Capture anti-clonotype Ab-Ag complex on insoluble support IMMUNOPRECIPITATION Wash away unbound protein Y Y Y Y Y Y Y Y Discovery of the T cell antigen receptor (TcR) Lyse cells and add anti-clonotype Ab that binds to unique T cell structures Elute Ag from Ab and analyse the clonotypically-expresssed proteins biochemically Principal component was a heterodimeric 90kDa protein composed of a 40kDa and a 50kDa molecule ( and chains) Several other molecules were co-immunoprecipitated.
T cell clone A T cell clone B T cell clone C C V C V C V Structure of the TcR polypeptides Intact TcR chain polypeptides Cyanogen bromide digestion of the and proteins Biochemical analysis of digestion products Polypeptides contain a variable, clone-dependent pattern of digestion fragments and a fragment common to all TcR
B T Cloning of the TcR genes • The experimental strategy • The majority of genes expressed by T and B lymphocytes will be similar • Genes that greatly differ in their expression are most likely to be directly related to the specialised function of each cell • Subtract the genes expressed by B cells from the genes expressed by T cells leaving only the genes directly related to T cell function
mRNA B T AAAAA AAAAA AAAAA AAAAA T cell single stranded cDNA AAAAA AAAAA Hybridise the cDNA and mRNA shared between T and B cells Discard hybrids Clone and sequence T cell- specific genes Cloning of TcR genes by subtractive hybridisation Digest unhybridised B cell mRNA Isolate non-hybridising material specific to T cells
32P 32P Restriction enzyme sites GERMLINE DNA V D J C REARRANGED DNA V D J C Analysis of T cell-specific genes Of the T cell-specific genes cloned, which cDNA encoded the TcR? Assumptions made after the analysis of Ig genes: TcR genes rearrange from germline configuration Ig gene probes can be used as TcR genes will be homologous to Ig genes Find two restriction sites that flank the TcR region Cut the T cell cDNA and placental (i.e. germline) DNA and Southern blot the fragments
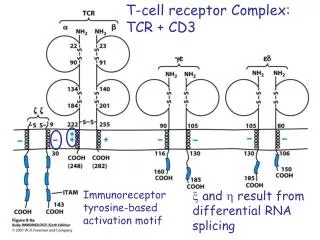
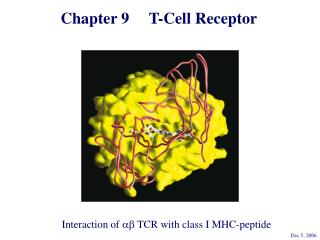
Each a and b chain consists of one ‘Ig-like’ N-terminal variable region (V), one Ig-like constant (C) domain, a hydrophobic transmembrane region, and a short cytoplasmic region. Thus the extracellular portion of the abheterodimer is structurally similar to the antigen-binding fragment (Fab) of an Ig, which is made up of the V and C regions of a light chain and the V region and one C region of a heavy chain. TCR
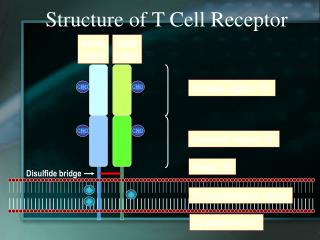
VH VH VL VL Fab CH CH CL CL CH CH Fc CH CH Transmembrane region Cytoplasmic tail The T cell antigen receptor Antigen combining site Resembles an Ig Fab fragment Va Vb Domain structure: Ig gene superfamily Carbohydrates Monovalent No alternative constant regions Ca Cb Never secreted Hinge Heterodimeric, chains are disuphide-bonded + + + Very short intracytoplasmic tail Positively charged amino acids in the TM region Antigen combining site made of juxtaposed Va and Vb regions 30,000 identical specificity TcR per cell
T cell antigen receptor diversity • TcRare highly variable in the individual • Diversity focused on small changes in the charge & shape presented at the end of the T cell receptor. • TcR diversity to the peptide antigens that bind to MHC molecules • Mechanisms of diversity closely related to T cell development • Random aspects of TcR construction ensures maximum diversity • Mechanisms of diversity generation similar to immunoglobulin genes
Generation of diversity in the TcR COMBINATORIAL DIVERSITY Multiple germline segments In the human TcR Variable (V) segments: ~70, 52 Diversity (D) segments: 0, 2 Joining (J) segments: 61, 13 The need to pair and chains to form a binding site doubles the potential for diversity JUNCTIONAL DIVERSITY Addition of non-template encoded (N) and palindromic (P) nucleotides at imprecise joints made between V-D-J elements SOMATIC MUTATION IS NOT USED TO GENERATE DIVERSITY IN TcR
Estimate of the number of human TcR and Ig Excluding somatic hypermutation Immunoglobulin TcR Element H 40 59 52 ~70 Variable segments 27 0 2 0 Diversity segments - - D segments in all 3 frames Yes Yes 6 9 13 61 Joining segments 2 (1)* 2 1 Joints with N & P nucleotides 2360 3640 No. of V gene pairs ~1013 ~1013 Junctional diversity ~1016** ~1016 Total diversity * Only half of human k chains have N & P regions **No of distinct receptors increased further by somatic hypermutation
L & V x70-80 J x 61 C TcR L & V x52 D1 Jb1 x 6 C1 D2 Jb2 x 7 C2 TcR Organisation of TcR genes TcR genes segmented into V, (D), J & C elements (VARIABLE, DIVERSITY, JOINING & CONSTANT) Closely resemble Ig genes (a~IgL and b~IgH) This example shows the mouse TcR locus
Vn V2 V1 J C Germline TcR Rearranged TcR 1° transcript TcR a gene rearrangement by SOMATIC RECOMBINATION Spliced TcR mRNA Rearrangement very similar to the IgL chains
Vn V2 V1 J C Vn+1 Productively rearranged TcR 1° transcript TcR a gene rearrangement RESCUE PATHWAY There is only a 1:3 chance of the join between the V and J region being in frame a chain tries for a second time to make a productive join using new V and J elements
L & V x52 D1 J C1 D2 J C2 Germline TcR TcR b gene rearrangement SOMATIC RECOMBINATION D-J Joining V-DJ joining Rearranged TcR 1° transcript C-VDJ joining Spliced TcR mRNA
D1 J C1 D2 J C2 V Germline TcR D-J Joining V-DJ joining 2nd chance at V-DJ joining Need to remove non productive rearrangement TcR b gene rearrangement RESCUE PATHWAY There is a 1:3 chance of productive D-J rearrangement and a 1:3 chance of productive D-J rearrangement (i.e only a 1:9 chance of a productive b chain rearrangement) Use (DJC)b2 elements
Va Ja 7 23 12 7 9 9 Jb 9 23 7 Db 12 7 7 12 9 9 Vb 9 7 23 V, D, J flanking sequences Sequencing upstream and downstream of V, D and J elements revealed conserved sequences of 7, 23, 9 and 12 nucleotides.
HEPTAMER - Always contiguous with coding sequence NONAMER - Separated fromthe heptamer by a 12 or 23 nucleotide spacer √ √ Jb Jb 9 9 23 23 7 7 Db Db 12 12 7 7 7 7 12 12 9 9 9 9 Vb Vb 9 9 7 7 23 23 Recombination signal sequences (RSS) 12-23 RULE – A gene segment flanked by a 23mer RSS can only be linked to a segment flanked by a 12mer RSS
12-mer = one turn 23-mer = two turns Intervening DNA of any length 23 12 Vb 7 9 7 Db Jb 9 Molecular explanation of the 12-23 rule
V4 V5 V3 V1 V3 V4 V2 V6 V2 V5 V6 V7 V8 V7 9 V9 D J V8 V9 9 23-mer • Heptamers and nonamers align back-to-back • The shape generated by the RSS’s acts as a target for recombinases 12-mer 7 7 D J V1 Molecular explanation of the 12-23 rule Loop of intervening DNA is excised • An appropriate shape can not be formed if two 23-mer flanked elements attempted to join (i.e. the 12-23 rule)
9 23 7 7 12 9 9 9 23 Coding joint Signal joint 12 V D J 7 7 V D J Junctional diversity Mini-circle of DNA is permanently lost from the genome Imprecise and random events that occur when the DNA breaks and rejoins allows new nucleotides to be inserted or lost from the sequence at and around the coding joint.
Looping out works if all V genes are in the same transcriptional orientation V1 V3 V4 V9 V2 D D J J V1 D J 9 7 23 9 23 7 How does recombination occur when a V gene is in opposite orientation to the DJ region? 12 7 9 V1 V3 V4 V9 V2 V4 D J 12 7 9 Non-deletional recombination
V4 and DJ in opposite transcriptional orientations 1. 2. D D D D J J J J 12 12 12 12 7 7 7 7 9 9 9 9 9 9 9 9 9 23 23 23 23 23 7 7 7 7 7 V4 V4 V4 V4 V4 3. 4. D J 12 7 9 Non-deletional recombination
1. 2. V4 Heptamer ligation - signal joint formation D J D J 12 12 12 7 7 7 9 9 9 12 7 3. 9 9 9 9 9 23 23 23 23 7 7 7 7 V4 V to DJ ligation - coding joint formation D J V4 4. V4 D J Fully recombined VDJ regions in same transcriptional orientation No DNA is deleted
V D J 9 7 23 12 7 9 V 9 7 23 J D 7 12 9 V 9 9 7 7 23 23 J D 7 7 12 12 9 9 Steps of TcR gene recombination Recombination activating gene products, (RAG1 & RAG 2) and ‘high mobility group proteins’ bind to the RSS The two RAG1/RAG 2 complexes bind to each other and bring the V region adjacent to the DJ region • The recombinase complex makes single stranded nicks in the DNA, the ends of each broken strand. • The nicks are ‘sealed’ to form a hairpin structure at the end of the V and D regions and a flush double strand break at the ends of the heptamers. • The recombinase complex remains associated with the break
V 9 7 23 J D 7 12 9 The hairpins at the end of the V and D regions are opened, and exonucleases and transferases remove or add random nucleotides to the gap between the V and D region DNA ligase IV joins the ends of the V and D region to form the coding joint and the two heptamers to form the signal joint. V V 9 7 23 J J D D 7 12 9 Steps of TcR gene recombination A number of other proteins, (Ku70:Ku80, XRCC4 and DNA dependent protein kinases) bind to the hairpins and the heptamer ends.
7 9 TCCACAGTG AG GTGTCAC V 23 V 9 7 23 J D 12 9 7 AT GTGACAC TA CACTGTG J D 7 12 9 TC AG V U J D AT TA U 7 9 TC AG CACAGTG GTGTCAC V 23 J D 12 9 7 AT TA GTGACAC CACTGTG Junctional diversity: P nucleotide additions The recombinase complex makes single stranded nicks at random sites close to the ends of the V and D region DNA. The 2nd strand is cleaved and hairpins form between the complimentary bases at ends of the V and D region.
V3 7 9 CACAGTG GTGTCAC 23 V2 V4 12 9 7 GTGACAC CACTGTG V5 V9 V8 V6 V7 U U D J AT TA U TC AG TC AG V V U J D AT TA Heptamers are ligated by DNA ligase IV V and D regions juxtaposed
Regions to be joined are juxtaposed The nucleotides that flip out, become part of the complementary DNA strand U U D D J J AT TA AT TA U U TC AG TC AG V V D J AT TA~TA TC~GA AG V Generation of the palindromic sequence Endonuclease cleaves single strand at random sites in V and D segment The nicked strand ‘flips’ out In terms of G to C and T to A pairing, the ‘new’ nucleotides are palindromic. The nucleotidesGA and TA were not in the genomic sequence and introduce diversity of sequence at the V to D join.
CACACCTTA Complementary bases anneal TTCTTGCAA CACACCTTA TC~GA V D J TA~TA Exonucleases nibble back free ends TTCTTGCAA D D J J AT TA~TA AT TA~TA TC~GA AG TC~GA AG V V DNA polymerases fill in the gaps with complementary nucleotides and DNA ligase IV joins the strands TC CACACCTTA TC~GA AG V V D D J AT TA~TA AG C TTCTTGCAA TA GTTAT AT Junctional Diversity – N nucleotide additions Terminal deoxynucleotidyl transferase (TdT) adds nucleotides randomly to the P nucleotide ends of the single-stranded V and D segment DNA CACTCCTTA TTCTTGCAA
V D J TCGACGTTATAT AGCTGCAATATA Junctional Diversity TTTTT TTTTT TTTTT Germline-encoded nucleotides Palindromic (P) nucleotides - not in the germline Non-template (N) encoded nucleotides - not in the germline Creates an essentially random sequence between the V region, D region and J region in beta chains and the V region and J region in alpha chains.
How does somatic recombination work? • How is an infinite diversity of specificity generated from finite amounts of DNA? • Combinatorial diversity and junctional diversity • How do V region find J regions and why don’t they join to C regions?12-23 rule • How does the DNA break and rejoin? • Imprecisely, with the random removal and addition of nucleotides to generate sequence diversity.
Vb Db Jb C 2x 1x DIVERSITY DIVERSITY Why do V regions not join to J or C regions? IF the elements of the TcR did not assemble in the correct order, diversity of specificity would be severely compromised Full potential of the beta chain for diversity needs V-D-J-C joining - in the correct order Were V-J joins allowed in the beta chain, diversity would be reduced due to loss of the imprecise join between the V and D regions
CDR3 CDR1 Location of junctional diversity TcR chain TcR chain CDR2 V-J Join V-D Join D-J join Variability Amino acid No. of TcR chain CDR = Complemantarity determining region
T-cell Receptor • T cells also express other membrane receptors that do not recognize antigen but participate in responses to antigens: these are collectively called accessory molecules.
Lck PTK Lck PTK TcR TcR CD8 CD4 3 2 MHC Class I MHC Class II CD4 and CD8 can increase the sensitivity of T cells to peptide antigen MHC complexes by ~100 fold T cell co-receptor molecules
CD8 binding site CD8 binding site MHC class I MHC class II CD8 and CD4 contact points on MHC class I and class II
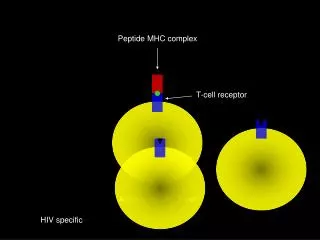
TcR CD3 CD3 TcR-CD3 complex The intracytoplasmic region of the TcR chain is too short to transduce a signal The CD3or (zeta)chains are required for cell surface expression of the TcR-CD3 complex and signalling through the TcR Signalling is initiated by aggregation of TcR by MHC-peptide complexes on APC
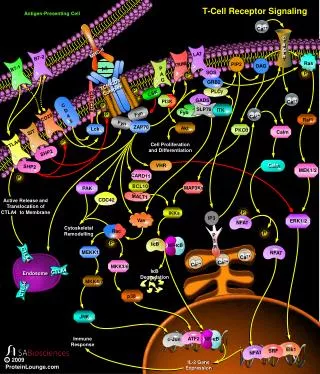
Transmission of signals from the cell surface to the nucleus Almost identical to transmission in B cells • T cell-specific parts of the signalling cascade are associated with receptors unique to T cells - TcR, CD3 etc. • Subsequent signals that transmit signals to the nucleus are common to many different types of cell. • The ultimate goal is to activate the transcription of genes, the products of which mediate host defence, proliferation, differentiation etc. • Once the T cell-specific parts of the cascade are complete, signalling to • the nucleus continues via three common signalling pathways via: • The mitogen-activated protein kinase (MAP kinase) pathway • An increase in intracellular calcium ion concentration mediated by IP3 • The activation of Protein Kinase C mediated by DAG
Simplified scheme linking antigen recognition with transcription of T cell-specific genes • MAP Kinase cascade • Small G-protein-activated MAP kinases found in all multicellular animals - activation of MAP kinases ultimately leads to phosphorylation of transcription factors from the AP-1 family such as Fos and Jun. • Increases in intracellular calcium via IP3 • IP3, produced by PLC-g, binds to calcium channels in the ER and releases intracellular stores of Ca++ into the cytosol. Increased intracellular [Ca++] activate a phospatase, calcineurin, which in turn activates the transcription factor NFAT. • Activation of Protein Kinase C family members via DAG • DAG stays associated with the membrane and recruits protein kinase C family members. The PKC, serine/threonine protein kinases, ultimately activate the transcription factor NFkB The activated transcription factors AP-1, NFAT and NFkB induce B cell proliferation, differentiation and effector mechanisms
Estimate of the number of human TcR and Ig Excluding somatic hypermutation Immunoglobulin TcR Element H 40 59 52 ~70 Variable segments 27 0 2 0 Diversity segments - - D segments in all 3 frames Yes Yes 6 9 13 61 Joining segments 2 (1)* 2 1 Joints with N & P nucleotides 2360 3640 No. of V gene pairs ~1013 ~1013 Junctional diversity ~1016** ~1016 Total diversity * Only half of human k chains have N & P regions **No of distinct receptors increased further by somatic hypermutation
Y T cell help Y Antigen presentation B B Anergy or deletion of anti-self cells Foreign antigen Y Y T T Antibody APC No T cell help Self Antigen Why do TcR not undergo somatic mutation? Affinity maturation due to somatic mutation