T Cell Receptor
T Cell Receptor. W. Robert Fleischmann, Ph.D. Department of Urologic Surgery University of Minnesota Medical School rfleisch@umn.edu (612) 626-5034. Objectives. To understand the central role of T cells To understand how T cell receptors are generated

T Cell Receptor
E N D
Presentation Transcript
T Cell Receptor W. Robert Fleischmann, Ph.D. Department of Urologic Surgery University of Minnesota Medical School rfleisch@umn.edu (612) 626-5034
Objectives • To understand the central role of T cells • To understand how T cell receptors are generated • To understand how T cell receptors function • To understand the T cell receptor complex and its function • To understand CD4 and CD8 co-receptors and their functions • To understand the alloreactivity of T cells
Central Role of T Cells • CD4+ helper T cells respond to antigen presentation to initiate the adaptive immune response. • Th1 cells that produce IFN- and IL-2 stimulate CD8+ cytotoxic T cells. • Th2 cells that produce IL-10 and IL-4 • Turn down production of Th1 cells • Stimulate mature B cells to divide and differentiate to become Ig-producing plasma cells.
Helper T Cells • Helper T cells recognize antigenic peptides that are presented by antigen-presenting cells. • Helper and cytotoxic T cells bind and recognize antigenic peptides via their T cell receptors. • Each helper and cytotoxic T cell bears a T cell receptor that recognizes one unique antigenic peptide. • In total, there exist helper T cells that have the capability of reacting with essentially every possible antigenic peptide. • The sum total of all antigenic peptides that a person’s helper T cells can recognize is called the person’s antigenic repertoire.
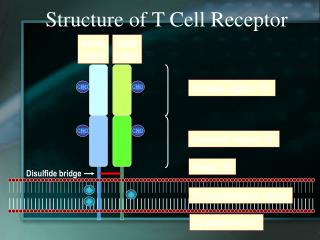
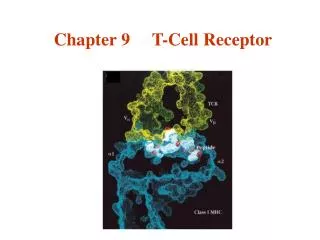
T Cell Receptors • T cell receptors (TCRs) are heterodimers • dimer • dimer • T cell receptors are highly specific for antigen. • In this way, T cell receptors are very analogous to Ig molecules on B cells. • cell receptors appear to recognize classes of antigens that are present on groups of pathogens • In this way, T cell receptors appear to be more analogous to Toll-like receptors of innate immunity (that are involved with pattern recognition) than they are with adaptive immunity.
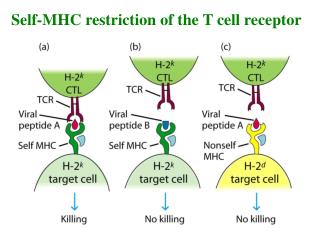
Recognition Specificity of TCR Kuby Recognizes self MHC Recognizes antigen Recognizes self MHC Fails to recognize antigen Fails to recognize MHC Cannot recognize antigen
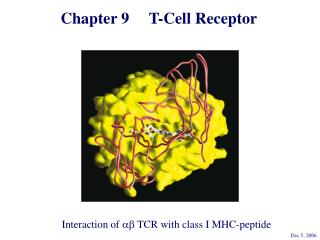
Roles of and T cells • T cells • Recognize antigen that is processed and presented by an MHC molecule. • Activate the adaptive immune response. • Can kill by granulysin and perforin.
Roles of and T cells • T cells • Require neither antigen processing nor presentation for antigen recognition. • Have very few variants. • May be more involved in the innate immune response than in the adaptive immune response (pattern recognition). • Recognize the microbial phospholipid antigen, 3-formyl-1-butyl pyrophosphate found on Mycobacterium tuberculosis, other bacteria, and parasites. • Persistent infections with Mycobacterium tuberculosis, Hemophilus influenzae, Plasmodium (malaria) and Leishmania parasites cause enhanced production of T cells that may kill by granulysin and perforin. • T cells are also prevalent in chronic autoimmune conditions, such as lupus myositis, and multiple sclerosis. • T cells produce a broad spectrum of cytokines and appear to recruit T cells to the site of pathogen invasion. • T cells may also act as antigen-presenting cells.
TCR Region of the Genome The TCR region of the genome encodes a number of sections of the TCR receptor that can be mixed and matched by recombination, analogous to Ig recombination.
TCR Gene Rearrangement • The chain, like the Ig light chain, is encoded by V, J, and C gene segments. • Rearrangement leads to a selection of a VJ combination attached to a single C gene segment. • The chain has a transmembrane cytoplasmic tail, so the TCR receptor is expressed as a membrane-bound chain. • The chain, like the Ig heavy chain, is encoded by V, D, J, and C gene segments. • Rearrangement leads to a selection of a VDJ combination attached to one of two C gene segments. • The chain has a transmembrane cytoplasmic tail, so the TCR receptor is expressed as a membrane-bound chain.
Mechanism of TCR Rearrangement • The pre-T cell expresses the recombination-activating genes, RAG-1 and RAG-2 recombinases. • The RAG-1/2 recombinase enzyme recognizes conserved recombination signal sequences flanking regions of introns between various V, D, and J coding sequences. • As for Ig rearrangement, the RAG-1/2 recombinase nicks one strand of the TCR DNA between the coding sequence and the recombination signal sequence. • The RAG-1/2 excises DNA loops. (“throw-away” loops of DNA have been found in thymocytes) • Each rearranged DNA sequence will encode a single type of TCR.
Allelic Exclusion • Each T cell will produce a single chain from just one of the chromosomal loci (allelic exclusion). • Two chains can be produced by a single cell. • Therefore two TCR sets may be expressed on a given T cell. • However, a single T cell will express a single antigen-binding specificity. How does this occur? • There are many MHC molecules that could possibly be expressed (and be recognized by TCRs). • Most TCRs will fail to recognize self and they will not survive passage through the thymus. • Only those T cells with a TCR that recognizes self will survive passage through the thymus. These T cells are said to be self-restricted T cells. • Thus, it is unlikely that two TCRs expressed on the same T cell will both be self-MHC restricted and, therefore, functional.
3294 27 3752 140 Kuby germ-line repertoire = (3,294) x (3,752) = 12,592,228; germ-line repertoire = (27) x (140) = 3,780
Generation of the TCR Repertoire • Combinatorial joining of V, D, and J region generates TCRs with many different antigen-binding specificities. • Increased diversity can be attained by alternative joining of D gene sequences to give VJ, VDJ, or VDDJ chains. • As is seen with Ig gene rearrangements, 0-6 nucleotides can be added at the junctions between V and D and between D and J. • P-region nucleotide addition: endonuclease cleavage and repair can lead to the addition of nucleotides that are palindromic. • N-region nucleotide addition: additional non-encoded nucleotides can be added by a terminal deoxynucleotide transferase. • Different from Ig gene rearrangements, somatic cell mutations do not appear to play a role in generating TCR diversity.
Kuby Note: RSS = recombination signal sequence
Importance of Lack of Somatic Mutation in Generation of Diversity • B cells generate additional diversity by somatic cell mutations that occur in the Ig region as the B cell is dividing and differentiating. This permits the generation and selection of Ig with enhanced antigen-binding affinity. • T cells lack the ability to generate somatic cell mutations. • This is good because the T cell rearrangements are selected in the thymus. • Lack of somatic cell mutations means that T cell specificity for antigen doesn’t change after the T cell leaves the thymus. • This reduces the chance of generating self-reactive TCRs when the T cell is activated (causing autoimmunity).
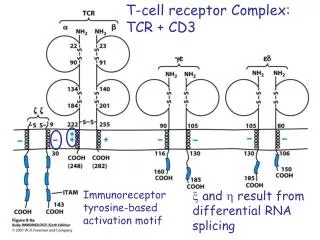
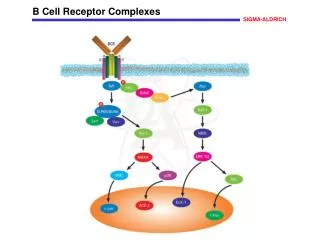
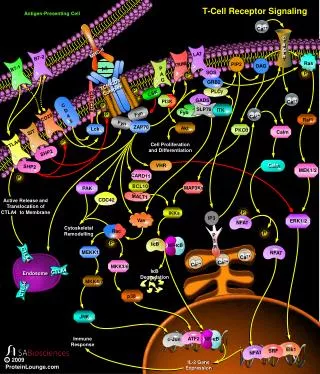
The TCR Complex • The TCR is associated with CD3, an antigen found on all T cells. • CD3 is composed of three dimers that aggregate around the TCR. • • • or where and are encoded from the same gene. Thus, they are products of the same mRNA that is differentially spliced at the 3’ end (COOH end of protein). • CD3 dimers all have a cytoplasmic signal transduction domain called ITAM (immunoreceptor tyrosine-based activation motif).
T cell Receptor Complexed with CD3 ITAM = immunoreceptor tyrosine-based activation motif Kuby
T cell Receptor Complexed with CD3 ITAM = immunoreceptor tyrosine-based activation motif Kuby
CD4 and CD8 Co-Receptors • Both CD4 and CD8 T cell co-receptors contain immunoglobulin-like domains. • CD4 molecule • A 55 kDa monomer, with 4 Ig-like domains • Binds to a conserved region on Class II antigen • Binds to signal transduction molecule p56lck and forms a bridge that also binds to the chains of CD3 • CD8 molecule • heterodimer or homodimer of 30-38 kDa monomers, with 1 Ig-like domain, held together by a disulfide bond • Binds to a conserved region on Class I antigen
Affinity of TCR for MHC/Peptide Increases with Binding of Co-receptors • The affinity of the TCR for the MHC/peptide complex is relatively modest. • CD4 or CD8 adds to binding affinity. • Other co-receptors add to binding affinity. • CD2 binds to B7 (co-stimulatory molecule) • LFA-1 (leukocyte functional antigen; aka, integrin) binds to ICAM-1 (intercellular adhesion molecule) • CD28 binds to B7 • CD45R binds to CD22
Binding Strength of T Cell Receptor • Binding strength is initially relatively weak. • Can be increased by co-receptor binding. • Lends additional specificity (CD4/CD8 binding). • Sends co-activation signals (binds to B7). • Stabilizes binding (CD45, LFA-1)
T Cell: APC Contacts Note: B7:CD28 interaction required for activation of CD8+ T cell (through activation of CD4+ T cell), but not for killing by CD8+ T cell
Alloreactivity of T Cells • CD4+ and CD8+ T cells are alloreactive to MHCs other than their cognate MHC molecule. • Alloreactive means that they immunologically react to MHCs other than their own because they are allogenic or genetically different. • Thus, MHCs are alloantigens and are the reason why allographs (transplants) are rejected. • MHCs may be directly recognized as foreign antigens (direct allorecognition). • MHCs bound to foreign proteins may be directly recognized as foreign antigens (direct allorecognition). • MHCs may be internalized, processed, and presented to the immune system bound to self-MHC (indirect allorecognition).