Central dogma: Information flow in cells
500 likes | 737 Views
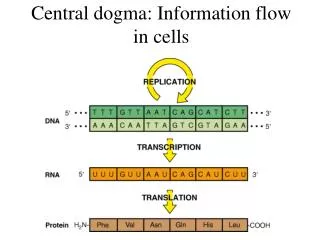
Central dogma: Information flow in cells. Nucleotides. Pyrimidine bases: Cytosine (C), Thymine (T), Uracil (U, in RNA) Purine bases: Adenine (A), Guanine (U). Prokaryotic gene coding. Eukaryotic processing of rRNA. DNA Replication: addition of a nucleotide. DNA duplex formation.

Central dogma: Information flow in cells
E N D
Presentation Transcript
Nucleotides • Pyrimidine bases: Cytosine (C), Thymine (T), Uracil (U, in RNA) • Purine bases: Adenine (A), Guanine (U)
Genetic Elements • Prokaryotes: Chromosome, plasmid, viral genome, transposable elements • Eukaryotes: Chromosomes, plasmid, mitochondrion or chloroplast genome, viral genome, transposable elements
Melting of DNA • Melting means separation of two strands from the heteroduplex • Melting temperature of DNA is dependent on the relative number of AT and GC pairs • Melted DNA can hybridize at temperatures below melting temperature • This process can be used to test relatedness between species (interspecies DNA-DNA hybridization) • It is also possible to reanneal DNA with rRNA to test relatedness of one species rRNA with the rRNA genes of another species
**DNA structure overview** • complementary strands (antiparallel) • 3 Angstrom separationof hydrogen bonds • sugar phosphate backbone held together with hydrogen bonding between bases • size is expressed in nucleotide bases pairs. E. coli has 4600 kbp. (E. coli chromosome is > 1mm, about 500X longer than the cell itself. How can the organism pack so much DNA into its cell? • each bp takes up to 0.34nm, and each helix turn is 10bp(or 34 Angstroms), therefore how long is l kb of DNA? and how many turns does it have? • inverted repeats, stem-loop, hairpins, sticky ends • supercoiled DNA (DNA-binding proteins) • relaxed, nicked circular DNA
DNA Organization • In prokaryotes: naked circular DNA with negative supercoiling • Negative supercoiling is introduced by DNA gyrase (topoisomerase II) • Topoisomerase I relaxes supercoiling by way of single-strand nicks • In eukaryotes: linear DNA packaged around histones in units called nucleosomes • The coiling around histones causes negative supercoiling
Initiation of DNA replication Origin of replication= oriC = ~300bp Templates, primers, polymerase, primase
Replication overview • 1. origin of replication+ 300 bases, recognized by specific initiation proteins = replication fork • 2. bidirectional, therefore leading and lagging strands • helicase unwinds the DNA a little (ATP-dependant) • single-strand binding protein prevents single strand from reannealing • Primase, DNA polymerase III and DNA polymerase I (also 5' to 3' exonuclease activity), ligase • Okazaki fragments • Topoisomerases, and supercoiling regulation • 3. Proofreading (3 to 5' exonuclease activity by DNA pol III)
Transcription • RNA plays an important role • tRNA, mRNA, rRNA • Name three differences between chemistry of RNA and DNA • RNA has both functional and genetic roles
Initiation of Transcription Pribnow box=tataat
More transcription • Polycistronic mRNA • How can mRNA be used in microbial ecology? • Antibiotics and RNA polymerases
RNA processing • Removal of introns • Ribozymes (nobel prize-Tom Cech and Sid Altman) • RNA-splicing enzymes • Origins of life? Which came first RNA or DNA?
The genetic code • Notice that the wobble base generally makes minor changes in the amino acid • AUG is the start code (formyl methionine) for bacteria • UAA, UAG, UGA are stop codons • Specific tRNA for each other codon
tRNA associated with codon ~60 specific tRNAs in prokaryotes
mRNA, tRNA and ribosomes Shine Dalgarno sequence GTP and Elongation Factors (EF)
Role of rRNA in protein synthesis • Structural and functional role • 16S rRNA involved in initiation • Base pairing occurs between ribosome binding sequence on the mRNA and a complementary seq on the 16S rRNA • 23S rRNA involved in elongation • Interacts with EFs
Overview of today • Summarized basic DNA structure • DNA replication • DNA sequencing • Transcription • RNA processing • Translation • Role of rRNA in protein synthesis