Chapter 2 Structure and Function of Nucleic Acids
Chapter 2 Structure and Function of Nucleic Acids. Introduction

Chapter 2 Structure and Function of Nucleic Acids
E N D
Presentation Transcript
Chapter 2 Structure and Function of Nucleic Acids • Introduction • Life depends on the genetic instructions. This hereditary information is passed on from a cell to its daughter cells at division, and from one generation of an organism to the next through the organisms’reproductive cells. Genes, the information-containing elements that determine the characteristics of a species as a whole and of the individuals within it.
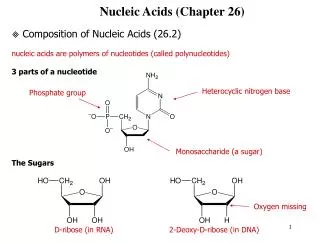
1 Chemistry of nucleic acids • Nucleotides are the subunits of DNA and RNA. • A nucleotide consists of a base, a five-carbon sugar, and one or more phosphate groups.
Purines and pyrimidines are nitrogen-containing heterocyclic compounds. • Their principal derivatives are nucleosides and nucleotides. Cytosine(C), thymine (T), and uracil (U) are called pyrimidine compounds; guaninie (G) and adenine (A) are purine compounds. Each nucleotide is named by reference to the unique base that it contains.
Diverse physiologic functions of Nucleotides • Specific nucleotides participate in reactions that fulfill physiologic functions as diverse as protein synthesis, nucleic acid synthesis, regulatory cascades, and intra- and intercellular signal transduction pathways. • A.Adenosine derivatives: AMP, ADP, ATP, and cAMP. ATP are the major biologic transducer of free energy. • B.Guanosine derivatives: cGMP serve as the principal second messenger in some cells. • C. Many coenzyme are nucleotide derives: NAD+, NADP+, FAD, etc.
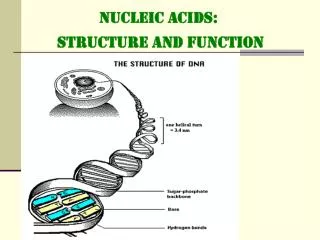
2 Deoxyribonucleic acid (DNA) The two strands of the double-helical molecule, each of which possesses a polarity, are antiparallel; ie, one strand runs in the 5’to 3’ direction and the other in the 3’ to 5’direction. The two strands, in which opposing bases are held together by hydrogen bonds, wind around a central axis in the form of a double helix. The genetic information resides in the sequence of nucleotides on one strand, the template strand. This is the strand of DNA that is copied during nucleic acid synthesis.
A always pairs with T, and G with C. This complementary base-paring enables the base pairs to be packed in the energetically most favorable arrangement in the interior of the double helix. • Each turn of DNA is made up of 10.4 nucleotide pairs and the center-to center distance between adjacent nucleotide pairs is 3.4 nm. The coiling of the two strands around each other creates two grooves in the double helix, major groove and minor groove. • B-DNA, A-DNA, and Z-DNA.
large groove small groove groove small groove largegroove Three kinds of DNA double-helix
Nucleotides absorb ultraviolet light • The conjugated double bonds of the enterocyclic bases of purines an pyrimidines, and polynucleotides absorbe ultraviolet light. Their spectra are pH-dependent. However, at pH 7.0 all the common nucleotides absorb light at a wavelength close to 260 nm.
DNA supercoiling • Packaging large DNA molecules to fit into cells requires DNA supercoiling. • Nucleosome formation • Chromatin and chromosome.
Circled DNA Supercoiled DNA
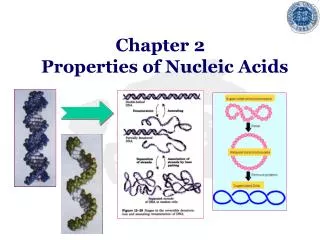
Denaturation and Renaturation of DNA • Denaturation (melting) and Renaturation of DNA • PCR (Polymerase Chain Reaction) • cDNA (complementary DNA) • Southern blotting (DNA/cDNA) • Northern blotting (DNA/RNA). • Western Blotting
Figure 4-8. The denaturation and renaturation of double-stranded DNA molecules.
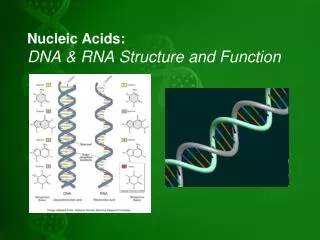
Ribonucleic acid (RNA) RNA is a polymer of purine and pyrimidine ribonucleotides linked together by 3’, 5’-phosphodiester bridges analogous to those in DNA. Although sharing many features with DNA, RNA possesses several specific differences. • (1) Bases are attached to ribose rather than 2’- deoxyribose. • (2) U, not T. • (3) Single strand. however, capable of folding back on itself like a hairpin. • (4) Others
Those cytoplasmic RNA molecules that serve as templates for protein synthesis are designated messenger RNAs, or mRNA. Many other cytoplasmic RNA molecules (ribosomal RNAs, or rRNA) have structural roles wherein they contribute to the formation of ribosomes ( the organellar machinery for protein synthesis) or serve as adapter molecules (transfer RNAs; tRNAs) for the translation of RNA information into specific sequences of polymerized amino acids.
Some RNA molecules have intrinsic catalytic activity. The activity of these ribozymes often involves the cleavage of a nucleic acid. An example is the role of RNA in catalyzing the processing of the primary transcript of a gene into mature messenger RNA. • In human cells, there are small nuclear RNA (snRNA) species that are not directly involved in protein synthesis but that may have roles in RNA processing and the cellular architecture. These relatively small molecules vary in size from 90 to about 300 nucleotides.
mRNA • This class is the most heterogeneous in size and stability. mRNAs, particularly in eukaryotes, have some unique chemical characteristics. The 5’terminal of mRNA is “capped”by a 7-methylguanosine triphosphate that is linked to an adjacent 2’-0 –methyl rebonucleotide at its 5’-hydroxyl through the three phosphates. The other end of most mRNA molecules, the 3’-hydroxyl terminal, has attached a polymer of adenylate residues 20-250 nucleotides in length.
The mRNA molecules present in the cytoplasm are not the RNA products immediately synthesized from the DNA template but must be formed by processing from a precursor molecule before entering the cytoplasm. Thus, in mammalian nuclei, the immediate products of gene transcription constitute a fourth class of RNA molecules. These nuclear RNA molecules are very heterogeneous in size and are quite large. The heterogeneous nuclear RNA (hnRNA) molecules may have a molecular weight in excess of 107, whereas the molecule weight of mRNA molecules is generally less than 2×106.
mRNA Structure of Eucaryotes Coding region 3‘-untranslation region 5‘-untranslation region
Cap structure (Only on some caps) 7-methylguanosine triphosphate cap
Figure 4-18. Structure of the 5′ methylated cap of eukaryotic mRNA.
Figure 4-19. Overview of RNA processing in eukaryotes using β-globin gene as an example. The β-globin gene contains three protein-coding exons (red) and two intervening noncoding introns (blue). The introns interrupt the protein-coding sequence between the codons for amino acids 31 and 32 and 105 and 106. Transcription of this and many other genes starts slightly upstream of the 5′ exon and extends downstream of the 3′ exon, resulting in noncoding regions (gray) at the ends of the primary transcript. These regions, referred to as untranslated regions (UTRs), are retained during processing. The 5′ 7-methylguanylate cap (m7Gppp; green dot) is added during formation of the primary RNA transcript, which extends beyond the poly(A) site. After cleavage at the poly(A) site and addition of multiple A residues to the 3′ end, splicing removes the introns and joins the exons. The small numbers refer to positions in the 147-aa sequence of β-globin.
Figure 4-30. Recognition of a tRNA by aminoacyl synthetases. Aspartyl-tRNA synthetase (AspRS) is a class II enzyme, and arginyl-tRNA synthetase (ArgRS) is a class I enzyme.
tRNA • tRNA molecules vary in length from 74-95 nucleotides. There are at least 20 species of tRNA molecules in every cell, at least one (and often several) corresponding to each of the 20 amino acids required for protein synthesis.
tRNA structure CCA end D arm Tψ arm Anticodon arm Anticodon
Figure 6-11. Amino acid activation. The two-step process in which an amino acid (with its side chain denoted by R) is activated for protein synthesis by an aminoacyl-tRNA synthetase enzyme is shown. As indicated, the energy of ATP hydrolysis is used to attach each amino acid to its tRNA molecule in a high-energy linkage. The amino acid is first activated through the linkage of its carboxyl group directly to an AMP moiety, forming an adenylated amino acid;the linkage of the AMP, normally an unfavorable reaction, is driven by the hydrolysis of the ATP molecule that donates the AMP. Without leaving the synthetase enzyme, the AMP-linked carboxyl group on the amino acid is then transferred to a hydroxyl group on the sugar at the 3' end of the tRNA molecule. This transfer joins the amino acid by an activated ester linkage to the tRNA and forms the final aminoacyl-tRNA molecule. The synthetase enzyme is not shown in these diagrams.
Figure 4-29. Aminoacylation of tRNA. Amino acids are covalently linked to tRNAs by aminoacyl-tRNA synthetases.
Figure 4-22. Assigning codons using synthetic mRNAs containing a single ribonucleotide. Addition of such a synthetic mRNA to a bacterial extract that contained all the components necessary for protein synthesis except mRNA resulted in synthesis of polypeptides composed of a single type of amino acid as indicated.
rRNA A ribosome is a cytoplasmic nucleoprotein structure that acts as the machinery for the synthesis of proteins from the mRNA templates. On the ribosomes, the mRNA and tRNA molecules interact to translate into a specific protein molecule information transcribed from the gene.
Figure 4-32. The general structure of ribosomes in prokaryotes and eukaryotes.
Figure 4-33. Two-dimensional map of the secondary structure of the small (16S) rRNA from bacteria, showing the location of base-paired stems and loops. In general, the length and position of the stem-loops are very similar in all species, although the exact sequence varies from species to species. The most highly conserved regions are represented as red lines, and the numbered stem-loops unique to prokaryotes are preceded by a P. Eukaryotic small (18S) rRNAs exhibit a generally similar pattern of stem-loops, although, as with prokaryotes, a few are unique.
Figure 4-25. Translation of nucleic acid sequences in mRNA into amino acid sequences in proteins requires a two-step decoding process. First, an aminoacyl-tRNA synthetase couples a specific amino acid to its corresponding tRNA. Second,a three-base sequence in the tRNA (the anticodon) base-pairs with a codon in the mRNA specifying the attached amino acid. If an error occurs in either step, the wrong amino acid may be incorporated into a polypeptide chain.