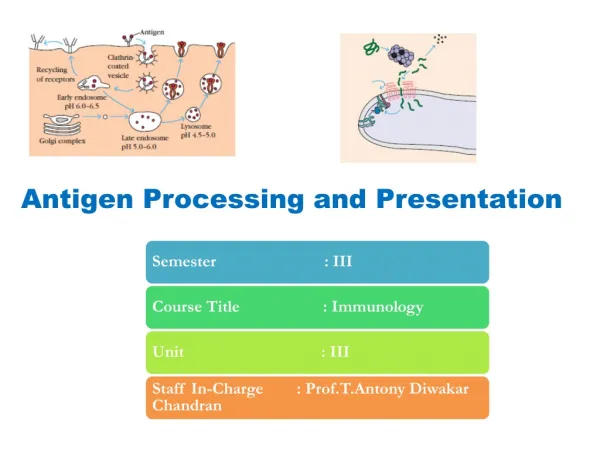
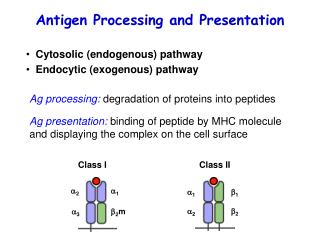
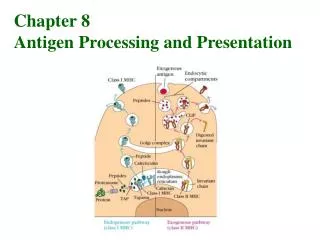
Antigen Processing and Presentation
Antigen Processing and Presentation. January 23, 2007. 2-1 Table 1 2-1 Fig 1 2-1 Fig 2 2-1 Fig 3, 4 2-1 Fig 5 2-1 Table II 2-1 Table III 2-2 Table I,II 2-2 Table III, IV, Fig 1 2-2 Table V, VI, VII 2-3 Fig 1,2 2-3 Fig 3,4 2-3 Fig 5 2-3 Fig 6 (no detail).

Antigen Processing and Presentation
E N D
Presentation Transcript
Antigen Processing and Presentation January 23, 2007
2-1 Table 1 • 2-1 Fig 1 • 2-1 Fig 2 • 2-1 Fig 3, 4 • 2-1 Fig 5 • 2-1 Table II • 2-1 Table III • 2-2 Table I,II • 2-2 Table III, IV, Fig 1 • 2-2 Table V, VI, VII • 2-3 Fig 1,2 • 2-3 Fig 3,4 • 2-3 Fig 5 • 2-3 Fig 6 (no detail)
4-1. Kirk Ziegler and Emil Unanue Identification of a macrophage antigen-processing event required for I-region-restricted antigen presentation to T lymphocytes. J. Immunol. 127:1869-1875. (1981).
Macrophages (M) • M are needed for CD4+ T cell activation • MHC restriction between T cell and M • M are known to degrade bacteria • General thought is that M simply degrade antigens • How is antigen “handled” by a M
Conclusions-I • First demonstration of antigen processing • An antigen processing period of ~15 was required, after which time all of the necessary signals for T cell activation were on the cell surface of the APC • Antigen processing required active APC metabolism • Subsequent, antigen processing was shown to require an acidic compartment (i.e. inhibited by treatment with chloroquine)
Jan Klein and the controversy over antigen processing “To avoid giving the impression of claiming that I was always right, I can provide an example wherein Time, the corrector upended my judgment. Several years ago, during my digression into immunology, I became involved in a controversy over antigen processing, the notion that foreign proteins must first be degraded inside a cell before they can be presented to T lymphocytes. One could say that I created the controversy by questioning whether the evidence available then warranted the universal acceptance of the antigen processing hypothesis and by claiming that our own data went against the hypothesis. On the latter point I was wrong. Antigen processing is now a well-established concept and Emil R. Unanue, Paul M. Allen, Howard M. Grey, Alain Townsend and others deserve all the credit for developing it. My attempt to deny antigen processing was perhaps my most spectacular, but not my only blunder. Nonetheless, I do not regret having challenged the antigen processing hypothesis, and if we had at hand today that knowledge which was available then, I would do so again...” Jan Klein, Alone on the heart of the earth: an immunogeneticist’s journey into the past. Adv. Cancer Research 63: 1-39, 1994.
The CD4-CD8 divide • CD8 cells were generated during a viral infection • CD8-target cell interaction was MHC restricted • CD4 cells were generated during a bacterial or model antigen challenge • CD4-APC interaction was MHC restricted Therefore the consensus was that CD4 and CD8 T cells were recognizing antigens that distinct in source (virus vs bacteria) and cell location (membrane vs. soluble)
Morrison 4-2. Lynda Morrison, Aaron Lukacher, Vivian Braciale, David Fan, and Tom Braciale Differences in antigen presentation to MHC class I- and class II-restricted influenza virus-specific cytolytic T lymphocyte clones. J. Exp. Med. 163:903-921. (1986).
Conclusions-II • First study to show that class I and class II processing pathways were distinct • Showed that class I pathway was not acidic, but required protein synthesis • Showed for the same viral antigen that both CD4 and CD8 T cells could be generated, thus uniting the view of how antigen is handled by APCs and target cells.
What was the nature of the antigen recognized by CD8 T cells? • Townsend-trained with Ita Askonas who collaborated with John Skehel @ Mill Hill, started his own lab at Oxford • Generated CD8+ T cells specific for influenza nucleoprotein (NP), which is not expressed on the surface of an infected cell • In a study previous to this, showed that NP produced in the cytoplasm was sufficient to charge the class I pathway, thereby establishing that the source of antigen was the cytoplasm
Townsend 4-3. Alain Townsend, Claes Öhlén, Judy Bastin, Hans-Gustaf Ljunggren, Linda Foster and Klas Kärre Association of class I major histocompatibility heavy and light chains induced by viral peptides. Nature 340:443-448. (1989).
RMA-S • Generated by Klas Karre to explore the missing self-hypothesis of NK cell activation • RBL-5 cell line-Rauscher virus induced H-2b lymphoma • EMS mutagenized and selected for low class I by multiple rounds of anti-class I + C’
Conclusions-III • RMA-S class I expression can be restored by peptides • Class I restoration is allele specific • RMA-S + peptide restores association of class I heavy chains and 2m • Proposed that the RMA-S defect was due to reduced peptide availability during class I folding • RMA-S and T2 cell lines were instrumental in further dissecting and characterizing the class I pathway