Rapid Detection of Extensively Drug-Resistant Tuberculosis Using Pyrosequencing
10 likes | 144 Views
This study evaluates the efficiency of pyrosequencing (PSQ) as a diagnostic tool for rapidly identifying mutations related to extensively drug-resistant tuberculosis (XDR-TB). We designed 8 assays targeting specific genes in the Mycobacterium tuberculosis to aid in the detection of drug resistance. Testing 166 isolates from high-prevalence regions, our findings suggest that PSQ provides reliable and timely results, showing over 90% agreement with traditional drug susceptibility testing methods, thus enhancing early detection and treatment strategies for XDR-TB.

Rapid Detection of Extensively Drug-Resistant Tuberculosis Using Pyrosequencing
E N D
Presentation Transcript
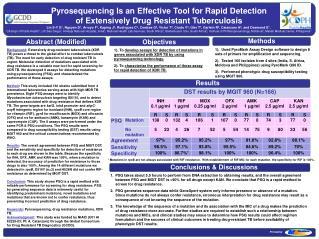
Pyrosequencing Is an Effective Tool for Rapid Detection of Extensively Drug Resistant TuberculosisLin S-Y G1, Nguyen D1, Arroyo F1, Kaping J2, Rodrigues C3, Coetzee G4, Victor T5, Crudu V6, Gler T7, Garfein R2, Catanzaro A2, and Desmond E1 . CA Dept of Public Health1, UC San Diego2, Hinduja National Hospital, India3, National Health Lab Services, South Africa4, Stellenbosch Univ, South Africa5, Institute of Phthisiopneumology, Moldova6, Makati Medical Center, Philippines7 Methods Abstract (Modified) Objectives 1). Used PyroMark Assay Design software to design 8 sets of primers for amplification and sequencing. 2). Tested 166 isolates from 4 sites (India, S. Africa, Moldova and Philippines) using PyroMark Q96 ID. 3). Performed phenotypic drug susceptibility testing using MGIT 960. Background: Extensively drug-resistant tuberculosis (XDR TB) poses a threat to the global effort to combat tuberculosis (TB). The need for early detection of drug resistant TB is urgent. Molecular detection of mutations associated with drug resistance is a valuable new tool for rapid screening for XDR TB. We developed 8 assays for detecting mutations using pyrosequencing (PSQ), and characterized the performance of these assays. Method: This study included 166 strains submitted from 4 international laboratories serving areas with high MDR TB prevalence. Eight PSQ assays were to identify Mycobacterium tuberculosis targeting IS6110, and to detect mutations associated with drug resistance that defines XDR TB. The gene targets are katG, inhA promoter and ahpC-oxyR intergenic region for isoniazid (INH), rpoB core region for rifampin (RIF), gyrA for moxifloxacin (MOX) and ofloxacin (OFX) and rrs for amikacin (AMK), kanamycin (KAN) and capreomycin (CAP). The 8 assays were performed under the same PCR & PSQ conditions. The PSQ results were compared to drug susceptibility testing (DST) results using MGIT 960 and the critical concentrations recommended by WHO. Results: The overall agreement between PSQ and MGIT DST, and the sensitivity and specificity for detection of resistance to each drug are shown in the table. Because the specificity for INH, OFX, AMK and KAN was 100%, when a mutation is detected, the accuracy of prediction for resistance to those drugs is also 100%. Among the 14 different mutations we detected in rpoB, D516Y (n=3) and H526N did not confer RIF-resistance as determined by MGIT DST. Conclusion: This study shows PSQ is a rapid method with reliable performance for screening for drug resistance. PSQ by generating sequence data is extremely useful for identifying predominant mutations, novel mutations and mutations that are known not to confer resistance, thus preventing incorrect prediction of drug resistance. Keywords: Pyrosequencing, drug resistance mutations, XDR TB. Acknowledgment: This study was funded by NIAID (U01 AI 82229-03; PI: A. Catanzaro) through the Global Consortium for Drug Resistant TB Diagnostics (GCDD). 1). To develop assays for detection of mutations in genes associated with XDR TB by using pyrosequencing technology. 2). Tocharacterize the performance of those assay for rapid detection of XDR TB. Results Objectives * Mutations in rpoB are not always associated with RIF resistance. With establishment of RIF MIC for each mutation, the specificity for RIF is 100%. Conclusions & Discussions • PSQ takes about 5.5 hours to perform from DNA extraction to obtaining results, and the overall agreement between PSQ and MGIT DST is >90% for all drugs except KAN. We conclude that PSQ is a rapid method to screen for drug resistance. • PSQ generates sequence data while GeneXpert system only informs presence or absence of a mutation. Since mutations do not always confer resistance, erroneous interpretation for drug resistance may result as a consequence of not knowing the sequence of the mutation. • The knowledge of the sequence of a mutation and its association with the MIC of a drug makes the prediction of drug resistance more accurate. Further studies are required to establish such a relationship between mutations and MICs, and clinical studies may ensue to determine how PSQ results could affect regimen formulation and the success of clinical outcomes in treating drug-resistant TB before availability of phenotypic DST results. Printed by