HYBRIDIZATION
240 likes | 880 Views
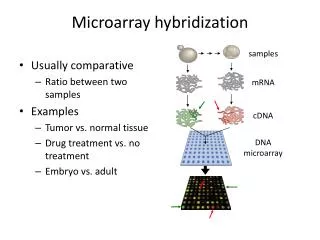
HYBRIDIZATION. By. Dr. Emad AbdElhameed Morad. Lecturer of Medical Microbiology and Immunology. Denaturation or melting: Separation of the two DNA strands from each other. Melting could be done by either: Heat (90-100 °c) Alkaline pH Low salt concentration

HYBRIDIZATION
E N D
Presentation Transcript
HYBRIDIZATION By Dr. Emad AbdElhameed Morad Lecturer of Medical Microbiology and Immunology
Denaturation or melting: Separation of the two DNA strands from each other. Melting could be done by either: • Heat (90-100 °c) • Alkaline pH • Low salt concentration • Renaturation or annealing: Reassociation of the two DNA strands to form a double helix. Annealing could be done by : • Lowering the temperature • Neutralization of pH • Raising the salt concentration
Nucleic acid probe: • A short known sequence of single stranded DNA or RNA designed to hybridize with the target nucleic acid. • Nucleic acid probes (known) are thus used in identification of microbes (unknown). • The probes are labeled to facilitate their detection following hybridization.
Hybridization based assays Solution phase hybridization Solid phase hybridization Dot blot In situ hybridization Blotting techniques FISH Southern Northern Western
Dot blot The bacterial cells are lysed to liberate DNA. The DNA is treated by alkali or heat to separate the two strands. The DNA strands are attached to nitrocellulose membrane. Labeled probe is then applied to the membrane. If the probe is complementary to the test DNA, it will form a hybrid duplex with the test DNA and remain on the membrane after washing. Formation of hybrid DNA is proved by detecting the label conjugated to the probe.
Stringency of hybridization The temperature, salt concentration and sequence homology forms the conditions of stringency for hybridization. Stringency increases as the salt concentration decreases and temperature increases. More stringent conditions permit hybridization of only highly homologous sequence. A single base mismatch in a stretch of 20 bases may prevent hybridization under the most stringent conditions. RNA-RNA duplexes are more stable than RNA-DNA duplexes, which are more stable than DNA-DNA duplexes.
Labels of probes Labeling can be done using radioactive or non-radioactive labels. The common radioactive isotopes used for labeling include P 32, S 35, I 125. Once hybridized, the labeled probes can be detected by scintillation counter or on X-ray autoradiography. Disadvantages include: Higher expense Difficulty in handling Health hazard
Radioactive labels of probes The upper lane is +Ve The lower lane is -Ve
The non-radioactive labels include biotin, digoxygenin , acridinium esters. • Biotin binds specifically to avidin. Avidin is tagged with enzymes. Addition of substrate results in production of colouredproduct, signaling the positive hybridization reaction. • Avidin may be conjugated with fluorescein. Hybridization is observed for fluorescence using UVL. • Detection of hybridization may be done by using fluorescein conjugated antibodies against biotin.
Digoxygenin labeled probes are detected by enzyme labeled anti-digoxygenin antibody and then using substrate to detect it. • Probes can also be labeled with acridinium ester. • Detection is achieved by the addition of hydrogen peroxide hydroxide, which results in hydrolysis of the ester linkage. • The light that is produced in this reaction is detected using a chemiluminometer.
In situ hybridization Hybridization process performed on intact tissues or cells fixed to a glass microscopic slide. Fluorescence in situ hybridization (FISH): In situ hybridization (ISH) using probe labeled with a fluorescent dye. Examination is done using fluorescent microscope.