Comparing Two Protein Sequences
Comparing Two Protein Sequences. Cédric Notredame. Our Scope. Look once Under the Hood. Pairwise Alignment methods are LIMITED. If You Understand the LIMITS they Become VERY POWERFUL. Pairwise Alignment methods are POWERFUL. Outline. -WHY Does It Make Sense To Compare Sequences.

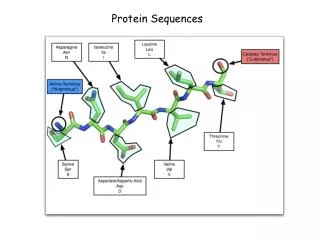
Comparing Two Protein Sequences
E N D
Presentation Transcript
Comparing Two Protein Sequences Cédric Notredame
Our Scope Look once Under the Hood Pairwise Alignment methods are LIMITED If You Understand the LIMITS they Become VERY POWERFUL Pairwise Alignment methods are POWERFUL
Outline -WHY Does It Make Sense To Compare Sequences -HOW Can we Compare Two Sequences ? -HOW Can we Align Two Sequences ? -HOW can I Search a Database ?
Why Does It Make Sense To Compare Sequences ? Sequence Evolution
Why Do We Want To Compare Sequences wheat --DPNKPKRAMTSFVFFMSEFRSEFKQKHSKLKSIVEMVKAAGER | | |||||||| || | ||| ||| | |||| |||| ????? KKDSNAPKRAMTSFMFFSSDFRS----KHSDL-SIVEMSKAAGAA EXTRAPOLATE Homology? SwissProt ??????
Why Does It Make Sense To Align Sequences ? -Evolution is our Real Tool. -Nature is LAZY and Keeps re-using Stuff. -Evolution is mostly DIVERGEANT Same Sequence Same Ancestor
Why Does It Make Sense To Align Sequences ? Same Sequence Same Function Same Origin Same 3D Fold Many Counter-examples!
An Alignment is a STORY ADKPKRPLSAYMLWLN ADKPKRPLSAYMLWLN ADKPKRPLSAYMLWLN Mutations + Selection ADKPKRPKPRLSAYMLWLN ADKPRRPLS-YMLWLN
ADKPKRPLSAYMLWLN ADKPKRPLSAYMLWLN ADKPKRPLSAYMLWLN Mutations + Selection ADKPKRPKPRLSAYMLWLN ADKPRRPLS-YMLWLN ADKPRRP---LS-YMLWLN ADKPKRPKPRLSAYMLWLN Deletion Insertion Mutation An Alignment is a STORY
Evolution is NOT Always Divergent… AFGP with (ThrAlaAla)n Similar To Trypsynogen N S AFGP with (ThrAlaAla)n Chen et al, 97, PNAS, 94, 3811-16 NOT Similar to Trypsinogen
Evolution is NOT Always Divergent AFGP with (ThrAlaAla)n Similar To Trypsynogen N S AFGP with (ThrAlaAla)n NOT Similar to Trypsinogen SIMILAR Sequences BUT DIFFERENT origin
Evolution is NOT always Divergent… Same Sequence Same Origin Same Function Same 3D Fold But in MOST cases, you may assume it is… Similar Function DOES NOT REQUIRE Similar Sequence Similar Sequence Historical Legacy
How Do Sequences Evolve Each Portion of a Genome has its own Agenda.
How Do Sequences Evolve ? Family KSKA Histone3 6.4 0 Insulin 4.0 0.1 Interleukin I 4.6 1.4 a-Globin 5.1 0.6 Apolipoprot. AI 4.5 1.6 Interferon G 8.6 2.8 Rates in Substitutions/site/Billion Years as measured on Mouse Vs Human (80Million years) Ks Synonymous Mutations, Ka Non-Neutral. CONSTRAINED Genome Positions Evolve SLOWLY EVERY Protein Family Has its Own Level Of Constraint
Different molecular clocks for different proteins--another prediction
How Do Sequences Evolve ? The amino Acids Venn Diagram C P L V Small A G G I Aliphatic C C T S D N K Y E F H Q W R Aromatic Hydrophobic Polar To Make Things Worse, Every Residue has its Own Personality
How Do Sequences Evolve ? + - - In the core, SIZE MATTERS On the surface, CHARGE MATTERS OmpR, Cter Domain In a structure, each Amino Acid plays a Special Role
How Do Sequences Evolve ? Big -> Big Small ->Small NO DELETION Charged -> Charged Small <-> Big or Small DELETIONS Accepted Mutations Depend on the Structure + - -
How Can We Compare Sequences ? Substitution Matrices
How Can We Compare Sequences ? Their Structure We Do Not Have Them !!! Their Function To Compare Two Sequences, We need:
How Can We Compare Sequences ? Same Sequence We will Need To Replace Structural Information With Sequence Information. Same Origin Same Function Same 3D Fold It CANNOT Work ALL THE TIME !!!
How Can We Compare Sequences ? How to derive that matrix? To Compare Sequences, We need to Compare Residues We Need to Know How Much it COSTS to SUBSTITUTE an Alanine into an Isoleucine a Tryptophan into a Glycine … The table that contains the costs for all the possible substitutions is called the SUBSTITUTION MATRIX
How Can We Compare Sequences ? Using Knowledge Could Work C P Small L V A G G Aliphatic I C C T S D N K Y E F H Q Aromatic W R Hydrophobic Polar But we do not know enough about Evolution and Structure. Using Data works better.
How Can We Compare Sequences ? Making a Substitution Matrix Observed Log Expected by chance -Take 100 nice pairs of Protein Sequences, easy to align (80% identical). -Align them… -Count each mutations in the alignments -25 Tryptophans into phenylalanine -30 Isoleucine into Leucine … -For each mutation, set the substitution score to the log odd ratio:
You’re kidding! … I was struck by a lightning twice too!! Garry Larson, The Far Side
How Can We Compare Sequences ? Making a Substitution Matrix The Diagonal Indicates How Conserved a residue tends to be. W is VERY Conserved Some Residues are Easier To mutate into other similar Cysteins that make disulfide bridges and those that do not get averaged
How Can We Compare Sequences ? Making a Substitution Matrix
How Can We Compare Sequences ? Using Substitution Matrix Given two Sequences and a substitution Matrix, We must Compute the CHEAPEST Alignment ADKPRRP---LS-YMLWLN ADKPKRPKPRLSAYMLWLN Deletion Insertion Mutation
Scoring an Alignment Raw Score TPEA ¦| | APGA Score = + 6 + 0 + 2 = 9 • Question: Is it possible to get such a good alignment by chance only? • Most popular Subsitution Matrices • PAM250 • Blosum62 (Most widely used) 1
Gap Opening Penalty Gap Extension Penalty gap Insertions and Deletions • Gap Penalties • Opening a gap is more expensive than extending it Seq A GARFIELDTHE----CAT ||||||||||| ||| Seq B GARFIELDTHELASTCAT
How Can We Compare Sequences ? Limits of the substitution Matrices They ignore non-local interactions and Assume that identical residues are equal ADKPKRPLSAYMLWLN They assume evolution rate to be constant ADKPKRPLSAYMLWLN ADKPKRPLSAYMLWLN Mutations + Selection ADKPKRPKPRLSAYMLWLN ADKPRRPLS-YMLWLN
How Can We Compare Sequences ? Limits of the substitution Matrices Substitution Matrices Cannot Work !!!
How Can We Compare Sequences ? Limits of the substitution Matrices I know… But at least, could I get some idea of when they are likely to do all right
How Can We Compare Sequences ? The Twilight Zone Similar Sequence Similar Structure Different Sequence Structure ???? 30% %Sequence Identity Same 3D Fold 30 Twilight Zone Length 100
How Can We Compare Sequences ? The Twilight Zone Substitution Matrices Work Reasonably Well on Sequences that have more than 30 % identity over more than 100 residues
How Can We Compare Sequences ? Which Matrix Shall I used Other Matrices Exist: BLOSUM 42 BLOSUM 62 BLOSUM 62 The Initial PAM matrix was computed on 80% similar Proteins It been extrapolated to more distantly related sequences. Pam 250 Pam 350
How Can We Compare Sequences ? Which Matrix Shall I use Choosing The Right Matrix may be Tricky… • GONNET 250> BLOSUM62>PAM 250. • But This will depend on: • The Family. • The Program Used and Its Tuning. • Insertions, Deletions? PAM: Distant Proteins High Index (PAM 350) BLOSUM: Distant Proteins Low Index (Blosum30)
HOW Can we Align Two Sequences ? Dot MatricesGlobal Alignments Local Alignment
Dot Matrices QUESTION What are the elements shared by two sequences ?
>Seq1 THEFATCAT >Seq2 THELASTCAT T H E F A T C A T T Window H E Stringency F A S T C A T Dot Matrices
Window size Sequences Stringency Dot Matrices
Window=1Stringency=1 Window=11Stringency=7 Window=25Stringency=15 Dot Matrices Strigency