Optimizing Chromatographic Methods: A Factorial Design Tutorial by Dr. Jeff Hughes
This tutorial, presented by Dr. Jeff Hughes of RMIT University, demonstrates how experimental design techniques, specifically factorial designs, can enhance chromatographic method development. The tutorial addresses crucial questions regarding factor influence on peak separation in chromatograms. Participants will explore two-level factorial designs, learn to identify significant factors, and maximize chromatographic response using tools like Excel and Minitab. The tutorial is aimed at researchers seeking to optimize High-Performance Liquid Chromatography (HPLC) methods.

Optimizing Chromatographic Methods: A Factorial Design Tutorial by Dr. Jeff Hughes
E N D
Presentation Transcript
What Shape is your Method In? A Tutorial on the Application of Experimental Designs to Development of Chromatographic Methods By Dr Jeff Hughes School of Applied Science, RMIT University Melbourne , Australia
The aim of this tutorial is to demonstrate how the methods of experimental design can be used to investigate chromatographic procedures In this tutorial we will look at using Factorial Designs to answer the following questions: Which factors significantly influence the of separation of peaks in our chromatogram? Which factor has the greatest influence on separation? RMIT University
Data for this tutorial is taken from “Chemometrics: Experimental Design” by Ed Morgan Calculations are demonstrated using Excel, but can be carried out using the commercial program Minitab ( a demo version can be downloaded from http://minitab.com) RMIT University
Factorial Designs • The most suitable type of design for screening • Each variable (factor) has a set number of possible levels or values • If there are k variables, each set at 2 possible levels (‘high’ and ‘low’) then there are 2k possible combinations • These designs are called two-level factorial designs. If all combinations are used they are called full factorial designs RMIT University
Screening • Aim – identify significant factors (variables) • A factor is ‘significant’ if its influence is greater than the ‘noise’ level (experimental error) • Usually carry out screening using reduced designs such as factorial or Plackett-Burman designs RMIT University
The trials in a factorial design can be represented as points on an n-dimensional cube (n=3 in this case) 1,-1,1 1,1,1 11,-1 1,-1,-1 -1,-1,1 -1,1,1 -1,-1,-1 -1,1,-1 RMIT University
Case Study – HPLC method • Aim: to optimise the separation of peaks in a HPLC analysis RMIT University
Define the Response • The CRF (chromatographic response function) is used to quantify separation of peaks. This function thus gives a single number to the ‘quality’ of a chromatogram. The aim is thus to maximise the CRF RMIT University
Define the Factors • The factors studied in this study were levels in the eluent of:- • Acetic Acid • Methanol • Citric Acid RMIT University
Experimental Domain RMIT University
Factorial design (Coded form) This design gives all combinations of the factors at 2 levels ‘+’, high ‘-’, low RMIT University
Factorial design (Uncoded) This table shows the actual levels of the variables used in the experiments. Normally the order of experiments is randomised but we will keep it in this structured forms so you can see the patterns Results are inserted here when the experiments are performed RMIT University
Factorial design (Uncoded) The CRF values are now inserted after the experiments (chromatographic runs) are carried out RMIT University
Analysis of the results - Excel • Calculate Main Effects – this calculates the effect on the response solely due to one factor • Main effects are the difference between average response at high level of the factor – average response at low level RMIT University
Calculation of Main Effects The main effects can also be calculated by multiplying the variable column by the CRF column pairwise , adding up the column and then dividing by 4 Average of the ‘high’ values of CRF for each variable e.g for AA = (9.5+10.7+8.8+11.7)/4 Average of ‘low’ values of CRF for each variable e.g for AA = (10+11+9.3+11.9)/4 The ’Main Effect’ is the difference between the ‘high’ and ‘low’ average e.g for AA = (10.19-10.55) RMIT University
Calculation of Interactions Interactions coefficients –found by multiplying the appropriate variable columns Interactions calculated by multiplying the CRF column and the appropriate variable interactions column. To get the interaction effect add up the column and divide by 4 RMIT University
Factorial Calculations using Minitab • The program ‘Minitab’ can be used to carry out calculations as follows:- • To set up the design: Stat > DOE > Factorial > Create Factorial Design Type of Design: 2 level factorial design 9default generators) Number of factors: 3 Designs: Full Factorial Factors see screen dump on next slide Options: do not randomize (normally should randomize but for the tutorial not randomizing makes it easier to see patterns in the layout) RMIT University
The generated FFD design as it should appear in Minitab Type the CRF responses here after performing the experiments RMIT University
Factorial Calculations using Minitab • To analyse the design: • Stat > DOE > Factorial > Analyse factorial Design • Click on ‘C8 CRF’ as the response . Accept the default values RMIT University
Factorial Calculations - Minitab • 31/07/2007 11:26:30 ———————————————————— • Welcome to Minitab, press F1 for help. • Results for: Worksheet 2 • Full Factorial Design • Factors: 3 Base Design: 3, 8 • Runs: 8 Replicates: 1 • Blocks: 1 Center pts (total): 0 • All terms are free from aliasing. • Design Table • Run A B C • 1 - - - • 2 + - - • 3 - + - • 4 + + - • 5 - - + • 6 + - + • 7 - + + • 8 + + + Minitab Output RMIT University
Factorial Fit: CRF versus Acetic Acid, Methanol, Citric Acid Estimated Effects and Coefficients for CRF (coded units) Term Effect Coef Constant 10.3625 Acetic Acid -0.3750 -0.1875 Methanol 1.9250 0.9625 Citric Acid 0.1250 0.0625 Acetic Acid*Methanol 0.1250 0.0625 Acetic Acid*Citric Acid 0.0250 0.0125 Methanol*Citric Acid 0.8250 0.4125 Acetic Acid*Methanol*Citric Acid 0.0250 0.0125 Note , however, coefficients are simply twice the effects so no new information Main Effects Interactions Coefficients – from fitting to a second order equation Y = bo+b1*x1+b2*x2+b3*x3+b12*x1*x2+b13*x1*x3+b23*x2*x3+b123*x1*x2*x3 Where x1 is acetic acid , x2 is methanol and x3 is citric acid RMIT University
What do the results tell us? • The main effects tell us which variable has the strongest effect on the response (CRF) – in this case methanol has the strongest effect on CRF • A negative effect means the response is reduced as the variable increases. The negative effect for acetic acid means that as we increase the concentration of acetic acid, the CRF gets smaller (and hence our separation is worse) RMIT University
What about interactions? • An interaction effect is where the effect on the response of one variable depends on the level of another variable. • In this study methanol and citric acid seem to have the largest interaction. RMIT University
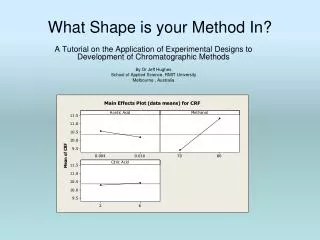
Main Effects and Interactions Plots • Main effects plots help to visually display the variable effect. They graph the average response at the high and low levels. The steeper the graph, the stronger the effect • The plots can be drawn in Miniab as follows:- • Stat > DOE >Factorial > factorial Plots • Tick ‘Main Effects Plots’ • Setup > Select CRF as the response and choose all 3 variables (>>) • The Interactions plots can be produced similarly (just select ‘Interactions’ instead of ‘Main Effects Plots’) RMIT University
CRF average at ‘high’ methanol CRF average at ‘low’ methanol Note that Methanol has the steepest slope, indicating the strongest effect RMIT University
The plots show there is an interaction effect with methanol and citric acid at high methanol CA has a Positive effect but at low methanol it has a negative effect on CRS RMIT University
Conclusions • Methanol has the largest effect on CRF • The Methanol effect strongly depends on the Citric Acid level. Citric acid has a positive effect at high Methanol but a negative effect at low Methanol • All 3 variables do seem to affect the result. Citric acid has the smallest main effect but large interaction effect • Hence probably can’t ‘screen out’ any of these variables from further study RMIT University
Significance – Normal Probability Plots • Normal Probability Plots are used to test whether data is normally distributed. • In our case, we can use such a plot to test for significance of the effects/coefficients • If the effects are not significant we expect variations just to be due to random error and this can be tested with the plots. It is only a guide, however, as we have no real estimate of the experimental error • In Minitab the plot can be generated: • Stat > DOE >factorial > Analyse Factorial Design > Graphs and Select Effects Plots (Normal) RMIT University
Effects due to random errors should be on a straight line. This plot indicates Methanol and the Methanol/Citric Acid interaction are significant effects RMIT University
Problems • We have not replicated any experiments so no determination of error. We cannot tell if the coefficients (effects) overall are significant (although normal probability plots help). We can only compare them to see which is the most significant • We also cannot test for curvature – i.e are the effects of the variables linear. A non-linear effect can be when the response at the high and low levels is similar but at intermediate values is much higher or lower. pH effects are often non-linear RMIT University
Solution? • Add centre points!! • Centre points are experiments with all variables set at 0 (coded) i.e. mid values • Replication of the centre point allows determination of error RMIT University
Results added for the centre points RMIT University
Estimated Effects and Coefficients for CRF (coded units) Term Effect Coef SE Coef T P Constant 10.362 0.05000 207.25 0.003 Acetic Acid -0.3750 -0.1875 0.05000 -3.75 0.166 Methanol 1.9250 0.9625 0.05000 19.25 0.033 Citric Acid 0.1250 0.0625 0.05000 1.25 0.430 Acetic Acid*Methanol 0.1250 0.0625 0.05000 1.25 0.430 Acetic Acid*Citric Acid 0.0250 0.0125 0.05000 0.25 0.844 Methanol*Citric Acid 0.8250 0.4125 0.05000 8.25 0.077 P is the probability a coefficient is not significantly different from zero i.e no effect on CRF. A low probability (< 0.05 at the 5% level) indicates high significance. The methanol effect is the only significant one at the 5% level although the methanol-citric acid effect is just above the 5% level RMIT University
The centre point responses are all on the linear response line. Thus no curvature is indicated. RMIT University
The Normal Plot shows also that Methanol is the only significant effect but Methanol/CA interaction is probably also significant RMIT University
Next Phase? • Factorial designs give indication of significant effects and interactions • Designs such as the Central Composite Design (CCD) can be used to find the best (optimal) settings of the variables and plot Response Surfaces • CCD designs involve adding extra points (trials) to the Factorial Designs RMIT University
Axial points Cube (factorial) points CCD design for 3 factors. The 8 factorial points are corners of the cube. In this study the cube points correspond to the FFD . 6 axial points are added to form a CCD RMIT University
Example of a CCD design and the generated response surface This is an example of a response surface plot for the optimization of capillary electrophoresis separation of a mixture of ranitidine-related compounds. Ranitidine is a drug used in the treatment of gastric and duodenal ulcers. The two variables being optimized are pH and applied voltage. The response variable used in the optimization is the logarithm of the CEF function, a parameter devised to assess the quality of a chromatogram (this is an alternative response function to the CRF). This function takes into account peak separation and total elution time (J. Chrom. A 766 245-54. Optimization of the capillary electrophoresis separation of ranitidine and related compounds. V.M. Morris, C. Hargraeves, K. Overall, P.J.Marriott, J.G.Hughes) RMIT University
Extra Information • Notes on factorial and central composite designs can be found at the author’s website: • Chemometrics in Australia This site also has an Excel spreadsheet which sets out the calculations for the FFD design used in this tutorial RMIT University
Author: Dr Jeff Hughes School of Applied Science, RMIT UniversityMelbourne , Australia http://www.rmit.edu.au/staff/hughes j RMIT University