ADAPTIVE IMMUNE RESPONSE
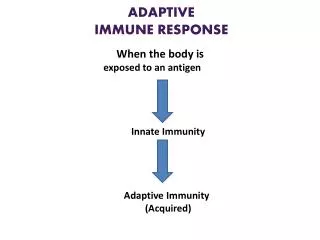
ADAPTIVE IMMUNE RESPONSE. Immune response to infections 1. Innate response Neutrophils, macrophages, NK cells at site of infection 2. Initiation of adaptive response iDCs phagocytose Ag and take to nodes DCs present Ag to T cells Get specific helper (CD4) T cells

ADAPTIVE IMMUNE RESPONSE
E N D
Presentation Transcript
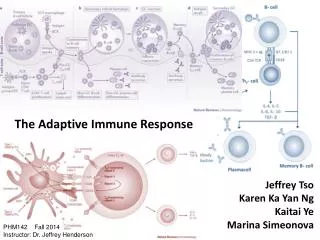
ADAPTIVE IMMUNE RESPONSE • Immune response to infections • 1. Innate response • Neutrophils, macrophages, NK cells at site of infection • 2. Initiation of adaptive response • iDCs phagocytose Ag and take to nodes • DCs present Ag to T cells • Get specific helper (CD4) T cells • “help” B cell activation = humoral immunity • Activate macrophages = CMI • Get specific cytotoxic (CD8) T cells • Kill virally infected cells = CMI
ADAPTIVE IMMUNE RESPONSET CELLS • Development: in thymus • Immunogenetic events produce multiple TCRs • Similar to B cells • TCR gene with multiple V, D, and J segments • Recognize 1015 epitopes • T cells with TCRs that react with self antigens deleted • Start with double negative cells: no TCR, CD4 or CD8 • Next get double positive cells: TCR, CD4, CD8 • Positive selection = weak recognition of MHCI or MHCII • Negative selection = if reaction is strong – apoptosis • ~2% of cells survive
ADAPTIVE IMMUNE RESPONSET CELLS • TCR complex • T cell receptor – similar to Fab of antibodies • Alpha and beta Ig-like chains with variable and constant domains • Hydrophobic transmembrane regions • CD3 proteins • Gamma + epsilon; delta + epsilon, each with one Ig-like domain • Transmembrane segment binds to TCR • Cytoplasmic domain of each has one ITAM • ITAM = immunoreceptor tyrosine-based activation notif; for signaling • Zeta chains = disulfide-liked homodimer, each protein has 3 ITAM • Transmembrane region with aspartate
ADAPTIVE IMMUNE RESPONSET CELLS • Co-receptors • CD4 protein binds to MHC-II on APC • CD4 T cells = helper T cells • MHC-II only on APCs • CD8 protein binds to MHC-I on target cells • CD8 T cells = cytotoxic T cells • MHC-I on all nucleated cells
ADAPTIVE IMMUNE RESPONSET CELLS • Accessory molecules = lymphocyte surface molecule distinct from Ag receptor complex; adhesion or signaling • CD45R, CD28, CD40L • Adhesion molecules = tighten interaction with APC • LFA (leukocyte function-associated antigen)-1, CD2 • Cytokine receptors: IL-1R, IL-2R, others
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • Antigen Presentation to T cells • MHC (major histocompatibility complex) 1 • all nucleated cells • MHC-encoded alpha (hvy) chain and beta 2 (lt) microglobulin • Polymorphic alpha 1 and alpha 2 domains for closed binding cleft • agretope • Conserved alpha 3 domains = binding site for CD8 • Beta 2 interacts noncovalently with alpha 3
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • Abbas Fig 4-4
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • MHC-II = dimer of alpha and beta subunits • APCs • Both chains MHC encoded • Alpha 1 and beta 1 domains variable and form open binding cleft • Most of variability on beta chain • Agretope • Alpha 2 and beta 2 folded into Ig domains • B2 Ig domain binds to CD4
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • Abbas Fig 4-6
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • Peptide presentation by MHC I molecules • MHC I molecule synthesized in endoplasmic reticulum (ER) • Cytoplasmic protein (including microbial) degraded by proteosome • Usually dealing with degraded (misfoldedetc) normal proteins • Degraded proteins marked by ubiquitin • To ER via TAP (transporter assoc w Ag processing) • Ag peptide binds to cleft in hvy chain of MHC-I • Peptide of 8 or 9 amino acids • MHC-I with peptide to cell membrane via golgi and exocytic vesicle • Uninfected cells usually coated with MHC-I with self proteins
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • Peptide presentation by MHC II molecules • Phagocytized proteins degraded in phagolysosomes by APCs • MHC-II molecule synthesized in ER • Invariant chain (Ii) placed in cleft • Ii provides stability and prevents MHC-II from binding with ER peptides • Lysosomal and exocytic vesicles merge • Invariant chain replaced with degraded peptide • cleft open at ends; longer peptides (30 aa) presented • Fused vesicle migrates to cell membrane
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • Antigen processing and presentation – Abbas Fig 5-8
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • Cross-presentation of Ag • iDCsphagocytise virally infected cells • Some peptides migrate to cytoplasm and processed to MHC I • Also, some cytoplasmic peptides from phagocytised cells processed to MHC II
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • Cross-presentation of Ag to CD8 T cells. Abbas Fig 5-7
ADAPTIVE IMMUNE RESPONSEantigen presenting cells • Antigen Presenting Cells (APCs) = Dendritic cells (DCs), B cells, and Macs • Dendritic cells = “only APC that can initiate Ag specific response” • Presumably from myeloid and lineage • Precursor DCs in blood; differentiate into immature DCs (iDC) • iDC recognized antigen with TLR and other receptors; activated • Pinocytosis/phagocytosis and cytokine production, now DC • DCs can no longer phagocytose; go to T-cell area of lymph nodes where they present Ag to T cells • Langerhan’s cells are skin DC
ADAPTIVE IMMUNE RESPONSEantigen presentation to T cells • T Lymphocytes • Recognize antigens (on dendritic cells) in peripheral lymphoid organs, resulting in expansion of Ag-specific lymphocyte pool • Differentiate into effector cells and memory cells • Effector lymphocytes become activated when they recognize Ag • Activation of T cells requires recognition of Ag displayed on APCs, costimulators, and cytokines produced by both APC and T cell
ADAPTIVE IMMUNE RESPONSE antigen presentation to T cells • Activation of CD4 T Cells • MHC II on APC binds with TCR • TCR recognizes foreign protein + MHC self proteins • Interaction strengthened by binding of CD4 to MHC II molecule • Signal transmitted via TCR/CD3 complex zeta proteins • Adhesion molecule interactions • LFA-3 on APC with CD2 on T cell • ICAM-1 on APC with LFA-1 on T cell • Costimulatory signals also required • B7 on APC to CD28 on T cell • cytokines • Partial activation without adequate costimulatory signal = Anergy or apoptosis; method to eliminate self-reacting cells or induce tolerance
ADAPTIVE IMMUNE RESPONSEHelper T cell Function • TH1 and TH2 CD4 (helper) T cells • TH0 cells mature into TH1 or TH 2 depending on • Nature and concentration of antigen • How antigen presented • Type of APC • Cytokines • TH1 = IgM, IgG, activated Macs • TH2 = humoral response; IgA, IgE
ADAPTIVE IMMUNE RESPONSECytotoxic T cells • Activation of CD8 T cells = cytotoxic T cells (CTLs) • Precursors in nodes bind TCR + CD8 to MHC-1 of APC + costim • TCR recognizes foreign protein in self MHC molecule • Specific clone expands by ~100,000 • Activated CTLs bind with target cell • Granulysin, granzymes and perforin released from granules = apoptosis • Also interaction of FasL on CTL with Fas on target = apoptosis • Apoptosis • Cell DNA and internal membranes fragment • Shrink to “apoptotic bodies” which are easily phagocytosed • “Clean” cell death as apposed to necrosis
HUMORAL IMMUNE RESPONSE • HIR = production of antibodies by plasma (B) cells • Some Definitions • Albumin, globulin = serum proteins • Gamma-globulin = immunoglobulin • Immunogen, antigen • Epitope (linear, conformational) • Hapten; carrier • Adjuvant (alum, Freunds, liposomes, CW components) • Immune tolerance (dose, co-stimulators, transplants ) • T (cell) independent antigen; T dependent Ag • Route of Ag administration • Immunoglobulin superfamily on next slide
HUMORAL IMMUNE RESPONSE • Immunoglobulin (antibody) structure
HUMORAL IMMUNE RESPONSE • Immunoglobulin structure • Heavy chain determines class/subclass IgM, G, A, D, E • Mu, gamma, alpha, delta, epsilon • Light chain (kappa, lambda) • Domains – constant and variable • One variable domain on each lt chain and one variable domain on each hvy chain • Three constant domains on hvy chain of IgG and IgA; four on IgM and IgE • One constant domain on lt chains • Variable domains on hvy/lt chains have 3 complementary determining regions (CDRs) – complements epitope structure
HUMORAL IMMUNE RESPONSE • Immunoglobulin structure • Fab portion; variable region • Fc portion; • C’ fixatin; • bind to Fc receptors on effector cells • Fc of IgG and IgA involved with transfer across placenta and mucosa • Hinge region • Papain = 2 Fab + Fc; Pepsin = F(ab)2 Fc • Membrane-spanning portion • Interchain disulfide bonds • Isotype = IgD, IgM, IgG1, IgG2, IgG3, IgG4, IgA1, IgA2, IgE • Allotype = variation of isotype in different individuals • Idiotype = protein sequences in variable region • Interacts with epitope
HUMORAL IMMUNE RESPONSE • Immunoglobulin Classes
HUMORAL IMMUNE RESPONSE • Membrane-associated antibodies • All isotypes may be membrane-associated on B cells • All membrane-associated AAs monomeric • Last CH region has 26 AAs with hydrophobic side chains • Membrane spanning region • Secreted antibodies • Found in blood and other extracellular fluids • IgG and IgE are monomeric • IgD negligible • IgM and IgA are multimeric • Have tail pieces on end of hvy chains which are bound to J chains
HUMORAL IMMUNE RESPONSE • Immunoglobulin Isotypes • IgD = <1% of serum Ig • membrane Ig on early B-cells • IgM = 5-10% of serum Ig • membrane Ig on early B-cells • IgD and IgM are only isotypes that can be expressed together on same cell • 5 day half-life in serum • Serum IgM pentameric; 10 Ag-binding sites; tail piece and j chain • Complement fixation; opsonin; bacteriolysis • First isotype after immunization; then wanes
HUMORAL IMMUNE RESPONSE • Immunoglobulin Isotypes • IgG = 85% of serum Ig • 4 subclasses; all monomeric • 23 day half-life • T cells required • Complement fixation; opsonin • Maternal IgG transported across placenta and neonatal intestinal epithelium • Second isotype after immunization; persists • Also anamnestic (booster) response
HUMORAL IMMUNE RESPONSE • Immunoglobulin Isotypes • IgA = 5-15% of serum Ig • 2 subclasses; both dimer, trimer? with tail piece and J chain • IgA transported across epithelial cells has additional secretory component • IgE = <1% of serum Ig • Bind to Fc receptors on Mast cells, basophils and eosinophils • Immediate hypersensitivity • Parasite infections
HUMORAL IMMUNE RESPONSE Neonatal Immunity: Neonates lack ability to mount effective immune response. Therefore, use maternal IgG and IgA IgG transported across placenta into fetal circulation IgG and IgA in breast milk ingested IgG and IgA neutralize organisms in gut IgG transported across gut epithelium into circulation Neonatal Fc receptor for IgG on placta and gut epithelium cells
HUMORAL IMMUNE RESPONSE • Immunogenetics • Chromosomes 2, 22, 14 have genes for kappa, lambda, hvy chains • Variable region genes upstream from constant region genes • Genetic recombination events to form variable region of hvy and lt chains: • V (variable) and J (joining) segments for light chains • ~ 300 V gene segments and ~5 J segments = ~ 1500 possible • V, D (diversity), and J segments for hvy chains • 300 - 1000 V genes, 12 D genes, and 6 J genes = <100,000 possible • Association of hvy and light chain = > 1,000,000 possible
HUMORAL IMMUNE RESPONSE • Immunogenetics (continued) • Class switching • Genes for constant region of hvy chain in set order • IgM, D, G3, G1, G2, G4, E, A1, A2 • Different constant region genes deleted in response to T cell cytokines
HUMORAL IMMUNE RESPONSE • Production of antibodies by B cell • B cell receptor = Ig molecule specific for non-self antigens • Cells with receptors for self antigens removed during development in bone marrow • Ig = IgD and IgM on naive B cells • B cell receptor complex • IgM or IgD + Ig alpha/Ig beta with immunoreceptor tyrosine-based activity motif (ITAM) • Signal transducing receptor initiates activation signal via Ig ITAM • Ultimately activates factors that induce transcription of genes
HUMORAL IMMUNE RESPONSE • Antibody response to T (cell) independent antigens • Surface Ig receptors recognize repeating sites on Ag • Long polysaccharides • Get cross-linking of many Ig receptors, activating signal cascade • Clonal expansion and plasma cell development of specific B cells • Disadvantages of T independent response • No T cell cytokines for class switching; IgM only • Ab has lower affinity for Ag (limited affinity maturation?) • No response in young (<2yr) children
HUMORAL IMMUNE RESPONSE • Antibody response to T (cell) dependent antigens • Ig receptors on B cell recognize Ag but cross-linking inadequate to activate cell • Therefore need second signal from T helper cell; thus • 1) Ag binds to Ig receptor on B cell as above • 2) Some bound Ag internalized, processed and presented in MHC-II surface molecule to T cell (which has receptor for this Ag); i.e., B cell functions as APC. T cell, so activated, secretes cytokines which are received by and stimulate the B cell • 3) The two signals stimulate the B cell to replicate, clonal expansion • A) Plasma cells which produce and secrete the Ig of the receptor • In nodes and bone marrow • B) Memory cells with the same receptor • Anamnestic (booster) response
HUMORAL IMMUNE RESPONSE • Activation of B cells: continued • A second signal may be provided by C’ receptor • Interaction of B cell with T cell via CD40 (B cell) – CD40L (T cell) • Class switching from influence of helper T cell cytokines • Initial = IgM • INFg = IgG • IL-4 = IgA, IgE • IL-5 = IgA (also induces eosinophil production) • Somatic mutation = increased affinity • Primary and Secondary responses • Primary immune response = strong IgM • Secondary immune response (second exposure to Ag) = faster, more IgG
HUMORAL IMMUNE RESPONSE • Antibodies inactivate microorganisms by • Agglutination • Neutralization • Antibody to toxins • Antibody to microbial surface molecules that bind to host cells • Opsonization • Natural killer cells have receptors for IgG • Eosinophils have receptors for IgG, IgA, and IgE • Complement fixation • Neutrophils, macrophages have receptors for C3b • Gram negative organisms susceptible to MAC