Background
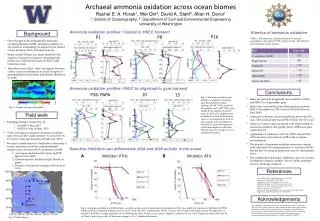
Ammonia oxidation profiles: Coastal to HNLC transect . Kinetics of ammonia oxidation. Background. P16. P1. P8. Table 1. The ammonia oxidation kinetics of natural populations. We varied [ 15 NH 4 + ] from 10 nM – 250 nM (n=3 per treatment, 3 time points).

Background
E N D
Presentation Transcript
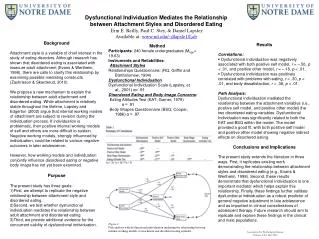
Ammonia oxidation profiles: Coastal to HNLC transect Kinetics of ammonia oxidation Background P16 P1 P8 Table 1. The ammonia oxidation kinetics of natural populations. We varied [15NH4+] from 10 nM – 250 nM(n=3 per treatment, 3 time points) • Once thought to be dominated by ammonia-oxidizing Bacteria (AOB), ammonia oxidation in the oceans is increasingly recognized to be carried out by Archaea (AOA, Thaumarchaeota) • Many oceanic biomes are under-sampled with respect to ammonia oxidation, including high nitrate, low chlorophyll regions (HNLC) and transition zones • Transition zones (Fig.1, stars) are regions between classic ocean environments and host hotspots of phytoplankton biodiversity and blooms (Ribalet et al. 2010) Archaeal ammonia oxidation across ocean biomesRachel E. A. Horak1, Wei Qin2, David A. Stahl2, Allan H. Devol11: School of Oceanography, 2: Department of Civil and Environmental EngineeringUniversity of Washington Ammonia oxidation profiles: HNLC to oligotrophic gyre transect Conclusions P26: PAPA 10 13 SeaWiFS Project, NASA/Goddard Space Flight Center Fig. 3. Ammonia oxidation rate profiles, inorganic N nutrients, and chlorophyll for select stations. 50 nM15NH4+ added to all samples, and 5 mM14NO2- was added to all samples in mixed layer. In 2012, rate samples were incubated on deck in the dark at near in situ temperature. In 2013, rate samples were incubated in the dark on a free-floating array at the same depth of collection. All rate data are average (n=3) ± 1 standard deviation. • Rates decreased in magnitude from coastal to HNLC, and HNLC to oligotrophic gyre. • Rates were consistently lower than previous studies: HNLC (Grundle et al. 2013) and at ALOHA (Dore & Karl 1996) • Ammonia oxidation was not significant above the NO2max. The maximal rate was 20-80 m below the NO2max • AOA amoA gene copies increased with depth similar to ammonia oxidation rate profile, but b-AOB amoA gene copies did not. • Application of inhibitors (ATU & PTIO) showed that AOA are more active than b-AOB in these marine environments • The kinetics of ammonia oxidation were most similar with cultivated AOA representative N. maritimus SCM1, but the Km for natural populations may be substantially lower • The combination of kinetics, inhibitors, and amoA copies is helpful to identify whether AOA or AOB contribute more to ammonia oxidation Fig. 1. Annual average chlorophyll Field work • Sampling during 2 cruises (Fig. 2): • GeoMICS: May 2012 • KM1314: Aug. & Sept. 2013 • 15NH4+ technique to measure ammonia oxidation rates. We used the “azide method” to convert N+N to N2O for mass spectrometry (Horak et al. 2013) • We used a combination of 3 methods to determine if oceanic ammonia oxidation is predominately carried out by Archaea (AOA) or Bacteria (AOB): • amoAgene and transcript copies (q-PCR; Mincer et al. 2007) • Domain-specific inhibitors (Qin, Horak, in prep) • Kinetics of ammonia oxidation (Horak et al. 2013) Selective inhibitors can differentiate AOA and AOB activity in the ocean A B Inhibitor: ATU Inhibitor: PTIO P26 PAPA 9 P16 P8 P1 10 References 11 12 Dore & Karl 1996 LimnolOceanog 41: 1619-1628 Horak et al. 2013 ISME J 7:2023-2033 Grundleet al. 2013 Mar Chem 155: 113-123 Martens-Habbena et al. 2009. Nature 461:976-981 Mincer, et al. 2007. Environ Microbiol 9:1162-1175 Qin et al. in revision, PNAS USA Qin, Horak, Martens-Habbena, et al. in prep. Environ Microbiol Ribalet et al. 2010 PNAS USA 107: 16571-16576 13 14 15 16 ALOHA Acknowledgements We would like to thank the ship’s crew and technicians of R/V T. Thompson and Kilo Moana, Chief Scientists Ginger Armbrust and Anitra Ingalls, and the Dimensions of Biodiversity science party for assistance. Special thanks to Laura Carlson, Katherine Heal, Davey French, Alex Nelson for outstanding lab and field assistance, and Andrew Schauer for mass spectrometry. Funding provided by NSF grants OCE-1046017 and OCE-1205232. Fig. 4. Ammonia oxidation in AOB cultures, an AOA isolate, and in natural marine populations (NO2 max depth) in response to inhibitor (A) PTIO, 2-phenyl-4,4,5,5,-tetramethylimidazoline-1-oxyl-3-oxide, (B) ATU, allylthiourea. HCA1 is a new AOA isolate from Puget Sound, WA (Qin et al. in revision). b-AOB is average inhibition of 7 b-AOB strains (Qin, Horak, et al. in prep). Depth for station P16 was 85 m. Depth for station P26 was 65 m. Note: x-axis is log scale. All data are average (n=3) ± 1 standard deviation. Fig. 2. Field stations