Chapter 11
Chapter 11. Membrane Transport of Small Molecules and the Electrical Properties of Membranes. I. Principles of membrane transport - protein-free lipid bilayers are highly impermeable to ions - two main classes of membrane transport proteins

Chapter 11
E N D
Presentation Transcript
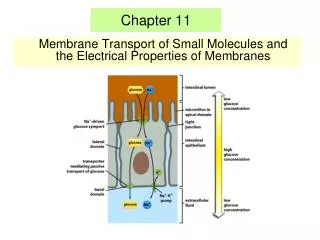
Chapter 11 • Membrane Transport of Small Molecules and the Electrical Properties of Membranes
I. Principles of membrane transport - protein-free lipid bilayers are highly impermeable to ions - two main classes of membrane transport proteins - transporters ( also called carriers or permeases) and channels - active transport – mediated by transporters coupled to an energy source - passive transport – mediated by all channels and many transporters II. Transporters and passive transport - glucose transporters III. Transporters and active transport - coupled transporters; ATP-driven pumps; light-driven pumps - uniporters; symporters; antiporters - Na+-glucose cotransporter – coupled transport; symporter - lactose permease - coupled transport; symporter - three classes of ATP-driven pumps - Ca2+ pump - Na+-K+ pump - ABC transporters IV. Ion channels - ion channels are selective and gated - membrane potential - K+- channel - aquaporins - neuron - voltage-gated Na+-channels - transmitter-gated ion channels - neuromuscular transmission
Ion concentration differences across the lipid bilayer are useful for: • Driving various transport processes • Conveying electrical signals in electrically excitable cells • Making most of the cell’s ATP • Cell signaling
The rate at which a molecule diffuses across a synthetic lipid bilayer depend on its size and solubility The smaller the molecule and the less polar it is, the more rapidly it diffuses across the bilayer
Permeability coefficients (in cm/sec) through synthetic lipid bilayers Product of the concentration difference (in mol/cm3) and permeability coefficient (in cm/sec) gives the flow of solute in moles per second per square centimeter of membrane
Two main classes of membrane transport proteins: Transporters and Channels All these proteins are multi-pass transmembrane proteins 1. Transporters bind to a specific solute and undergo a series of conformational changes. 2. Channel proteins interact with the solute much more weakly; form aqueous pores; transport at a much faster rate.
An electrochemical gradient combines the membrane potential and the concentration gradient
A conformational change in a transporter could mediate the passive transport of a solute
The kinetics of simple diffusion and transporter-mediated diffusion
Transporters and active transport Cells carry out active transport in three main ways: (1) Coupled transporters couple the uphill transport of one solute across the membrane to the downhill transport of another. (2) ATP-driven pumps couple uphill transport to the hydrolysis of ATP. (3) Light-driven pumps, found mainly in bacteria and archaea, couple uphill transport to an input of energy from light.
Cells drive active transport in three ways The actively transported molecule is shown in yellow and the energy source is shown in red - ion-driven transporters mediate secondary active transport - ATP-driven transporters mediate primary active transport
Three types of transporter-mediated transport Uniports, symports, and antiports are used for both passive and active transport
Active transport Coupled transport – the transport of one solute strictly depends on the transport of a second. It involves either the simultaneous transfer of a second solute in the same direction (symporters) or the transfer of a second solute in the opposite direction (antiporters) The tight coupling between the transport of the two solutes allows these carriers to harvest the energy stored in the electrochemical gradient of one solute, typically an ion, to transport the other
The glucose-Na+ symport protein uses the electrochemical Na+ gradient to drive the import of glucose
The active transport of many sugars and amino acids into bacterial cells is driven by the electrochemical H+ gradient across the plasma membrane E. coli lactose permease
During the transport cycle, some of the helices undergo sliding motions causing them to tilt. An alternative opening and closing of the crevice between the helices exposes the binding sites for lactose and H+, first on one side of the membrane and then the other
An asymmetric distribution of carrier proteins underlies the transcellular transport of solutes
The Ca2+ pump Structures of the unphosphorylated, Ca2+-bound state (left) and the phosphorylated, Ca2+-free state (right) have been determined by x-ray crystallography
Model showing how ATP binding and hydrolysis cause drastic conformational changes, bringing the nucleotide-binding and phosphorylation domains into close proximity. This change is thought to cause a 90° rotation of the activator domain, which leads to a rearrangement of the transmembrane helices. The rearrangement of the helices disrupts the Ca2+-binding cavity and releases the Ca2+ into the lumen of the SR.
The Na+-K+ pump transports ions in a cyclic manner The binding of cytosolic Na+ (1) and the subsequent phosphorylation by ATP of the cytosolic face of the pump (2) induce the protein to undergo a conformational change that transfers the Na+ across the membrane and releases it on the outside (3). The linkage of the phosphate to an aspartic acid in the protein drives the conformational change. The binding of K+ on the extracellular surface (4) and the subsequent dephosphorylation (5) return the protein to its original conformation, which transfers the K+ across the membrane and releases it into the cytosol (6). The changes in conformation are analogous to the A ⇌ B transitions of transporters, except that here the Na+-dependent phosphorylation and the K+-dependent dephosphorylation of the protein cause the conformational transitions to occur in an orderly manner, enabling the protein to do useful work. For simplicity, only one Na+- and one K+-binding site are shown. In the real pump in mammalian cells, there are three Na+- and two K+-binding sites. The net result of one cycle of the pump is therefore to transport three Na+ out of the cell and two K+ in. Ouabain inhibits the pump by preventing K+ binding.
The Na+ - K+ pump is required to maintain osmotic balance and stabilize cell volume
Auxiliary transport system associated with transport ATPases in bacteria with double membranes The transport ATPases belong to the ABC transporter supefamily
Examples of a few ABC proteins Nature Structural & Molecular Biology 11, 918 - 926 (2004)
A typical ABC transporter consists of four domains – two highly hydrophobic domains and two ATP-binding catalytic domains ATP binding leads to dimerization of the two ATP-binding domains and ATP hydrolysis leads to their dissociation.
The ATP switch model for the transport cycle of an ABC transporter The schematic is for a drug exporter. The drug-binding site (red) is high-affinity and faces the inner leaflet of the membrane. Step I: the transport cycle is initiated by binding of substrate to its high-affinity site on the TMDs from the inner leaflet of the membrane. The affinity of the NBDs for ATP is increased, effectively lowering the activation energy for closed dimer formation. Two molecules of ATP bind, cooperatively, to generate the closed dimer. Step II: the closed NBD dimer induces a conformational change in the TMDs such that the drug-binding site is exposed extracellularly and its affinity is reduced, releasing the bound drug. Step III: ATP is hydrolyzed to form a transition-state intermediate. Hydrolysis of the two ATP molecules is normally sequential, although for some ABC transporters only one ATP may be hydrolyzed. Step IV: sequential release of Pi, and then ADP, restores the transporter to its basal configuration. Nature Structural & Molecular Biology 11, 918 - 926 (2004)
Ion channels Can transport up to 100 million ions per second, a rate 105 times greater than that mediated by a carrier protein
Among their many functions, ion channels control the pace of the heart, regulate the secretion of hormones into the bloodstream, and generate the electrical impulses underlying information transfer in the nervous system.
Ion channels are ion-selective (ion selectivity) and fluctuate between open and closed states (gated)
Ion channels, like enzymes, have their specific substrates: potassium, sodium, calcium, and chloride channels permit only their namesake ions to diffuse through their pores. The ability of channels to discriminate among ions is called ion selectivity.