10.2
10.2. 14.7. GTI1. B3 C-limited (CLB). B3 Replete (RB). B3 N-limited (NLB).

10.2
E N D
Presentation Transcript
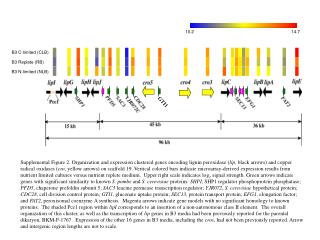
10.2 14.7 GTI1 B3 C-limited (CLB) B3 Replete (RB) B3 N-limited (NLB) Supplemental Figure 2. Organization and expression clustered genes encoding lignin peroxidase (lip, black arrows) and copper radical oxidases (cro, yellow arrows) on scaffold 19. Vertical colored bars indicate microarray-derived expression results from nutrient limited cultures versus nutrient replete medium. Upper right scale indicates log2 signal strength. Green arrows indicate genes with significant similarity to known S. pombe and S. cerevisiae proteins: SHPI, SHP1 regulator phosphoprotein phosphatase; PFD5, chaperone prefoldin subunit 5; SAC3 leucine permease transcription regulator; YJR072, S. cerevisiae hypothetical protein; CDC28, cell division control protein; GTI1, gluconate uptake protein; SEC13, protein transport protein; EFG1, elongation factor; and FAT2, peroxisomal coenzyme A synthesis. Magenta arrows indicate gene models with no significant homology to known proteins. The shaded Pce1 region within lipI corresponds to an insertion of a non-autonomous class II element. The overall organization of this cluster, as well as the transcription of lip genes in B3 media had been previously reported for the parental dikaryon, BKM-F-1767. Expression of the other 16 genes in B3 media, including the cros, had not been previously reported. Arrow and intergenic region lengths are not to scale.