Separation: Agarose gel electrophoresis-I
210 likes | 510 Views
Section G-II Gene Manipulation G3 Electrophoresis G4-1 Ligation G4-2 Transformation and G4-3 Analysis of recombinants. -. +. DNA sample. buffer solution. Migration of DNA. Agarose gel. Separation: Agarose gel electrophoresis-I.

Separation: Agarose gel electrophoresis-I
E N D
Presentation Transcript
Section G-IIGene Manipulation G3 Electrophoresis G4-1 Ligation G4-2 Transformation and G4-3 Analysis of recombinants Yang Xu, College of Life Sciences
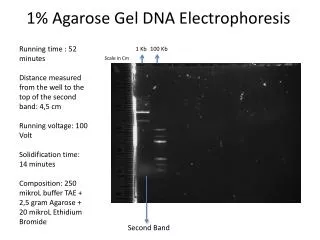
- + DNA sample buffer solution Migration of DNA Agarose gel Separation: Agarose gel electrophoresis-I Definition: Agarose is a polysaccharide derived from seaweed, which forms a solid gel when dissolved in aqueous solution at concentrations between 0.5 and 2% (w/v). Principle: • Movement: When an electric field is applied to an agarose gel in the presence of a buffer solution, DNA fragments move through the gel towards the + electrode, because DNA negatively charged. • Movement rate: The movement rate is dependent on fragment sizes and shapes of the DNA and the electrophoresis may be used to separate mixtures of DNA fragments by the rate. Yang Xu, College of Life Sciences
Separation: Agarose gel electrophoresis-II General procedures: • The DNA samples are placed in wells in the gel surface. • the power supply is switched on and the DNA is allowed to migrate through the gel in separate lanes or tracks. • The added dye (EB) also migrates, and is used to find the progress. • The DNA is stained by the EB in the gel, the DNA shows up as an orange band on illumination by UV light. Yang Xu, College of Life Sciences
- + Separation: Agarose gel electrophoresis-III Result reading: see p103 Fig 4 • Tracks M: A set of linear marker DNA fragments of known sizes; • Track U: Undigested plasmid DNA consists of two bands. The lower band consists of negatively supercoiled plasmid DNA. The upper band is the nicked DNA, which has lower mobility; • Track B: digestion with BamHI, containing a single fragment; • Track E: digestion with EcoRI, containing five fragments, the third largest fragment contains the gene X, which could be purified from the gel and ready for ligation into a new vector. Yang Xu, College of Life Sciences
Purification of fragments • Purposes: Agarose gels may also be used preparatively to isolate specific fragments for use in subsequent ligation and other cloning experiments. • Purification method: Fragments are cut from the gel, and treated by one of a number of procedures to purify the DNA away from the contaminating agarose. • Example: If we assume that the EcoRI fragment, which contains the gene X, is the target DNA for a subcloning experiment, then the third largest fragment (in track E of Fig. 4a) could be purified from the gel, ready for ligation into a new vector (see Topic G4). Yang Xu, College of Life Sciences
G4-1 Ligation (DNA Recombination) • DNA ligation • Recombinant DNA molecules • Alkaline phosphatase Yang Xu, College of Life Sciences
DNA ligation • Purpose: To insert a target DNA fragment into a vector. • Function of DNA ligases: they will repair (ligate) a break in one strand of a dsDNA molecule. • Energy sources: • DNA ligase from E. coli uses NAD+as energy source; • T4 DNA ligase uses ATP (more commonly used). • Different ends: • Cohesive ends: Ligases are efficient at sealing the broken phosphodiester bonds for cohesive ends, • Blunt ends: just T4 ligase can even ligate one blunt end to another, but with rather lower efficiency. Yang Xu, College of Life Sciences
E S E E S E E E Ori EcoRI E E A B Recombinant DNA molecules-I • Making of the recombinant DNA: • Target DNA: may be a single fragment isolated from agarose gel, or a mixture of many fragments from genomic DNA. • Vector DNA: the new vector DNA can be cut with the same enzyme. If the target DNA has been prepared by digestion with same enzyme (EcoRI). • Recombinant molecules: The products are circular molecules with the target fragment inserted at the EcoRI site of the vector molecule with either orientation. Yang Xu, College of Life Sciences
Recombinant DNA molecules-II • A problem: The recreation of the original vector plasmid, by circularization of the linear vector alone, is a competing side reaction which can make problems on the recombinant identification. • Solution one--Paired distinct restriction: One solution is to prepare both the target DNA and the vector DNA using a pair of distinct restriction enzymes, such that they have non-compatible cohesive ends at either end. The likelihood of ligating the vector into a circle is then much reduced. Yang Xu, College of Life Sciences
E S E G -OH HO -AATTC CTTAA -OH HO -G E E G -OH P HO-AATTC P CTTAA -OH HO -G Alkaline phosphatase Solution two: Linear vector: Alkaline phosph-atase removes phosphate groups from the 5'-ends of DNA molecules. The linear vector will hence be unable to ligate into a circle, since no phosphates are available. Target DNA: A ligation with a target DNA insert can still proceed, since one phosphate is present to ligate one strand at each cut site. The nicks in the other strands will be repaired when transformated in E.coli. Yang Xu, College of Life Sciences
G4 -2 Transformation • Transformation • Selection of plates • Transformation efficiency • Growth and storage of transformants Yang Xu, College of Life Sciences
Transformation-I • Definition: It is the process of take-up of foreign DNA (normally plasmids) by bacteria. Plasmids are cloned by transferring into E. coli with defined genetic properties. • Purpose: The recombinant and other plasmid formed by ligation must be isolated from one another and replicated by a host. • Hosts: The most common hosts are strains of special E. coli, which must not express a restriction-modification system. The host cell can accept only one kind of foreign plasmid. • Competent cells: It was discovered that E. coli cells treated with solutions containing Ca2+ ions were rendered susceptible to take up exogenous DNA. Cells pretreated with Ca2+, in order to render them able to take up DNA, are known as competent cells. Yang Xu, College of Life Sciences
plasmid Transformation-II Transformation process: Taking up DNA: A solution of a plasmid molecule is combined with a suspension of competent cells for a period (30-40min), to allow the DNA to be taken up. Recovering: The mixture is then heat-shocked at 42 ºC for 1-2 min. This step induces enzymes involved in the repair of DNA, which allow the cells to recover from unusual conditions of the transformation process, and increases the efficiency. Transformation: The cells are then incubated in a growth medium and finally spread on an agar plate and incubated until single colonies of bacteria grow. Taking up DNA Recovering Transformation Yang Xu, College of Life Sciences
Selection of plates A problem:If all the competent cells present in a transformation reaction were allowed to grow on an agar plate, then many thousands or millions of colonies would result. Hence, a method for the selection of clones containing a plasmid is required. Principle: This is almost always provided by the presence of an antibiotic resistance gene on the plasmid vector, for example the -lactamase gene (ampr) conferring resistance to ampicillin. If the transformed cells are grown on plates containing ampidllin, only those cells which are expressing -lactamase (due to the presence of a transformed plasmid) will survive and grow. Notice: If a ligation mixture (but not a single plasmid molecules) had been used for the transformation, we would not know at this stage which clones contain recombinant plasmids with a target fragment incorporated (p106 Fig. 1). Yang Xu, College of Life Sciences
Transformation efficiency Definition: The quality of competent cells may be measured by determining the transformation efficiency, defined as the number of colonies formed per g pure plasmids. Range and usage: Transformation efficiencies can range: • 103 colonies/g plasmids: can be used for crude transformation protocols, which would only be appropriate for transferring an intact plasmid to a new host strain (subcloning); • 105 colonies/g plasmids: would be enough for a simple cloning experiment of the kind outlined in this book. • >108 colonies/g plasmids: can be used for generation of libraries; Yang Xu, College of Life Sciences
Growth and storage of transformants Growth: • Method: Single colonies from a transformation plate are trans-ferred to broth and grown overnight until the stationary phase. • Notice: The broth must be included with the antibiotic, and used to select the transformants on the original plate, to maintain the selection for the presence of the plasmid. Storage: • Method: It is normal practice to prepare a stock of each culture at this stage, by freezing a portion of the culture in the presence of glycerol. • Reason: The effect of glycerol stock is to protect the cells from ice crystal formation. The stock will enable the same strain/plasmid to be grown and prepared again if and when it is required. Yang Xu, College of Life Sciences
G4 -3 Analysis of Recombination • Screening transformants • Gel analysis • Fragment orientation Yang Xu, College of Life Sciences
Screening transformants Transformant definition: It is the cloned bacteria which contain a recombinant plasmid, with inserted target fragment. Purpose: Once a set of transformant clones have been produced in the experiment, the first requirement is to know which clones contain a recombinant plasmid, with inserted target fragment. Methods: 1. In the case of a simple subcloning experiment, the normal method of screening is to prepare the plasmid DNA from a number of clones and analyze it by agarose gel electrophoresis. 2. Commercial plasmids have been designed to facilitate this process, and are described in Topic H1. 3. In many cases, such as the screening of a DNA library, it will then be necessary to identify the interest clone among thousands or even hundreds of thousands of others (Topic I3). Yang Xu, College of Life Sciences
- + Gel analysis Application: • Simple gel analysis: Recombinant plasmids can usually be simply distinguished from recreated vectors by the relative sizes of the plasmids, and • Gel analysis of restriction digests: Further analysis need to follow the restriction digests. Page 109 Fig. 4 shows a gel map representing the analysis of the plasmids in Fig. 1. • Tracks corresponding to the vector plasmid and to recombinants are indicated. • Tracks U: The larger size of the recombinant plasmid is seen by comparing the undigested plasmid samples, containing super-coiled and nicked bands (see Topic G3); • Track E: The excision from the recombinant is seen in the EcoRI digest. Yang Xu, College of Life Sciences
A small large H H S S B Fragment orientation Orientation: The orientation of the fragment can be determined using a restriction digest with: • An enzyme (SalI) which is known to cut asymmetrically within the insert sequence, together with • Another enzyme (HindIII) which cuts at some specified site in the vector. Illustration: This is illustrated using a double digest with: • The patterns expected from two orientations, A and B, of inserted fragment are illustrated in Fig. 4 (tracks H/S). Yang Xu, College of Life Sciences
That’s all for Section G Yang Xu, College of Life Sciences