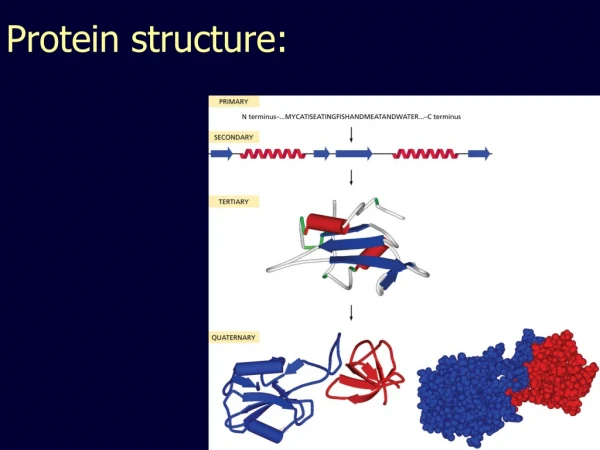
Protein Structure Similarity
720 likes | 1.18k Views
Protein Structure Similarity. Computation of Best Matches. Two “simultaneous” subproblems Find maximal correspondence set C Find alignment transform T Chicken-and-egg issue: Each subproblem is relatively simple: If we knew C, we could compute T

Protein Structure Similarity
E N D
Presentation Transcript
Computation of Best Matches Two “simultaneous” subproblems • Find maximal correspondence set C • Find alignment transform T Chicken-and-egg issue: • Each subproblem is relatively simple: • If we knew C, we could compute T • If we knew T, we could get C by proximity • But the combination is hard !!!
Only requires computing 6 parameters Computation of Best Matches Two “simultaneous” subproblems • Find maximal correspondence set C • Find alignment transform T Chicken-and-egg issue: • Each subproblem is relatively simple: • If we knew C, we could compute T • If we knew T, we could get C by proximity • But the combination is hard !!!
Find Alignment Transform • Two sets of points A= {a1,…,an} and B = {b1,…,bn} • Correspondence pairs (ai, bi) • Find T = arg minT RMSD(A,T(B)) • O(n) closed-form solution[Arun, Huang, and Blostein, 87] [Horn, 87] [Horn, Hilden, and Negahdaripour, 88]
O(n) SVD-Based Algorithm • T combines translation t and rotation R, such that T(bi) = t + R(bi) • b = (Σi=1,...,nbi)/n [mean of the bi’s] • Place the origin of coordinate system at b • minT RMSD(A,T(B)) simplifies to (up to some constants): • t and R can be computed separately • t = a[mean of the ai’s] [Arun, Huang, and Blostein, 87]
O(n) SVD-Based Algorithm • A3n = [a1-a, ..., an-a]B3n = [b1-b, ..., bn-b] • Compute SVD decomposition of 3×3 correlation matrix BAT: BAT = UDVTwhere D is a diagonal matrices with decreasing non-negative entries (singular values) along the diagonal • If det(U)det(V) = 1 then S = I, else S = diag(1,1,-1) • R = USVT [Arun, Huang, and Blostein, 87]
O(n) SVD-Based Algorithm • A3n = [a1-a, ..., an-a]B3n = [b1-b, ..., bn-b] • Compute SVD decomposition of 3×3 correlation matrix BAT: BAT = UDVTwhere D is a diagonal matrices with decreasing non-negative entries (singular values) along the diagonal • If det(U)det(V) = 1 then S = I, else S = diag(1,1,-1) • R = USVT [Arun, Huang, and Blostein, 87]
[Arun, Huang, and Blostein, 87] rotation matrix • [Horn, 87] quaternion
Guess small correspondence set Compute T Update correspondence set(correspondence from proximity) Apply T Trial-and-Error Approach to Protein Structure Comparison
Trial-and-Error Approach to Protein Structure Comparison • Set CS to a seedcorrespondence set (small set sufficient to generate an alignment transform) • Compute the alignment transform T for CS and apply T to the second protein B • Update CS to include all pairs of features that are close apart • If CS has changed, then return to Step 2else return (CS,T)
Trial-and-Error Approach to Protein Structure Comparison - result= nil - Iterate N times: • Set CS to a seedcorrespondence set (small set sufficient to generate an alignment transform) • Compute the alignment transform T for CS and apply T to the second protein B • Update CS to include all pairs of features that are close apart • If CS has changed, then return to Step 2else result result {(CS,T)} - Return result
Seed Generation from Fragment • From distance matrices E.g., DALI [Holm and Sander, 1996]
45 40 85 1 Using Distance Matrices (DALI) • Distances are invariant to rigid-body transformations • DALI [Holm and Sander, 1996] looks for similar hexapeptides by searching for similar 7x7 Ca-Ca distance matrices
Seed Generation from Fragment • From distance matrices E.g., DALI [Holm and Sander, 1996] • From secondary structure elements (SSE’s) E.g., LOCK [Singh and Brutlag, 1996] • From voting scheme (using geometric hashing) E.g., 3dSEARCH [Singh and Brutlag, 2000]
LOCK A.P. Singh and D.L. Brutlag. Hierarchical Protein Structure Superposition Using Both Secondary and Atomic Representations. Proc. ISMB, pp. 284-293, 1997. LOCK2:J. Shapiro and D.L. Brutlag. FoldMiner: Structural Motif Discovery Using an Improved Superposition Algorithm. Protein Science, 13:278-294, 2004. http://motif.stanford.edu/lock2/
LOCK • Two levels of features: SSEs and Ca atoms • Stage 1 (SSE alignment):Initial alignment is computed using SSEs represented as vectors • Stage 2 (atom alignment):Alignment is refined using Ca atoms represented as points
Rationale for LOCK • Using types of features is an effective way to reduce combinatorial explosion and computation • SSEs, which are responsible for most of the stability and functionality of the proteins, are more meaningful and better conserved than types of atoms and amino-acids • If 2 structures are similar, some of their SSEs should form similar substructures • Drawback: It narrows down the set of possible applications, e.g., can’t find small motifs at atomic level
Vector-Based Representation b-strands loops a-helices One vector per SSE (helix, strand, loop)
Vector-Based Representation • DSSP [Kabsch and Sander, 1983] classifies residues into helices/strands • For a-helix starting at residue i:Xorigin= (0.74Xi + Xi+1 + Xi+2 + 0.74Xi+3)/3.48where Xi is the position of the Ca atom of residue i (angle between two consecutive residues is 100dg factor 0.74) • Similar computation for Xend and for b-strand
Assume that i and p have been aligned. What is the score of the alignment of k and r? 2Mi S(di) = - Mi 1+(di/di0)2 Scoring Similarity • Position-independent differences: • |angle(i,k)-angle(p,r)| • |angle(i,j)-angle(p,q)| • |angle(j,k)-angle(q,r)| • |distance(i,k)-distance(p,r)| • |length(k)-length(r)| • Position-dependent differences: • angle(k,r) • distance(k,r) • Scores are additive Maximal score Score = S S(di) Value of di forwhich score is 0
E.g., using start, middle, and end points of vectors Stage 1: SSE Alignment • For every pair of SSE vectors of protein A, find all pairs of vectors in B that align well using orientation-independent scores seed correspondence sets • For each correspondence set: • Find alignment transform and apply it to B • Find correspondence set with maximal score (record transform T and correspondence set CS that yields maximal score)
(i,p), (j,q) (k,t) (l,r) (m,r) (k,r) (k,s) (m,s) (m,t) (l,s) (l,t) (m,s) (m,t) (l,t) (m,t) (m,t) Stage 1: SSE Alignment • A = (i, j, k, l, m) • B = (p, q, r, s, t) • Seed correspondence {(i,p),(j,q)} • Simultaneous gaps in both structures are not allowed (not in SCOP2) • Terminate a path when score of new correspondence is negative • Re-compute new transform with each new correspondence (?)
Stage 2: Atom (Core) Alignment • Construct correspondence pairs of atoms : • Atom i of A corresponds to atom j of T(B) iff i is the closest atom in A to j and j is the closest atom in T(B) to i • The distance between i and T(j) is <e (3Å) • Prune correspondence set to largest subset of correspondence pairs that follow backbone alignment constraint • Re-compute T to be the transform that minimizes the RMSD of the atoms in the correspondence set • Iterate 1-2-3 until RSMD converges
Experimental Results • 685 protein structures from PDB such that each pair has less than 25% sequence identity • 3 families of folds (based on SCOP classification): - myoglobins (11 structures) – ~20% amino acid identity- TIM barrels (50 structures)- immunoglobulins (38 structures) • Goal: Given one query protein in each family, find the other members of the family (3×685 = 2055 alignments) • Method: For each query, sort the 685 structures by score (computed by LOCK). Select the top k proteins. Count members of family (true positives) and non-members (false positives)
Myoglobins (11) TIM-barrels (50) Immunoglobulins (38)
Alignment of 50 TIM barrels a-helices in red b-strands in yellow
Alignments of 31 Immunoglobulins Only b-strands are shown
Running Time • ~ 1ms per seed correspondence • ~ 1h to search 10,000 protein structures • ~ 100s of days to compare all pairs of proteins in PDB • Geometric hashing to speedup stage 1
Seed Generation from Fragment • From distance matrices E.g., DALI [Holm and Sander, 1996] • From secondary structure elements (SSE’s) E.g., LOCK [Singh and Brutlag, 1996] • From voting scheme (using geometric hashing) E.g., 3dSEARCH [Singh and Brutlag, 2000]
Voting Scheme with Hash Table • Many-to-many comparison requires a better organization of computation to avoid repeating the same computation again and again • Pre-computation: Index proteins in hash table • Query phase: Voting scheme using hash table • Several variants on this theme 3d-Lookup [Holm and Sander, 1995] 3dSEARCH [Singh 2002]
Voting Scheme with Hash Table • Many-to-many comparison requires a better organization of computation to avoid repeting the same computation again and again • Pre-computation: Index proteins in hash table • Query phase: Voting scheme using hash table • Several variants on this theme 3d-Lookup [Holm and Sander, 1995]3dSEARCH[Singh 2002]
Indexing Target Structures in Hash Table (3dSEARCH [Singh 2002]) • Hash table: 3-D regular grid of cubic bins (~2Å) • For each target structure For each pair of vectors (i,j) • Compute a coordinate system • Place an entry for each other vectork into the bin containing the coordinates of the midpoint of the vector (or average of coordinates of origin, middle, and end points). Store ID of coordinate system + k’s orientation and type (a or b) in the entry.
v u v u Grid is same for all coordinate systems
v v u u Grid is same for all coordinate systems
Indexing Target Structures in Hash Table (3dSEARCH [Singh 2002]) • Hash table: 3-D regular grid of cubic bins (~2Å) • For each target structure For each pair of vectors (i,j) • Compute a coordinate system • Place an entry for each other vectork into the bin containing the coordinates of the midpoint of the vector (or average of coordinates of origin, middle, and end points). Store ID of coordinate system + k’s orientation and type (a or b) in the entry. • Grid is sparsely occupied hash table • A structure with n SSEs contributes n(n-1)(n-2) entries. Each vector is represented (n-1)(n-2) times • 10,000 structures with 10 SSEs each yield ~7M entries
Voting Using Hash Table Given a query structure • For each pair of vectors (i,j) • Compute a coordinate system • For each other vector k • Retrieve the bin accessed by this vector and the neighboring bins • For every entry (vector) in those bins that has the same orientation and type as k, add a vote for the coordinate system stored in the entry • Sort target structures based on max number of votes received by any of its coordinate systems • Small number of target structures. Use LOCK for better alignment • Hours of pure LOCK are reduced to seconds
Advantages of Voting System • Very efficient in practice for many-to-many comparisons • Can establish correspondence between partial, disconnected substructures • Parallel implementation is straightforward • Independent of the order in which vectors are considered • Drawback (?): May establish correspondences that do not satisfy the backbone sequence constraint
Problem #4: Find Pharmacophore in Ligands • Given: • Collection of N (= 5 to 10) small flexible ligands with similar activity (binding at same sites) Inhibitor binding to HIV protease Benzamidine binding to beta-Trypsin (3ptb)
Problem #4: Find Pharmacophore in Ligands • Given: • Collection of N (= 5 to 10) small flexible ligands with similar activity (binding at same sites) • A set of low-energy conformations (dozens to few hundreds) for each ligand
Problem #4: Find Pharmacophore in Ligands • Given: • Collection of N (= 5 to 10) small flexible ligands with similar activity (binding at same sites) • A set of low-energy conformations (dozens to few hundreds) for each ligand • Find a substructure (pharmacophore) that has a match in at least one conformation of each ligand
O O O H
O O O H
O O O O O O H H pharmacophore
Pharmacophore and Rational Drug Design • Pharmacophore identification is a form of “reverse engineering” to get a model of a binding site • A pharmacophore can be used to modify ligands into more potent drugs and/or to screen large databases of ligands for “leads”
Three Simultaneous Problems • Conformations? • Correspondence? • Transform? • But ligands are small molecules