Plant metabolomics database
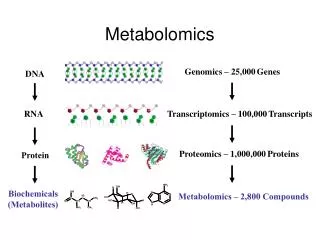
Metabolomics is described as comprehensive analysis in which all the metabolites of an organism are identified and quantified. It becomes even more challenging in the plants because it is estimated that the number of metabolites in the plant kingdom are more than 200 000 . Database organization

Plant metabolomics database
E N D
Presentation Transcript
Metabolomics is described as comprehensive analysis in which all the metabolites of an organism are identified and quantified. It becomes even more challenging in the plants because it is estimated that the number of metabolites in the plant kingdom are more than 200 000 . Database organization The data is organized into a mySQL database as per the Armet standards. One biological experiment consists of many target experiments. Each target experiment is performed at a different lab so the metadata of every target experiment is recorded at each lab along with the results.The biological metadata is recorded only once per experiment . Metadata: Metadata is the detailed information about an experiment which includes information about the Experiment Design, Sample Preparation, and Instrument Settings etc IMPORTANCE: Results of an experiment can be reproduced and validated. Plant metabolomics database
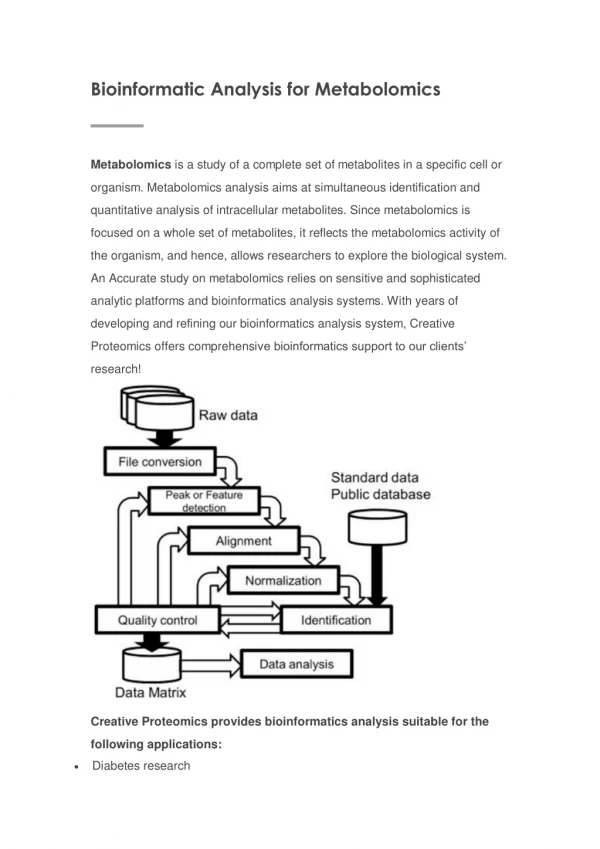
Databases and Tools ArMet (http://www.armet.org/ A framework for the description of plant metabolomics experiments and their results. ArMet encompasses the entire timeline of a plant metabolomics experiment. Architecture • Admin. Data that identify experiments and the users involved in them. • Biological Source. Genotype, provenance and identification data that describe the biological source material used in metabolomics experiments. • Growth. Data that describe the environments in which the biological source material is developed. • Collection. Data that describe the gathering of samples from biological source material. • Sample Handling. Data that describe the bulking/division and storage of material gathered during a collection event. • Sample Preparation. Data that describe the preparation of samples for presentation to an analytical instrument. • Analysis Specific Sample Preparation. Data that describe analysis/instrument specific sample preparation procedures. • Instrumental Analysis. Data that describe the analysis of the metabolite content of samples by analytical instruments. • Metabolome Estimate. Results sets for samples together with associated data pre-processing protocol information.
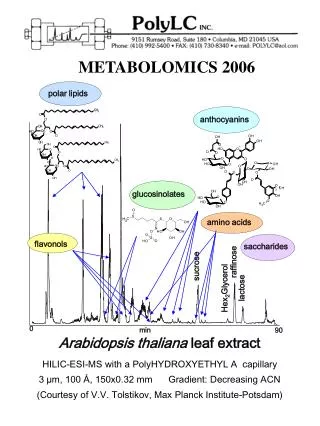
AraCyc (http://www.Arabidopsis.org/tools/aracyc/): AraCyc is a tool for visualizing biochemical pathways of Arabidopsis thaliana. The software allows querying and the graphical representation of biochemical pathways and expression data. Search AraCycThis link allows to search or browse the information (Pathways, Compounds, Enzymes...) contained in the AraCyc database OMICS Viewertool to overlay your data from gene expression, proteomic, or metabolomic experiments on a metabolic map Metabolic MapThe Metabolic map provides you with a 'bird's eye' view of Arabidopsis metabolism currently stored in AraCyc. Comparative AnalysisDraws summary tables comparing the metabolic profiles of Arabidopsis thaliana col. and E. coli K12 according to the information contained in AraCyc and EcoCyc Data Downloads download the AraCyc database files from the Plant Metabolic Network website.
Metabolomics Portal Pathway and Compound Databases Plant Metabolic Network (PMN) • Metabolic pathways from a large number of plants are comprehensively cataloged in PMN's PlantCyc database. PlantCyc includes experimentally supported,computationally predicted, and hypothetical pathways and enzymes. PMN is also a gateway to species-specific pathway databases for several plants, including Arabidopsis, rice, tomato, medicago, and poplar. . • KEGG Pathways Catalogs all kinds of pathways, including metabolic pathways. • ChemMine (NSF2010 project) • A compound database that also provides tools for compound clustering and other types of analysis. • PubChem • A compound database. • KEGG Ligand • A compound database. • ChEBI • A compound database. The compounds are classified by an ontological classification system. ChEBI uses nomenclature, symbolism and terminology endorsed by IUPAC and IUBMB. • KNApSAcK • A compound database that links metabolites to species, focusing specifically on plants.
MetNet is a suite of open-source of software to visualize, explore, statistically analyze and model mRNA, protein, and metabolite profiling data in the context of a metabolic and regulatory network map of Arabidopsis . Tools: MetNetDB is curator tool/Database of a metabolic and regulatory networkmap that contains a growing map of Arabidopsis entities (genes, RNAs, polypeptides, protein complexes and metabolites) and the catalytic and regulatory interactions between them FCModeler is a tool to graph and model data allowing submission of organelle-specific data and visualization of sub-cellular compartmentalized metabolites. GeneGobi analyzes datasets statistically and visually. MetNet (http://metnet.vrac.iastate.edu/ )
Metabolite Profiling Data NSF2010 Metabolomics Allows access to Arabidopsis metabolite profiling data. The Golm Metabolome Database Allows access to mass spectra and retention time index libraries, and metabolite profiling experiments. SpinAssign Performs batch-assignments of NMR peaks. Standard Spectrum Allows access to mass spectra data for compounds measured through mass spectrometry or NMR.
Data Analysis Tools • BL-SOM • This "Batch-learning self-organizing map" allows integrated analysis of a range of "omics" data. Two-dimensional feature maps can be generated linking genes and metabolites. • Cytoscape • An open source software platform for visualizing molecular interaction networks. • MapMan • A user-driven tool that displays large datasets (e.g. gene expression data from Arabidopsis Affymetrix arrays) onto diagrams of metabolic pathways. • PRIMe • The Platform for RIKEN Metabolomics provides a number of web-based tools for integrated analysis of metabolomic, transcriptomic, and other data. • Scripps Center for Mass Spectrometry • A variety of tools for peak alignment of MS and NMR data.
Human metabolomics database Human Metabolome Database (HMDB) is currently the most complete and comprehensive curated collection of human metabolite and human metabolism data in the world HMDB is a multi-purpose bioinformatics–cheminformatics–medical informatics database with a strong focus on quantitative , analytic or molecular-scale information about metabolites, their associated enzymes or transporters and their disease-related properties collection of experimental metabolite concentration data compiled from hundreds of mass spectra (MS) and Nuclear Magnetic resonance (NMR) metabolomic analyses performed on urine, blood and cerebrospinal fluid samples HMDB is divided into synoptic summary tables which, in turn, are linked to more detailed ‘MetaboCards’
HMDB's structure similarity search tool (ChemQuery) is the equivalent to BLAST for chemical structures HML refers to the Human Metabolite Library. This is a repository of all purchased, synthesized and isolated metabolites that have been acquired by the HMP team. HMDB from the perspective of a medical geneticist or a clinical chemist are its rich content and extensive linkage to metabolic diseases, to normal and abnormal metabolite concentration ranges (in many different biofluids), to mutation/SNP data and to the genes, enzymes, reactions and pathways associated with many diseases of interest Recent addition to the HMDB is a series of SimCell (SimCell is a metabolic simulation software package that allows complex metabolic pathways to be modeled at a cellular level and for ‘real-time’ movies of the enzymatic processes to be generated and graphed).
HORA suite (Human blOod Range vAlidator) consists of a Java application used to validate the metabolomic analysis of human blood against a database that stores the normal plasma and serum range concentrations of metabolites. The goal of HORA is to find the metabolites that are outside the normal range and to show those not present in the list provided by the user, for different thresholds of concentration. Moreover it supplies a graphical interface to manage the data. The software can also be used to compare different metabolomic techniques. HORA is open-source software and it can be accessed at http://www.paternostrolab.org
REFERENCES Nucleic Acids Res. 2007 January HMDB: the Human Metabolome Database David S. Wishart,1,5,7* Dan Tzur,1 Craig Knox,1 Roman Eisner,1 An Chi Guo,1 Nelson Young,1 Dean Cheng,1 Kevin Jewell, Springerlink HORA suite: a database and software for human metabolomics Stefania Bruschi1, Diego Calzolari1, Laurence Coquin1 and Giovanni Paternostr GMD@CSB.DB: the Golm Metabolome Database Joachim Kopka1, Nicolas Schauer1, Stephan Krueger1, Claudia Birkemeyer1, 1Max Planck Institute of Molecular Plant Physiology, Am Mühlenberg 1, 14476 Golm, Germany and 2Institute of Plant Sciences, Swiss Federal Institute of Technology, 8092, Zurich, Switzerland Received on October 20, 2004; revised on November 16, 2004; accepted on December 15, 2004 Database Resources in Metabolomics: An Overview Received: 15 January 2009 / Accepted: 15 April 2009 # Springer Science + Business Media, LLC 2009