Antibody Structure
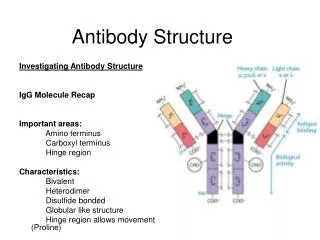
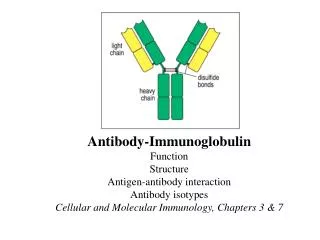
Antibody Structure. Investigating Antibody Structure IgG Molecule Recap Important areas: Amino terminus Carboxyl terminus Hinge region Characteristics: Bivalent Heterodimer Disulfide bonded

Antibody Structure
E N D
Presentation Transcript
Antibody Structure Investigating Antibody Structure IgG Molecule Recap Important areas: Amino terminus Carboxyl terminus Hinge region Characteristics: Bivalent Heterodimer Disulfide bonded Globular like structure Hinge region allows movement (Proline)
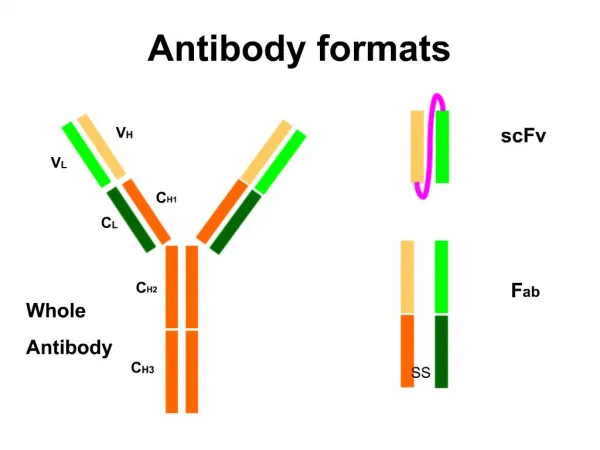
Antibody Functional Parts Structure of Ig was known Investigation of functional groups What parts of the Ig did what? Proteases used to digest below disulphide bond on heavy chain Pepsin Papain F(ab)2 still bivalent, therefore can bind 2 sites Therefore can still act like and antibody If reducing agent used, forms monovalent Ab structure---no globular structure can form Note: FC region is responsible for secondary biological function Ex. Ability to cross the placenta Ability to fix compliment
Antibody Diversity We know that there is a high level of diversity in the IRS; where does this come from? How do we get specificity region and biological function incorporated into Ab? Initial studies • Dr. Hood injected Balb c mice with multiple myeloma and precipitated out the light chains of IgG • Found out that although mice were genetically identical, the IgG samples from the mice were not. • After sequencing, found that variations lay in specific area. • Hood suggested that 2 genes encode for a single polypeptide chain giving rise to a variable and constant region • Notice that there is a high mutation rate in the V region • Therefore variability is created allowing a response to a wide range of antigens.
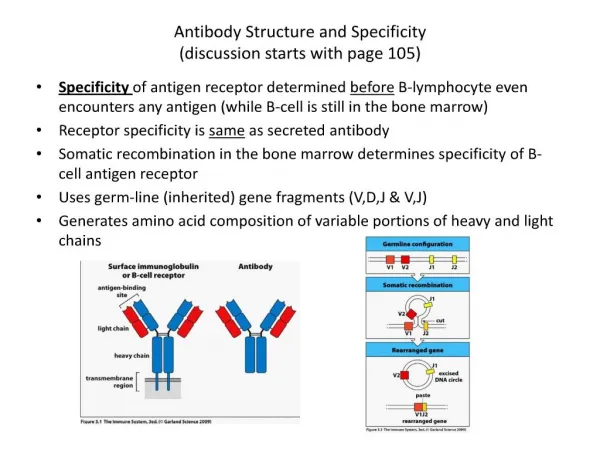
Antibody Diversity How do you make a variable and constant region on the same molecule? Dreyer and Bennet Proposed the 2-gene Model • Suggested that two separate genes encode a single Ig heavy or light chain. One gene for the the V region and the other for the C region. • Proposed that these two genes come together at the DNA level to form continuous message that can be transcribed and translated into a single Ig heavy or light chain. • Proposed that multiple V region genes were carried in the germ line, but only single copies of C region genes exist. • By postulating a single C region gene for each Ig class and subclass, the model could also account for the conservation of necessary biological effector functions while allowing for evolutionary diversification of V-region genes.
Antibody Diversity • Further Studies of Antibodies • Since there is such a diverse pool of Ig’s in the body, the best way to study them is by isolating the H chain • You wouldn’t isolate out the light chains as all Ig ‘s share light chain types (kappa and lambda). • The heavy chain is what sets them all apart.
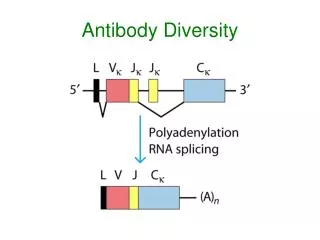
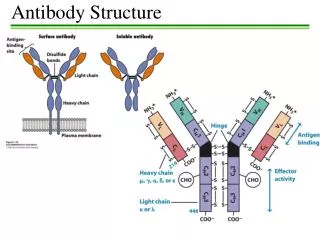
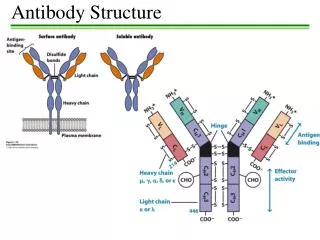
Ig Heavy Chains Separation of heavy chains • Separation column used • Heavy chains of particular classes will elute at different MW • In constant region of heavy chain there are 5 isotypes (µ,δ,γ,ε,α) • The heavy chains of the antibodies determine what class it is (ie IgA= α) • Each class can have either k or λ light chains Make-up of heavy and light chains k or λ light chain families contain V J C gene segments Rearranged V J segments encode the variable region of the light chain. The heavy chain family contains V D J C genes segments Rearranged V D J segments encode the variable region of the heavy chain. In Humans k = 70% In Mice k = 95% λ = 30% λ = 5%
Ig Heavy Chains Variation within C region of heavy chain Classes Subclass Light Chain IgA α α1 α2 κ λ IgD δ δ κ λ IgG γ γ1 γ2 γ3 γ4 κ λ IgE ε ε κ λ IgM µ µ κ λ • How many different genes are responsible for coding for Ig’s? __different genes encoding for Ig’s = diversity • What part of the Ab molecule does the 20 Ab recognize? • How many sub forms of IgG are there?
cys cys Ig Diversity How did these subtle differences arise? V C1 C2 • Since common elements are seen, there is a lot of homology • Indication that there is a common ancestor
γ2 ε2 δ2 β2 Ab Diversity Definition of a subclass. Hemoglobin (Hb) Over our lifetime there are 4 types of Hb I. Embryonic – α2ε2 II. Fetal -- α2 γ2 III. Adult – major α2 β2 IV. Adult – minor α2 δ2 • 5 genes involved here • High homology α1 α2 Gene duplication
Ab Subclasses • Phylogenetic tree based on • Size • Function • Homology • Events that will give rise to diversity • Gene duplication • Inversion • Translocation • Deletion β ε α δ γ
Where do the differences exist on the Antibody? 1. Isotypic determinants are constant region determinants that distinguish each Ig class and subclass within a species. 2. Allotypic determinants are subtle amino acid differences encoded by different alleles of isotype genes.Allotypic differences can be determined by comparing the same Ab class among inbred strains 3. Idiotypic determinants are generated by the conformation of the amino acid sequences of the L and H chain V regions. Each determinant = idiotope. The sum of these = idiotype.
There are two types of allotypic differences: 1. Simple Allotypic differences Ex k light chain k 1 k 2 C region Single amino acid difference @ 1 point 2. Complex Allotypic differences ex 2 alleles that are different by 30% What is the mechanism by with alleles can differ by 30%?
Events that will give rise to diversity Unequal Crossing Over Haptoglobin Binds to Hb and scavenges Fe Hp1 --- 83 aa Hp2 --- 142aa Primates only have Hp1 Humans have both Hp1 and Hp2 How could this have developed? α 1 83aa Hp1 α 2 142aa Hp2 sequence starts at 12 and repeats Mechanism α 1 83aa Hp1 71aa α 1 83aa 12aa α 2 142aa Hp2 Misalignment and crossing within the gene gives rise to α2 α2 is a better scavenger of Fe (evolution) Expansion and contraction of gene families