Single Nucleotide Polymorphism And Association Studies
Single Nucleotide Polymorphism And Association Studies. Stat 115 Dec 12, 2006. Outline. Definition and motivation SNP distribution and characteristics Allele frequency, LD, population stratification SNP discovery (unknown) and genotyping (known) SNP association studies

Single Nucleotide Polymorphism And Association Studies
E N D
Presentation Transcript
Single Nucleotide PolymorphismAnd Association Studies Stat 115 Dec 12, 2006
Outline • Definition and motivation • SNP distribution and characteristics • Allele frequency, LD, population stratification • SNP discovery (unknown) and genotyping(known) • SNP association studies • Case control studies, and family based association studies • Issues related to association studies
Polymorphism • Polymorphism: sites/genes with “common” variation, less common allele frequency >= 1%, otherwise called rare variant and not polymorphic • First discovered (early 1980): restriction fragment length polymorphism • Some definitions: • Locus: position on chromosome where sequence or gene is located • Allele: alternative form of DNA on a locus
Fundamental rules of genetics • Law of Segregation: a diploid parent is equally likely to pass along either of its two alleles P(pass copy 1) = P(pass copy 2) = ½ • Law of Random Union gametes unite in a random fashion, so allele A1 is no more likely to unite with allele A1 than A2, for example P(offspring is A1A1) = P(father passes A1) × P(mother passes A1) P(offspring is A1A2) = P(father passes A1) × P(mother passes A2)+ P(mother passes A1) × P(father passes A2) Slides from Karin S. Dorman
Hardy-Weinberg Equilibrium • Consider a single locus where there are two alleles segregating in a diploid population. Make the Hardy-Weinberg (HW) assumptions: • No difference in genotype proportions between the sexes. • Synchronous reproduction at discrete points in time (discrete generations) • Infinite population size (so that small variabilities are erased in the average) • No mutation. • No migration • No selection • Random mating Slides from Karin S. Dorman
Deriving HWE • Let genotypes at generation t be P11(t), P12(t), and P22(t). Then, • Genotype in the next generation will be • And p1(t+1)=p1(t); p2(t+1)=p2(t) • So in one step it returns to the equilibrium! Slides from Karin S. Dorman
A simple example • Consider this “population” Slides from Karin S. Dorman
SNP • Three classes of polymorphic markers: • Biallelic: SNPs and indels, less informative but more frequent & stable • Multiallelic: micro and mini satellites, more dynamic, high copy number loci have high mutation rate • Combination of above two • Single Nucleotide Polymorphism • Occasionally short (1-3 bp) indels are considered SNPs too • Come from DNA-replication mistake individual germ line cell, then transmitted
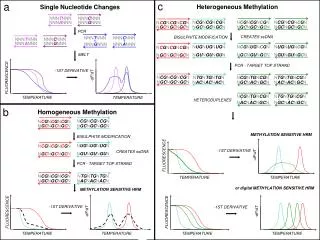
ATGGTAAGCCTGAGCTGACTTAGCGT-AT ATGGTAAACCTGAGTTGACTTAGCGTCAT SNP SNP indel SNPs result from replication errors and DNA damage They are a ‘polymorphic’ bit state at a nucleoside address What are Single Nucleotide Polymorphisms (SNPs)?
Why Should We Care • Personalized Medicine • Aithal et al., 1999, Lancet • Warfarin anticoagulant drug • CYP2C9 gene metabolizes warfarin, CYP2C9*1 (wild type) has two allelic variants: CYP2C9*2 & CYP2C9*3 (both single AA change) • Patients with variant alleles are poor warfarin metabolisers, often at higher risk of bleeding • Disease gene discovery • Association studies • Chromosome aberrations (copy number changes)
Disease resistant population Disease susceptible population Genotype all individuals for thousands of SNPs ATGATTATAG geneX ATGTTTATAG Resistant people all have an ‘A’ at position 4 in geneX, while susceptible people have a ‘T’ (A/T are the SNPs)
SNP Applications in Medicine • Gene discovery and allele mapping • Association-based (drug) candidate • polymorphism testing of a trait pool • Diagnostics / risk profiling • Drug response prediction • Homogeneity testing / study design • Gene function identification
Population Assignment– assessing competing hypotheses • The likelihood ratio method • Definition of competing hypotheses is essential Adapted from a slide of Steve DiFazio
Hypothesis testing in statistics … • Null hypothesis – assumed true unless there is an overwhelming evidence against it. • P-value – under the null hypothesis assess how “odd” aparticular aspect of the data is – the probability of seeing values as extreme or more extreme than the one we saw. • Using the likelihood ratio to find an effective aspect of the data to tell the two hypotheses apart – a way to guide your choice
SNP Distribution • Most common, > 1 SNP / 1KB • Balance between mutation introduction rate and polymorphism lost rate • Most mutations lost within a few generations • Often more transitions (A/G, C/T) than transversions (A/T, A/C, G/T, G/C) • In non-coding regions, often fewer SNPs at more conserved regions • In coding regions, often more synonymous than non-synonymous SNPs
SNP Characteristics: Allele Frequency Distribution • Most alleles are rare (minor allele frequency < 10%) • Allele frequency in different genomes have a large variation • Human > 1 SNP / 600-1KB, • Fly and maize have an order of magnitude greater number of polymorphism (1 SNP / 50-100 bp) • Nucleotide diversity is positively correlated with recombination rate
International HapMap Project • The International HapMap project is a recent, large-scale effort to facilitate GWAS studies: • Phase 1: 269 samples, 1.1 M SNPs • Phase 2: 270 samples, 3.9 M SNPs • Phase 3: 1115 samples, 1.6 M SNPs • Phase 3 platforms: • Illumina Human1M (by Wellcome Trust Sanger Institute) • Affymetrix SNP 6.0 (by Broad Institute)