The Role of HrasG12V Mutations in Skin Papilloma Development: Ki67 Staining and Allelic Imbalance Analysis
This study examines the impact of HrasG12V mutations on skin papilloma development in mice. Sections of mouse skin and papillomas from wild type (WT) and HrasG12V mice treated with TPA were analyzed. Haematoxylin and eosin (H&E) staining and immunohistochemistry (IHC) for Ki67 revealed hyperplastic epidermal changes and increased Ki67 labeling in HrasG12V mice. Additional analyses included laser-capture microdissection to isolate cellular samples and PCR amplification to study Hras allelic fractions. The findings highlight the role of Hras mutations in oncogenic signaling and cellular proliferation.

The Role of HrasG12V Mutations in Skin Papilloma Development: Ki67 Staining and Allelic Imbalance Analysis
E N D
Presentation Transcript
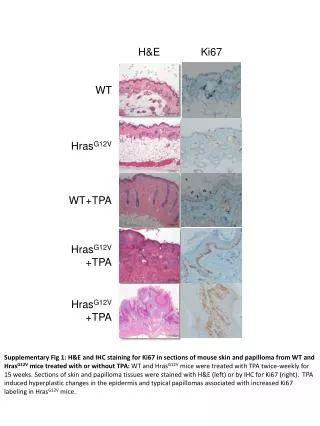
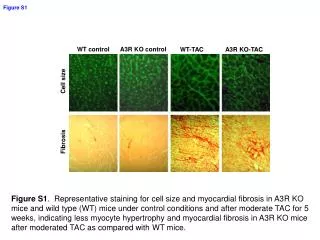
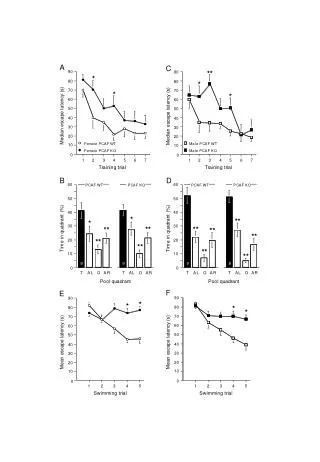
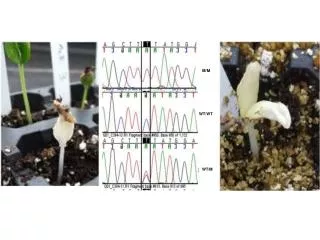
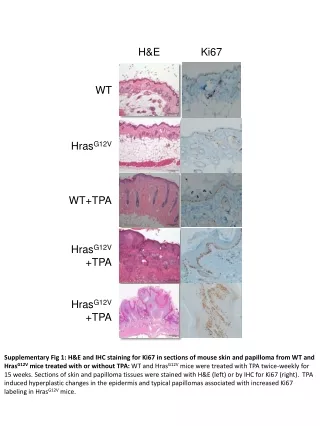
H&E Ki67 WT HrasG12V WT+TPA HrasG12V +TPA HrasG12V +TPA Supplementary Fig 1: H&E and IHC staining for Ki67 in sections of mouse skin and papilloma from WT and HrasG12V mice treated with or without TPA: WT and HrasG12V mice were treated with TPA twice-weekly for 15 weeks. Sections of skin and papilloma tissues were stained with H&E (left) or by IHC for Ki67 (right). TPA induced hyperplastic changes in the epidermis and typical papillomas associated with increased Ki67 labeling in HrasG12V mice.
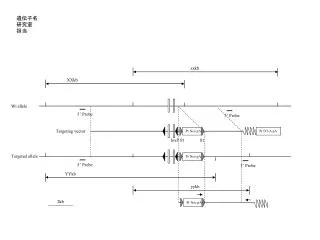
Codons 61 Codons 12 A T C T G G A G G G T C T A G A Allele 1 G T C A A G A G T C T G T A G G Allele 2 Supplementary Fig 2: G12V and Q61L mutations are located on different Hras alleles in tumors harboring mutations at both sites: PCR was performed to amplify genomic DNA with primers bracketing the region encoding the G12V and G61L mutations of Hras from three papilloma DNAs harboring mutations at both sites (see lanes with asterisk in Fig 2). Products were then subcloned into pcDNA3 and sequenced. A representative trace of two clones from the same tumor shows two Hras alleles, each containing one of the mutations.
Capped Before After Normal hyperplasia papilloma Supplementary Fig 3: Laser-capture microdissection of mouse skin tissues: Frozen sections were subjected to laser-capture microdissection to isolate histologically normal, hyperplastic and papilloma cells for analysis of Hras allelic fractions (see Fig 2D). A representative photomicrograph before and after microdissection as well as of the capped tissues is shown.
A B Supplementary Fig 4: HRAS allelic imbalance in papilloma correlates with mRNA expression of HRAS: A) RT-PCR products from RNA isolated from 5 different papillomas from TPA treated HrasG12V mice as well as RT-PCR of RNA isolated from two WT and two HrasG12V MEFs were incubated with or without GSU I, which digests only WT HRAS cDNA. B) Quantitation of copy number data from the above experiments. Total HRAS cDNA in the absence of GSU I was normalized to 100%.
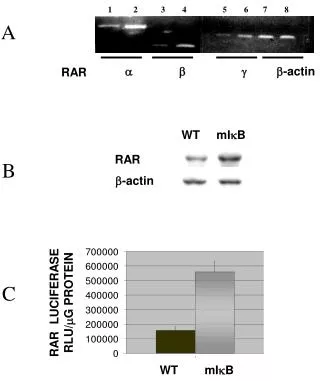
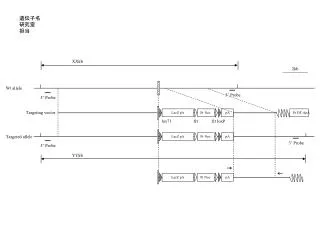
A C643 (G13R) Hth83 (Q61R) T24 (G12V) Hth83 WT C643 WT B EagI - + - + - + - + C Supplementary Fig 5: HRAS allelic imbalance in human cancer cell lines with HRAS mutations: A) HRAS gene sequence traces from cancer cell lines with HRAS mutation: C643 (G13R, thyroid), Hth83 (Q61R, thyroid) and T24 (G12V, bladder). HRAS mutant peak is higher than WT allele in C643 and Hth83 cells. T24 has HRAS G12V homozygous mutation. B) PCR products from DNA isolated from C643 or Hth83 cells were incubated with or without EagI, which digests only mutant HRAS DNA. C) Quantitation of HRAS alleles from the above experiments. Total HRAS DNA in the absence of EagI was normalized to 100%. Data represents mean + SEM of 3 independent experiments.
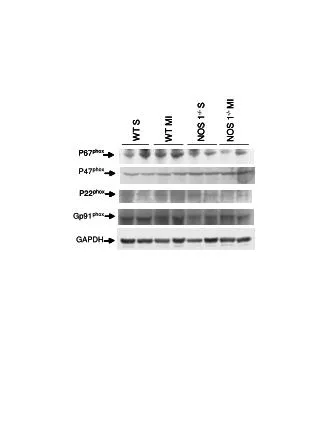
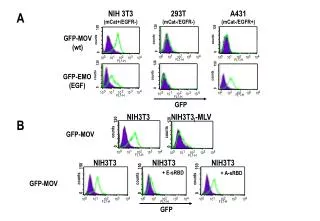
A Hth83 Cal-62 S 6 24 48 72 S 6 24 48 72 HRAS/KRAS pMEKS217/221 b-actin Hth-83 Cal-62 B S N H K S N H K KRAS HRAS NRAS pMEK pERKT202/Y204 pAKTS473 b-actin Supplementary Fig 6: Oncogenic RAS knockdown abrogrates MAPK signaling in cancer cell lines with RAS mutation. A) oncogenic Ras knockdown using RNAi in Hth-83 (HRAS mutation) and Cal62 ( KRAS mutation) cell lines for indicated time. Lysates were subjected to Western blotting with indicated antibodies. There are a marked decrease pMEK after oncogenic Ras knockdown over time in both cell lines. S: scramble B) isoform-specific Ras knockdown using RNAi in Hth-83 and Cal62 cells. Cell lystaes were subjected to Western blotting and detected with different antibodies. Only oncogenic Hras knockdown but not WT Kras and Nras diminished pMEK and pERK in Hth83 cells with Hras mutation. Only oncogenic Kras knockdown significantly diminished pMEK and pERK in Cal62 cells with Kras mutations. S:scramble, N:Nras, H:Hras, K:Kras. Presented data are representative for two independent experiments.
Cal62 Hth7 Hth83 Supplementary Fig 7: Oncogenic Ras knockdown inhibits cell proliferation. The indicated cell lines were transfected with indicated isoform-specific ras siRNA or scramble siRNA. Cells were counted at day 0 to day5. Each data point represents the mean plus SD of triplicates obtained in one of two independent experiments.