Neurodegenerative Diseases: Mechanisms of Aggregate Formation in Trinucleotide Repeat Disorders
This review examines neurodegenerative diseases linked to aggregate formation, focusing on Trinucleotide Repeat Disorders (TRDs). TRDs can be categorized into polyglutamine diseases, caused by CAG repeats, and non-polyglutamine diseases. Key examples include Dentatorubral-pallidoluysian atrophy (DRPLA), Huntington’s disease (HD), and various types of spinocerebellar ataxia (SCA). The review discusses how protein aggregation contributes to pathological mechanisms, including neuronal toxicity, apoptosis, and transcriptional dysregulation, ultimately leading to motor dysfunction, cognitive decline, and other symptoms in affected individuals.

Neurodegenerative Diseases: Mechanisms of Aggregate Formation in Trinucleotide Repeat Disorders
E N D
Presentation Transcript
Review the neurodegenerative diseases which are associated with aggregate formation and discuss how the aggregates may contribute to pathology Stuart Adams
Trinucleotide Repeat Disorders • Two broad groups of TRD’s. • Those caused by CAG repeats; coding glutamine – called polyglutamine diseases. • Those caused by other repeats – called non-polyglutamine diseases. • Polyglutamine repeats cause protein aggregation.
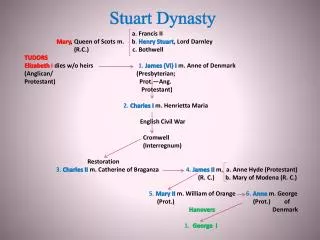
disease abbreviation gene chr protein Dentatorubropallidoluysian Atrophy DRPLA DRPLA (ATN1) 12 Atrophin-1 Huntington’s HD HTT 4 huntington Spinobulbar Muscular Atrophy SBMA AR X Androgen receptor Spinocerebellar Ataxia Type 1 SCA1 SCA1 6 Ataxin-1 Spinocerebellar Ataxia Type 2 SCA2 SCA2 12 Ataxin-2 Spinocerebellar Ataxia Type 3 SCA3 SCA3 14 Ataxin-3 Spinocerebellar Ataxia Type 6 SCA6 SCA6 19 Alpha-1A subunit of nerve calcium channel Spinocerebellar Ataxia Type 7 SCA7 SCA7 3 Ataxin-7 Polyglutamine diseases
DRPLA - phenotype • Ataxia and choreoathetosis • Dementia • Epilepsy • Young affected people may have progressive intellectual decline
DRPLA - gene • In asymptomatic individuals 6-35 CAG copies. • 48-93 copies in affected individuals. • 35-48 copies of CAG may or may not lead to an affected individual.
DRPLA - protein • Atrophin-1 is a nuclear protein with putative nuclear localizing signals • Mutant proteins aggregation and apoptotic cell death • Mutant proteins are more toxic to cells than full-length proteins
DRPLA - protein • Expanded polyglutamine stretches interact with: - TATA-binding protein (TBP)-associated factors (TAFII130) - CREB-binding protein (CBP) • Results in the suppression of CREB-dependent transcriptional activation that is vital for neuronal survival and plasticity
Huntington’s - phenotype • Motor disability with chorea • Mental disturbances including cognitive decline, changes in personality, and/or depression • Family history consistent with autosomal dominant inheritance
Huntingtin’s - gene • In asymptomatic individuals 26 or less CAG copies. • 40 or more copies in affected individuals. • 27-39 copies of CAG may or may not lead to an affected individual.
Huntingtin’s - protein • Probably not essential for normal function • Huntingtin is widely expressed with no obvious differences in the regional distribution of the mutant and wild-type protein. • Mutant proteins are more toxic to cells than full-length proteins
Huntingtin’s - protein From Biochem J. (2008) 412
SBMA - phenotype • X-linked, only males affected • Gradually progressive neuromuscular disorder • Weakness and atrophy of the proximal muscles. • Gynecomastia, testicular atrophy, and reduced fertility as a result of mild androgen insensitivity.
SBMA – AR gene • Androgen receptor gene (AR) • In asymptomatic individuals 9-34 CAG copies. • 38 or more copies in affected individuals. • 37 copies of CAG is a reduced penetrance allele
SBMA – AR protein • Transcription factor • Member of the steroid receptor superfamily • Individuals with SBMA produce an AR protein that contains a polyglutamine tail at the N-terminal end. • Polyglutamine tail alters the conformation of the AR protein (or an N-terminal peptide fragment) to produce neurodegeneration in SBMA. • The AR protein is expressed in the brain, spinal cord and muscle
SCA1 – phenotype Atrophy of cerebellum Loss of co-ordination Speech difficulty Decreased sensation in limbs (peripheral neuropathy) Loss of pyramidal nerve cells
SCA1 – gene Asymptomatic individuals have 6-44 CAG copies with stabilizing CAT codons present With diseased individuals the CAT codons are absent and alleles have 39-81 copies of CAG.
SCA1 – ataxin 1 protein Facilitates maneuvering nerve cell connections. Symptoms not directly caused by loss or normal ataxin-1 function. Ataxin-1 sequesters LANP protein (involved in cell communication)
SCA2 – phenotype/gene/protein As per SCA1 plus non-existent reflexes and eye focus problems. 14-31 CAG copies is asymptomatic. 36-64 copies = disease 31-36 copies individuals may or may not develop disease. Protein function unknown. Might be involved in protein-protein interactions.
SCA3 – phenotype/gene/protein As per SCA1 plus bulging eyes, small contractions of facial muscles and general rigidity 12-43 CAG copies is asymptomatic. 56-86 copies = disease 44-55 copies – relevance unknown Protein function unknown. Normally resides in cytoplasm
SCA6 – phenotype/gene Slowly progressing ataxia 4-18 CAG copies is asymptomatic. 21-33 copies = disease 19-20 copies – may or may not get disease Codes for a subunit of the calcium channels
SCA7 – phenotype/gene/protein Symptoms as SCA1 plus vision problems 4-19 CAG copies is asymptomatic. 37-306 copies = disease 20-29 copies – relevance unknown 30-36 copies – may or may not get disease Normal ataxin-7 function unknown.
Protein aggregation and neuronal inclusions From www.stanford.edu
Protein aggregation and neuronal inclusions • Aggregates are a feature of all polyglutamine diseases. • Many suggestions as to the pathogenesis of polyglutamine diseases and the role of the mutant proteins. From www.stanford.edu
Protein aggregation and neuronal inclusions • Glutamine is a polar molecule • Overabundance causes links to form within and between proteins • Molecules stick together forming strands held together by hydrogen bonds
Protein aggregation and neuronal inclusions • No folding into functional protein • Develop into tangled rigid groupings • Aggregates accumulate and interfere with nerve cell function by entrapping key cell regulatory factors.
Protein aggregation and neuronal inclusions • Is aggregation a cause or consequence of neurodegenerative diseases? • Are aggregates harmful or part of a cellular defence mechanism?
Perturbation of transcription • Excess glutamines lead to protein bundling called neuronal inclusions (NI) or inclusion bodies. • Research advances suggest that the polyglutamine tract region is proteolytically processed and a polyglutamine containing peptide fragment is retained in the nucleus, where it forms NIs. • Once in the nucleus, polyglutamine-expanded AR peptide fragments may cause pathology by interfering with transcriptional co-activators such as the CREB-binding protein (CBP) or PGC-1alpha, specificity protein 1 (Sp1) and neural protein BDNF.
Perturbation of transcription - CBP • The altered proteins sequesters CBP away from nuclear DNA • CBP can no longer activate transcription. • Fewer proteins produced – nerve cell death. • CBP has histone acetyltransferase activity – important for allowing TF access to DNA • Histone deacetylase (HDAC) inhibitors used for therapy. • Engineered form of CBP without glutamine repeats also possible therapy?
Perturbation of transcription – Sp1 • Sp1 is a sequence-specific transcription activator binding to GC-rich regions of DNA. • Interaction between N-terminal mutant protein and Sp1 interferes with Sp1-driven gene regulation. • Mutant Htt interacts with Sp1 disrupting Sp1-TAF130 complex altering expression of several neuronal target genes. • However studies show Sp1 reduction may be neuroprotective so situation is more complex (Qui et al. 2006).
Perturbation of transcription – BDNF • BDNF important for survival of striatal neurons. • Expression of BDNF regulated by REST/NRSF which binds to the promoter region. • Wild type protein binds and sequesters cytosolic REST/NRSF, limiting it’s translocation to the nucleus and allowing BDNF transcription. • Mutant protein doesn’t bind REST/NRSF leading to it’s accumulation in the nucleus and transcriptional represession of BDNF
Perturbation of transcription – PGC-1α • PGC-1α is a transcriptional co-activator involved in different metabolic programmes. • Mainly acts as a regulator of mitochondrial biogenesis and respiration. • Reduced levels of PGC-1α expression found in human and mouse HD brains. • PGC-1α expression is regulated by CREB-TAF4 complex which is impaired by mutant htt. • Also hypothesised that mutant protein binds directly to PGC-1α, affecting it’s ability to up regulate expression of it’s downstream targets (Cui et al. 2006)
Impairment of UPS in HD • Protein aggregates may impair ubiquitin proteosome system (UPS) function – conflciting evidence for this (Jana et al. 2001, Bowman et al. 2005 and Bett et al. 2006). • Proteosomes normally break down proteins tagged with ubiquitin. • Mutant protein continues to be tagged by ubiquitin for degradation. • Protein aggregates may occupy the proteosomes inhibiting their normal activity.
Summary Neurodegenerative diseases caused by increased CAG copies leading to mutant proteins with polyglutamine repeats Polyglutamine repeats cause protein aggregation Protein aggregation may affect neuronal development directly or indirectly by interacting with transcription factors or UPS.