Chapter 6. Multiple sequence alignment methods
Chapter 6. Multiple sequence alignment methods. Outline. What a multiple alignment means Scoring a multiple alignment Multidimensional Dynamic Programming Progressive alignment methods Multiple alignment by profile HMM training. Multiple alignment.

Chapter 6. Multiple sequence alignment methods
E N D
Presentation Transcript
Outline • What a multiple alignment means • Scoring a multiple alignment • Multidimensional Dynamic Programming • Progressive alignment methods • Multiple alignment by profile HMM training (C) 2000, 2001 SNU CSE Biointelligence Lab
Multiple alignment • Biologists produce high quality multiple alignments by hand using expert knowledge of protein sequence evolution. • Highly conserved regions • Buried hydrophobic residues • Influence of protein structure • Expected patterns of insertions and deletions (C) 2000, 2001 SNU CSE Biointelligence Lab
Multiple alignment • Manual multiple sequence alignment is tedius. • Automatic MSA methods are needed. • In general, an automatic method must have a way to assign a score so that better MSA get better scores. • Scoring a multiple alignment and searching over possible alignments should be distinguished. • In probabilistic modelling, scoring function is primary concern. • One of goals in probabilistic modeling is to incorporate as many of an expert’s evaluation criteria as possible into scoring procedure. (C) 2000, 2001 SNU CSE Biointelligence Lab
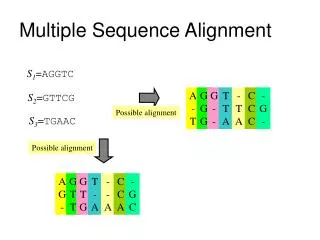
What a multiple alignment means • In a multiple sequence alignment, homologous residues among a set of sequences are aligned together in columns. • ‘Homologous’ is meant in both the structural and evolutionary sense. • Ideally, a column of aligned residues occupy similar three-dimensional structural positions and all diverge from a common ancestral residue. (C) 2000, 2001 SNU CSE Biointelligence Lab
What a multiple alignment means • Manually aligned example-10 imunoglobulin superfamily • A crystal structure of 1tlk(telokin) is known • The telokin structure and alignments to other related seqyences reveal conserved characteristics of the I-set immunoglobulin superfamily fold, including eight conserved β-strands and certain key residues in the sequences, such as two completely conserved cysteines in the b and f strands which form a disulfide bond in the core of the folded structure. (C) 2000, 2001 SNU CSE Biointelligence Lab
What a multiple alignment means (C) 2000, 2001 SNU CSE Biointelligence Lab
What a multiple alignment means • Except for trivial cases, it is not possible to create a single ‘correct’ multiple alignment. • Given pair of divergent but clearly homologus protein sequences, usually only 50% of the individual residues were superposable. • The Globin family, often used as a ‘typical’ problem in computational work, is in fact exceptional:almost the entire structure is convserved among divergent sequences. • Even the definition of ‘structurally superposable’ is subjective and can be expected to vary among experts. (C) 2000, 2001 SNU CSE Biointelligence Lab
What a multiple alignment means • Our ability to define a single ‘correct’ alignment will vary with the relatedness of the sequences being aligned. • An alignment of very similar sequences will generally be unambiguous, but there alignments are not of great interest to us. • For cases of interest, there is no objective way to define an unambiguously correct alignment. • Usually a small subset of key residues will be identifiable which can be aligned unambiguously for all the sequences in a family almost regardless of sequence divergence. • Core structal elements will also tend to be conserved and meaningfully alignable. (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment • Two important features of multiple alignments • Some positions are more conserved than others. • The sequences are not independent, but instead are related by a phylogenetic tree. (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment • An idealised way • Specifty a complete probabilistic model of molecular sequence evolution. • The probability of a multiple alignment can be calculated using evolutionary model. • We don’t have enough data to build such a model • Workable approximation:partly or entirely ignore the phylogenetic tree while doing some sort of position-specific scoring. (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment • Simplifying assumption • Individual columns of an alignment are statistically independent. • Then scoring function can be written as • Mi: column i of the multiple alignment m • S(mi):the score for column i • G:an function for scoring the gaps that occur in the alignments. • Unspecified function-affine scoring function can be used (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment-Minimum Entropy • Minimum Entropy • More variability in an alignment will be described by a higher entropy. Exactly matching sequences will have 0 entropy (completely organized) • To find the best alignment we want to have the minimum entropy. (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment-Minimum Entropy • Minimum entropy • Counting the residues in each column • Probability of residue a in column I (ML estimate) • Probability of a column(independence assumed) • Entropy is the negative log of the probability of the column. (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment-Minimum Entropy • Treating columns as statistically independent-Leaving out knowledge of phylogeny. • Actually very similar to HMM without gap information • The assumption that the sequences are independent can be reasonable if representative sequence of a sequence family s carefully chosen. • A variety of tree-based wdighting schemes have been proposed to deal with this problem to partially compensate for the defects of the sequence independence assumption. (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment-Sum of Pairs • Sum of pairs • Standard method of scoring multiple alignment • Similarity to HMM formulation • Do not use phylogenetic tree • Assumes statistical indepedence for the columns. • Not HMM formulation though (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment-Sum of pairs • Sum of pairs • Columns are scored by SP function using a substitution scoring matrix such as a PAM or BLOSUM matrix. • Use linear gap function or score affine gaps separately. • Sum N(N-1)/2 pairwise scores (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment-Sum of pairs • Problem of Sum of pairs • Sum of scores are not probabilistic correct extension to log-odds score. • Correct log-odds score extension: • SP score: • Evolutionary events are over-counted, a problem which increases as the number of sequemces increases. (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment-Sum of pairs • Example • an alignment of N sequences which all have leucine(L) at a certain position. • BLOSUM50 s(L,L)=5 • The SP score of the column is 5N(N-1)/2 • If instead there were one glycine(G) and N-1 Ls • BLOSUM50 s(G,L)=-4 • The SP score of the column is worse than the score for a column of all Ls by a fraction of 9(N-1) / 5N(N-1)/2 =18/5N (C) 2000, 2001 SNU CSE Biointelligence Lab
Scoring a multiple alignment-Sum of pairs • Difference is 18/5N • Relative difference between score between the correct and incorrect allignment decreases with the no. of sequences • Yet, if we have MORE evidence that L is conserved then an outlier out to DECREASE the score more. (C) 2000, 2001 SNU CSE Biointelligence Lab
Multidimensional Dynamic Programming • It is possible to generalise pairwise DP alignment to the alignment of N sequences. (C) 2000, 2001 SNU CSE Biointelligence Lab
Multidimensional Dynamic Programming • Assumptions • The columns of an alignment are statistically independent • The gaps are scored with a linear gap cost • Then the overall score S(m) for an alignment can be calculated as a sum of the scores for each column. (C) 2000, 2001 SNU CSE Biointelligence Lab
Multidimensional Dynamic Programming (C) 2000, 2001 SNU CSE Biointelligence Lab
Multidimensional Dynamic Programming • Simplifying the notation (C) 2000, 2001 SNU CSE Biointelligence Lab
Multidimensional Dynamic Programming • Straightforward Multidimensional DP • Pros • It can find optimal solution. • Arbitary column scoring function can be used • Only assumption is that column scores are independent. • Cons • There are 2^N-1 gap combinations for each entry • Huge computational complexity-O(2^N L^N) (C) 2000, 2001 SNU CSE Biointelligence Lab
Multidimensional Dynamic Programming-MSA • MSA can reduce the volume of the multidimensional dynamic programing matrix that needs to be examined • Optimally align up to 5-7 protein sequences of reasonable length(200-300 residues) (C) 2000, 2001 SNU CSE Biointelligence Lab
Multidimensional Dynamic Programming-MSA • Assumptions • SP scoring system • The score of a multiple alignment is the sum of the scores of all pairwise alignment defined by the multiple alignment. • Then the score of the complete alignment is given by • Let be the optimal pairwise alignment of k,l (C) 2000, 2001 SNU CSE Biointelligence Lab
Multidimensional Dynamic Programming-MSA • We can obtain a lower bound on the score of any pairwise alignment that can occur in the optimal multiple alignment. • Assume that we have a lower bound σ(a) on the score of the optimal multiple alignment, then for optimal multiple alignment a • We only need to consider pairwise alignment of k and l that score better than • A good bound σ(a) can be obtained by any fast heurist algorithm • Optimal pairwise alignment can be found using dynamic programming (C) 2000, 2001 SNU CSE Biointelligence Lab
Multidimensional Dynamic Programming-MSA • Now find the complete set of coordinate pairs (ik,il) such that the best alignment of xk to xl through (ik,il) scores more than • The costly multidimensional dynamic programming algorithm can be restricted to evaluate only cells in the intersection of all theses sets: I,e, cels (i1,i2,…iN) for which (ik,il) is in for all k,l. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods • Most commonly used approach • Works by constructing a succession of pairwise alignmensts. • Initially, two sequences are chosen and aligned by standard pairwise alignment;this alignment is fixed. • Then, a third sequence is chosen and aligned to the first alignment • This process is iterated until all sequences have been aligned. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods • Basically heuristic • It does not separate the scoring and optimising. • It does not directly optimise any global scoring function. • Fast and efficient, Generates reasonable result (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods • Differences between PA algorithms • The way that they choose the order to do the alignment • Whether the progression involves only alignment of sequences to a single growing alignment or whether subfamilies are built up on a tree structure and,at certain points, alignments are aligned to alignments. • Procedure used to align and score sequences or alignments against existing alignmetns. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods- Feng-Doolittle progressive multiple alignment • Calculate a diagonal matrix of N(N-1)/2 distances between all pairs of N sequences by standard pairwise alignment. Compute a distance matrix D=-log(S) • Construct a Guide tree from the distance matrix using a clustering algorithm • Starting from the first node added to the tree, align the child nodes. Repeat for all other nodes in the order that they were added to the tree. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods-Feng-Doolittle progressive multiple alignment • Converting alignment scores to distances • Doesn’t need to be accurate-the goal is only to create an approximate guide tree, not an evolutionary tree. • In phylogenetic tree construction, more care must be taken (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods-Feng-Doolittle progressive multiple alignment • Clustering • Done with The Fitch-Margooliash algorithm • Sequence-Sequence alignments • Done with usual pairwise dynamic programming. • A sequence is added to an existing group by aligning it pairwise to each sequence in the group in turn. • The highest scoring pairwise alignment determines how the sequence will be aligned to the group. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods-Feng-Doolittle progressive multiple alignment • ‘Once a gap,always a gap’ rule • After an alignment is completed, gap symbols are replaced with a neutral X character. • This rule allows pairwise sequenc alignments to be used to guide the alignment of sequences to groups or groups to groups; otherwise, any given pairwise sequence alignment would not necessarily be consistent with the pre-existing alignment of a group. • Desirable side effect:encouraging gaps to occur in the same columns in subsequent pairwise alignments. • Not needed in profile-based progressive alignment algorithms (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods • A problem with the Feng-Doolittle approach • all alignments are determined by pairwise sequence alignments. • It is advantageous to use position-specific information from the group’s multiple alignment to align a new sequence to it. (e.g. degree of sequence conservation) • Many progressive alignment methods use pairwise alignment of sequences to profiles or of profiles to profiles as a subroutine which is used many times in the process. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods • Linear gap scoring case • s(-,a)=s(a,-)=-g and s(-,-)=0 • Two profiles: sequence 1..n and n+1… N • Global alignment is • The first two sums are unaffected by the global alignment(s(-,-)=0) • Therefore the optimal alignment of the two profiles can be obtained by only optimising the last sum with the cross terms, which can be done exactly like a standard pairwise alignment. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods-CLUSTAW • Profile-based progresive multiple alignment • Works in much the same way as the Feng-Doolitle method except for its carefully tuned use of profile alignment methods. • Uses various heuristics. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods-CLUSTAW • Construct a distance matrix of all N(N-1)/2 pairs by pairwise dynamic programming. • Construct a guide tree by a neighbour-joining clustering algorithm. • Progressively align at nodes in order of decreasing similarity, using sequence-sequence, sequence-profile, and profile-profile alignment. • Scoring is basically SP. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods-CLUSTAW • Heuristics used • Sequences are weighted to compensate for biased representation in large subfamilies. • The substitution matrix is chosen on the basis of the similarity expected of the alignment. • Position-specific gap-open penalties are used. • Gap penalties are increased if there are no gaps in a column but gaps occur nearby in the alignment. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods-Iterative refinement methods • Problem with progressive alignment • Subalignments are frozen. • Once a group of sequemces has been aligned, their alignment to each other cannot be changed at a later stage as more data arrive. • Iterative refinement methods attempt to circumvent this problem. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods-Iterative refinement methods • Iterative refinement method • An initial alignment is generated • Then one sequence (or a set of sequences) is taken out and realigned to a profile of the remaining aligned sequences. • If a meaningful score is being optimized, this either increases the overall score or results in the same score. • Another sequence is chosen and realigned, and so on, until alignment does not change • Guaranteed to converged to a local maximum. (C) 2000, 2001 SNU CSE Biointelligence Lab
Progressive alignment methods-Iterative refinement methods • Barton-Sternberg multiple alignment • Find the two sequences with the highest pairwise similarity and align them using standard pairwise DP alignment. • Find the sequence that is most similar to a profile of the alignment of the first two, and align it to the first two by profile-sequence alignment. Repeat until all sequences have been included in the multiple aligment. • Remove sequence x1 and realign it to a profile of the other aligned sequences x2,… xN by profile-sequence alignment. Repeat for sequences x2…xN. • Repeat the previous realignment step a fixed number of times, or until the alignment score converges. (C) 2000, 2001 SNU CSE Biointelligence Lab
Multiple alignment by profile HMM training • Sequence profiles could be recast in probabilistic form as profile HMMs. • Profile HMMs could simply be used in place of standard profiles in progressive or iterative alignment methods. • Ad hoc SP scoring scheme can be replaced by more explicit profile HMM assumption. • Profile HMMs can also be trained from initially unaligned sequences using the Baum-Welch EM (C) 2000, 2001 SNU CSE Biointelligence Lab
Multiple alignment by profile HMM training- Multiple alignment with a known profile HMM • Before we estimate a model and a multiple alignment simultaneously we consider the simpler problem of obtaining a multiple alignment from a known model. • When we have a multiple alignment and a model of a small representative set of sequences in a family, and we wish to use that model to align a large member of other family members altogether. (C) 2000, 2001 SNU CSE Biointelligence Lab
Multiple alignment by profile HMM training- Multiple alignment with a known profile HMM • We know how to align a sequence to a profile HMM-Viterbi algorithm • Construction a multiple alignment just requires calculating a Viterbi alignment for each individual sequence. • Residues aligned to the same profile HMM match state are aligned in columns. (C) 2000, 2001 SNU CSE Biointelligence Lab
Multiple alignment by profile HMM training-Multiple alignment with a known profile HMM • Given a preliminary alignment, HMM can align further sequences. (C) 2000, 2001 SNU CSE Biointelligence Lab
Multiple alignment by profile HMM training- Multiple alignment with a known profile HMM (C) 2000, 2001 SNU CSE Biointelligence Lab
Multiple alignment by profile HMM training- Multiple alignment with a known profile HMM • Importance difference with other MSA programs • Viterbi path through HMM identifies inserts • Profile HMM does not align inserts • Other multiple alighment algorithms align the whole sequences. (C) 2000, 2001 SNU CSE Biointelligence Lab