Role of Pin1 in JNK1 Activation and IL-2 Production through H2O2 Treatment
80 likes | 210 Views
This study investigates the involvement of the Pin1 WW domain and T183 of JNK1 in their interaction under oxidative stress conditions. Using HEK.293 cells, we demonstrate how H2O2 affects Pin1's interaction with JNK1 and its subsequent effect on IL-2 production. Several co-transfection assays were conducted with FLAG-tagged Pin1 mutants and HA-JNK1, utilizing immunoprecipitation and immunoblotting techniques. The findings reveal the significance of Pin1 in JNK1 activation and its implications for cytokine production in T cells.

Role of Pin1 in JNK1 Activation and IL-2 Production through H2O2 Treatment
E N D
Presentation Transcript
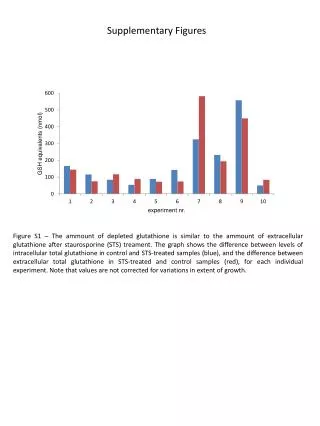
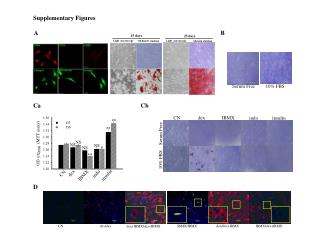
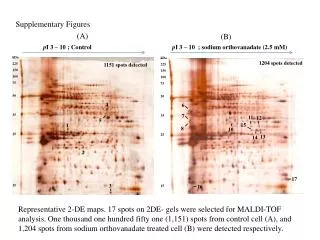
SupplementaryFigures SupplementaryFigure 1. T183 of JNK and WW domain of Pin1 are involved in the interaction. (a)HEK 293 cells transfected with plasmids as indicated were treated with 1 mM H2O2 for 1 h or left untreated. Immunoprecipitation and immunoblotting were performed. (b) HEK 293 cells were co-transfected with FLAG-tagged Pin1 WT or two deletion mutants of Pin1, WW or PPIase (amino acid residues 1-54 or 39-163, respectively) expression plasmids along with the HA-JNK1 plasmid, and exposed to 1 mM H2O2 for 1 h. Immunoprecipitation and immunoblotting were performed. FLAG-tagged Pin1 WW domain was not shown due to its small size. (c) HEK 293 cells were co-transfected with FLAG-tagged Pin1 WT or two point mutants of Pin1 (W34A or C113A) expression plasmids along with the HA-JNK1 plasmid, and exposed to 1 mM H2O2 for 1 h. Immunoprecipitation and immunoblotting were performed. SupplementaryFigure 2. H2O2 has no effect on Pin1 activity. Pin1-/- MEF cells were transiently transfected with FLAG-only or FLAG-Pin1 expression plasmids and then lysates from H2O2-treated for 1 h or untreated Pin1-/- MEF cells were immunoprecipitated with anti-FLAG beads. The immunoprecipitates were incubated with active JNK1, c-Jun, and [γ-32P] ATP for 10 min at 30°C. SupplementaryFigure 3. (a) Pin1does not interact with TCFb1.After transfection, HEK 293 cells were treated with 1 mM H2O2 for 1 h. Lysates were immunoprecipitated and immunoblotted with appropriate antibodies.(b) JNK1 phosphorylates Ser-232 of TCFb1.Active recombinant JNK1 was incubated with TCFb1 (WT, T242A, or S232/T242A) in 20 µl of kinase reaction buffer. The products of kinase reactions were separated by SDS-PAGE. The gels were dried and exposed to film. SupplementaryFigure 4. The data shown in Figure 4b were quantified as a percentage of phospho-JNK1.
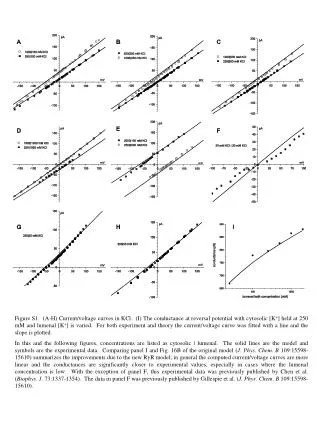
SupplementaryFigure 5. Pin1-induced JNK1 activity is stable. GST, GST-Pin1, or GST-Pin1 (C113A) proteins were pulled down with GST beads and then each bead complex was incubated with active JNK1 for 1 h at room temperature. After removal of GST-fusion proteins by centrifugation, the supernatants were subjected to SDS-PAGE and then immunoblotted with antibodies as indicated. The same supernatant samples were also subjected to in vitro kinase assays. SupplementaryFigure 6. Pin1 enhances JNK1/TCFβ1-mediated IL-2 production. After 16 h stimulation with PMA/ionomycin, supernatants were analyzed for IL-2 production using an ELISA assay. The results presented are representative of three independent experiments.Error bars indicate ±SD. Statistical significance was determined by the Student t test. *P <0.05 and **P <0.01 compared with control (TCFβ1 WT alone). #P <0.05 compared with control (TCFβ1 WT + JNK1). SupplementaryFigure 7. Hypothetical model for the role of Pin1 in JNK1 activation. The docking domain and the catalytic pocket of inactive JNK1 are depicted by rectangular and skewed triangular indentations, respectively. First, JNK1 is phosphorylated by several stresses, including UV, TNF-α and oxidative stress (H2O2), and partially activated by phosphorylation-induced structural changes in the docking domain and the catalytic pocket. The docking domain and the catalytic pocket of partially active JNK1 are indicated by oval and triangular indentations, respectively. Then, Pin1 binds to the pThr 183-Pro motif of JNK1 and induces subsequent cis-trans isomerization of the peptidyl-prolyl bond in the motif (inset). This isomerization induces further conformational change in the docking domain for substrate binding to become fully active. The docking domain of fully active JNK1 is depicted by pentagonal indentation. The JNK1 recognition site and the phosphoacceptor region of the target substrate are indicated by pentagonal and triangular extrusions, respectively. Double arrows denote up-regulation.
Pin1 Pin1 SupplementaryFigure 1 a b Y185F T183A Y185F T183A WT WT HA-JNK1 PPIase + + + + + + FLAG-Pin1 H2O2 WW WT - - + - + - + FLAG-Pin1 + + + + HA-JNK1 JNK1 IB: HA IB: HA JNK1 IB: FLAG Pin1 IP: FLAG IB: FLAG Pin1 p-JNK1 IB: p-JNK C113A WT FLAG-Pin1 IP: FLAG JNK1 IB: HA HA-JNK1 + + + IB: HA JNK1 JNK1 IB: HA Pin1 IB: FLAG IB: FLAG lysate IB: FLAG Pin1 IP: FLAG lysate IB: p-JNK p-JNK1 JNK1 IB: HA c IB: FLAG W34A lysate
Pin1 SupplementaryFigure 2 FLAG-Pin1 H2O2 - + + + - + [32P]-c-Jun Fold: 1.0 1.8 1.7 Pin1 IB: FLAG IB: tubulin tubulin SupplementaryFigure 3 a b + + + + + + HA-Pin1 S232A, T242A FLAG-c-Jun + + - - - - FLAG- TCFb1 (WT) - - + + - - T242A WT FLAG- TCFb1 (T242A) - - - - + + H2O2 - + - + - + [32P]- TCFb1 JNK kinase assay IB: HA TCFb1 Coomassie staining c-Jun IB: FLAG TCFb1 IP: FLAG IB: HA Pin1 c-Jun IB: FLAG TCFb1 lysate
SupplementaryFigure 4 Relative p-JNK level (%) (min)
SupplementaryFigure 5 Pin1 WT GST Pin1 C113A 0 20 40 60 0 20 40 60 0 20 40 60 (min) [32P]-c-Jun Fold 1.6 1.6 1.4 1.2 1.0 1.0 1.0 1.0 1.1 1.0 0.9 0.9 IB : p-JNK IB : JNK supernatant IB : GST input input input
SupplementaryFigure 6 1000 # 750 ** IL-2 (pg/ml) 500 * 250 0 TCFb1 WT - + - - - + - - - + - - - + - - - - - TCFb1 (S232A) - - + - - - + - - - + - - - + - - - - TCFb1 (T242A) - - - + - - - + - - - + - - - + - + + TCFb1 (S232A/T242A) - - - - + - - - + - - - + - - - + - - JNK1 - - - - - - - - - + + + + + + + + + + Pin1 WT - - - - - + + + + - - - - + + + + - - Pin1 (W34A) - - - - - - - - - - - - - - - - - + - Pin1 (C113A) - - - - - - - - - - - - - - - - - - +
P P SupplementaryFigure 7 TNF-α, oxidative stress, and UV Partially active Inactive P JNK1 JNK1 Pin1 P JNK1 Target Weak interaction Pin1 P JNK1 Isomerization Pin1 conformational change and full activation JNK1 JNK1 Pin1 JNK1 JNK1 Binding affinity Target JNK signaling