BLAST and Multiple Sequence Alignment
BLAST and Multiple Sequence Alignment. Announcements Quiz #3 on Thurs., May 17 on lectures presented April 26, May 3 and May 15 Writing assignments due May 24 at the beginning of class. Learning objectives-Learn the basics of BLAST and Psi-BLAST and CLUSTAL W

BLAST and Multiple Sequence Alignment
E N D
Presentation Transcript
BLAST and Multiple Sequence Alignment • Announcements • Quiz #3 on Thurs., May 17 on lectures presented April 26, May 3 and May 15 • Writing assignments due May 24 at the beginning of class. • Learning objectives-Learn the basics of BLAST and Psi-BLAST and CLUSTAL W • Workshop-Use of Psi-BLAST to determine sequence similarities. • Homework-Due May 20
BLAST • Basic Local Alignment Search Tool • Speed is achieved by: • Pre-indexing the database before the search • Parallel processing • Uses a hash table that contains neighborhood words.
Neighborhood words • The program declares a hit if the word taken from the query sequence has a score >= T when a scoring matrix is used. • This allows the word size (W) to be kept high (for speed) without sacrificing sensitivity. • If T is increased user the number of background hits is reduced and the program will run faster.
Which Program should one use? • Most researchers use methods for determining local similarities: • Smith-Waterman (gold standard) • FASTA • BLAST } Do not find every possible alignment of query with database sequence. These are used because they run faster than S-W
What are the different BLAST programs? • blastp • compares an amino acid query sequence against a protein sequence database • blastn • compares a nucleotide query sequence against a nucleotide sequence database • blastx • compares a nucleotide query sequence translated in all reading frames against a protein sequence database • tblastn • compares a protein query sequence against a nucleotide sequence database dynamically translated in all reading frames • tblastx • compares the six-frame translations of a nucleotide query sequence against the six-frame translations of a nucleotide sequence database. Please note that tblastx program cannot be used with the nr database on the BLAST Web page.
Identify Unknown Protein BLASTP; General protein comparison. Use ktup=2 for speed; ktup=1 for sensitive search. When to use a particular program Smith-Waterman Slower than FASTA3 and BLAST but provides maximum sensitivity TBLASTN Use if homolog cannot be found in protein databases; Approx. 33% slower Psi-BLAST Finds distantly related sequences. It replaces the query sequence with a position-specific score matrix after an initial BLASTP search. Then it uses the matrix to find distantly related sequences Problem Program Explanation
When to use a particular program (cont. 1) Problem Program Explanation Identify new orthologs TBLASTN:TBLASTX Use PAM matrix <=20 or BLOSUM90 to avoid detecting distant relationships. Search EST sequences w/in the same species. Always attempt to translate your sequence into protein prior to searching. Identify EST Sequence BLASTX;TBLASTX Identify DNA Sequence BLASTN Nucleotide sequence comparision
Filtering Repetitive Sequences • Over 50% of genomic DNA is repetitive • This is due to: • retrotransposons • ALU region • microsatellites • centromeric sequences, telomeric sequences • 5’ Untranslated Region of ESTs Example of ESTs with simple low complexity regions: T27311 GGGTGCAGGAATTCGGCACGAGTCTCTCTCTCTCTCTCTCTCTCTCTC TCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTC
Filtering Repetitive Sequences (cont. 1) • Programs like BLAST have the option of filtering out low complex regions. (Called Masking) • Repetitive sequences increase the chance of a match during a database search
PSI-BLAST • PSI-position specific iterative • a position specific scoring matrix (PSSM) is constructed automatically from multiple HSPs of initial BLAST search. Normal E value is used • The PSSM is created as the new scoring matrix for a second BLAST search. Low E value is used E=.001. • Result-1) obtains distantly related sequences 2) finds the important residues that provide function or structure.
Multiple alignment • Learning objectives-Understand usefulness of multiple alignment. Become familiar with ClustalW algorithm. Understand the difference between ClustalW and PSI-BLAST.
Steps to multiple alignment Create Alignment Edit the alignment to ensure that regions of functional or structural similarity are preserved Find conserved motifs to deduce function Structural Analysis Design of PCR primers Phylogenetic Analysis
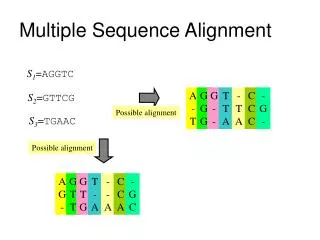
Multiple Sequence Alignment • Collection of three or more protein (or nucleic acid) sequences partially or completely aligned. • Aligned residues tend to occupy corresponding positions in the 3-D structure of each aligned protein.
Practical use of MSA • Helps to place protein into a group of related proteins. It will provide insight into function, structure and evolution. • Helps to detect homologs • Identifies sequencing errors • Identifies important regulatory regions in the promoters of genes.
Clustal W (Thompson et al., 1994) • CLUSTAL=Cluster alignment • The underlying concept is that groups of sequences are phylogenetically related. If they can be aligned then one can construct a tree. • Step1-pairwise alignments • Step2-create a guide tree • Step3-progressive alignment
Flowchart of computation steps in Clustal W (Thompson et al., 1994) Pairwise Alignment: Calculation of distance matrix Creation of unrooted Neighbor-Joining Tree Rooted NJ Tree (guide tree) and calculation of sequence weights Progressive alignment following the Guide Tree
Step 1-Pairwise alignments Compare each sequence with each other and calculate a distance matrix. A - B .87 - C .59 .60 - Different sequences Each number represents the number of exact matches divided by the sequence length (ignoring gaps). Thus, the higher the number the more closely related the two sequences are. A B C In this distance matrix, sequence A is 87% identical to sequence B
Step 1-Pairwise alignments Compare each sequence with each other and pairwise alignment scores human EYSGSSEKIDLLASDPHEALICKSERVHSKSVESNIEDKIFGKTYRKKASLPNLSHVTEN 480 Dog EYSGSSEKIDLMASDPQDAFICESERVHTKPVGGNIEDKIFGKTYRRKASLPKVSHTTEV 477 mouse GGFSSSRKTDLVTPDPHHTLMCKSGRDFSKPVEDNISDKIFGKSYQRKGSRPHLNHVTE 476 • SeqA Name Len(aa) SeqB Name Len(aa) Score • human 60 2 dog 60 76 • 1 human 60 3 mouse 59 57 • 2 dog 60 3 mouse 59 49
human:0.07429 dog:0.15904 mouse:0.3494 Guide Tree Sreal(ij) – Srand(ij) Sident(ij) – Srand(ij) Seff = x 100 Step 2-Create Guide Tree Use the Distance Matrix to create a Guide Tree to determine the “order” of the sequences. Distance from random sequence H - D 76 - M 57 49 - Different sequences H D M Branch length proportional to estimated divergence between dog and other sequences D = -ln(Seff) ( human:0.07429, dog:0.15904, mouse:0.34944);
human:0.07429 dog:0.15904 mouse:0.3494 Guide Tree Step 3-Progressive Alignment Align human and dog first. Then add mouse to the previous alignment. In the closely aligned sequences gaps are given a heavier weight (positive value) than gaps in more diver- gent sequences. “once a gap always a gap” Why a heavier weight for the closely aligned sequences? Because those gaps suggest separations between functional or structural entities. In more divergent sequences gaps may be produced as an artifact of sequences that are dissimilar.
Gap treatment • Short stretches of 5 hydrophilic residues often indicate loop or random coil regions (not essential for structure) and therefore gap penalties are reduced reduced for such stretches. • Gap penalties for closely related sequences are lowered compared to more distantly related sequences (“once a gap always a gap” rule). It is thought that those gaps occur in regions that do not disrupt the structure or function. • Alignments of proteins of known structure show that proteins gaps do not occur more frequently than every eight residues. Therefore penalties for gaps increase when required at 8 residues or less for alignment. This gives a lower alignment score in that region. • A gap weight is assigned after each aa according the frequency that such a gap naturally occurs after that aa in nature
Amino acid weight matrices • As we know, there are many scoring matrices that one can use depending on the relatedness of the aligned proteins. • As the alignment proceeds to longer branches the aa scoring matrices are changed to accommodate more divergent sequences. The length of the branch is used to determine which matrix to use and contributes to the alignment score.
Flowchart of computation steps in Clustal W (Thompson et al., 1994) Pairwise Alignment: Calculation of distance matrix Creation of unrooted Neighbor-Joining Tree Rooted NJ Tree (guide tree) and calculation of sequence weights Progressive alignment following the Guide Tree
Example of Sequence Alignment using Clustal W Asterisk represents identity : represents high similarity . represents low similarity
Multiple Alignment Considerations • Quality of guide tree. It would be good to have a set of closely related sequences in the alignment to set the pattern for more divergent sequences. • If the initial alignments have a problem, the problem is magnified in subsequent steps. • CLUSTAL W is best when aligning sequences that are related to each other over their entire lengths • Do not use when there are variable N- and C- terminal regions • If protein is enriched for G,P,S,N,Q,E,K,R then these residues should be removed from gap penalty list. (what types of residues are these?) Reference: http://www-igbmc.u-strasbg.fr/BioInfo/ClustalW/