Local alignment and BLAST
Local alignment and BLAST. Usman Roshan BNFO 601. Local alignment. Global alignment recursions: Local alignment recursions. Local alignment traceback. Let T(i,j) be the traceback matrices and m and n be length of input sequences. Global alignment traceback:

Local alignment and BLAST
E N D
Presentation Transcript
Local alignment and BLAST Usman Roshan BNFO 601
Local alignment • Global alignment recursions: • Local alignment recursions
Local alignment traceback • Let T(i,j) be the traceback matrices and m and n be length of input sequences. • Global alignment traceback: • Begin from T(m,n) and stop at T(0,0). • Local alignment traceback: • Find i*,j* such that T(i*,j*) is the maximum over all T(i,j). • Begin traceback from T(i*,j*) and stop when T(i,j) <= 0.
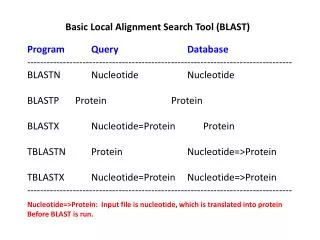
BLAST • Local pairwise alignment heuristic • Faster than standard pairwise alignment programs such as SSEARCH, but less sensitive. • Online server: http://www.ncbi.nlm.nih.gov/blast
BLAST • Given a query q and a target sequence, find substrings of length k (k-mers) of score at least t --- also called hits. k is normally 3 to 5 for amino acids and 12 for nucleotides. • Extend each hit to a locally maximal segment. Terminate the extension when the reduction in score exceeds a pre-defined threshold • Report maximal segments above score S.
Finding k-mers quickly • Preprocess the database of sequences: • For each sequence in the database store all k-mers in hash-table. • This takes linear time • Query sequence: • For each k-mer in the query sequence look up the hash table of the target to see if it exists • Also takes linear time