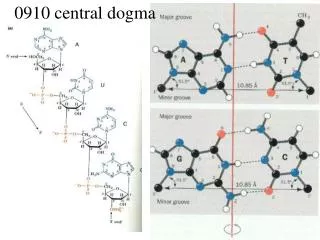
Central Dogma
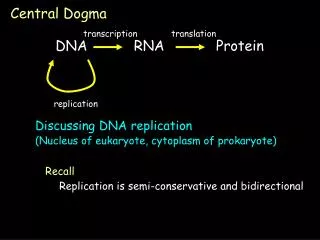
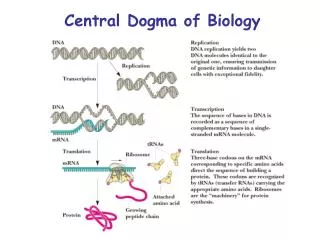
DNA Replication, Recombination, and Repair. Central Dogma. Central Dogma. DNA Replication – process of producing identical copies of original DNA strand separation followed by copying of each strand fixed by base-pairing rules. DNA replication is bidirectional.

Central Dogma
E N D
Presentation Transcript
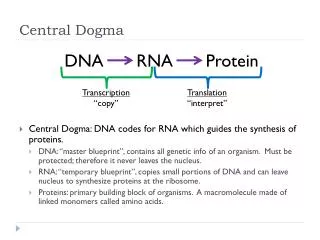
DNA Replication, Recombination, and Repair Central Dogma
DNA Replication – process of producing identical copies of original DNA • strand separation followed by copying of each strand • fixed by base-pairing rules
DNA replication is bidirectional. • involves two replication forks that move in opposite direction
DNA replication requires unwinding of the DNA helix. • expose single-stranded templates • DNA gyrase– acts to overcome torsional stress imposed upon unwinding • helicases– catalyze unwinding of double helix • disrupts H-bonding of the two strands • SSB (single-stranded DNA-binding proteins)– binds to the unwound strands, preventing re-annealing
Primer RNA primes the synthesis of DNA. Primase synthesizes short RNA.
DNA replication is semidiscontinuous • DNA polymerase synthesizes the new DNA strand only in a 5’3’ direction. Dilemma: how is 5’ 3’ copied? • The leading strand copies continuously • The lagging strand copies in segments called Okazaki fragments (about 1000 nucleotides at a time) which will then be joined by DNA ligase
Overall: each of the two DNA duplexes contain one “old” and one “new” DNA strand (semi-conservative) and half of the new strand was formed by leading strandand the other half by lagging strand.
DNA Polymerase = enzymes that replicate DNA • All DNA Polymerases share the following: • Incoming base selected in the active site (base-complementarity) • Chain growth 5’ 3’ direction (antiparallel to template) • Cannot initiate DNA synthesis de novo (requires primer) First DNA Polymerase discovered – E.coli DNA Polymerase I (by Arthur Kornberg and colleagues) Arthur Kornberg 1959 Nobel Prize in Physiology and Medicine Roger D. Kornberg 2006 Nobel Prize in Chemistry http://www.nobelprize.org
3’ 5’ exonuclease activity - removes incorrect nucleotides from the 3’-end of the growing chain (proofreader and editor) - polymerase cannot elongate an improperly base-paired terminus • proofreading mechanisms • Klenow fragment – removes mismatched nucleotides from the 3’’ end of DNA (exonuclease activity) • detection of incorrect base • incorrect pairing with the template (weak H-bonding) • unable to interact with the minor groove (enzyme stalls)
DNA Ligase = seals the nicks between Okazaki fragments • DNA ligase seals breaks in the double stranded DNA • DNA ligases use an energy source (ATP in eukaryotes and archaea, NAD+ in bacteria) to form a phosphodiester bond between the 3’ hydroxyl group at the end of one DNA chain and 5’-phosphate group at the end of the other.
Eukaryotic DNA Replication • Like E. coli, but more complex • Human cell: 6 billion base pairs of DNA to copy • Multiple origins of replication: 1 per 3000-30000 base pairs • E.coli 1 chromosome • Human 23 • E.coli circular chromosome; • Human linear
DNA Recombination = natural process of genetic rearrangement • recombinases • Holliday junction – crosslike structure
Mutations • Substitution of base pair • transition • transversion • Deletion of base pair/s • Insertion/Addition of base pair/s Macrolesions: Mutations involving changes in large portions of the genome DNA replication error rate: 3 bp during copying of 6 billion bp
Agents of Mutations • Physical Agents • UV Light • Ionizing Radiation • Chemical Agents • Some chemical agents can be classified further into • Alkylating • Intercalating • Deaminating • Viral
UV Light Causes Pyrimidine Dimerization • Replication and gene expression are blocked
Chemical mutagens • 5-bromouracil and 2-aminopurine can be incorporated into DNA
Deaminating agents • Ex: Nitrous acid (HNO2) • Converts adenine to hypoxanthine, cytosine to uracil, and guanine to xanthine • Causes A-T to G-C transitions
Acridines • Intercalate in DNA, leading to insertion or deletion • The reading frame during translation is changed
DNA Repair • Direct repair • Photolyase cleave pyrimidine dimers • Base excision repair • E. coli enzyme AlkA removes modified bases such as 3-methyladenine (glycosylase activity is present) • Nucleotide excision repair • Excision of pyrimidine dimers (need different enzymes for detection, excision, and repair synthesis)
QUIZ • Draw the structure of any nitrogenous base of your picking. (1 pt) • What is the difference between the glycosidic bond and the phosphodiester bond? (2 pts) • Give the reason why DNA utilizes the deoxyribose while RNA uses the ribose. (2 pts) • Enumerate all the enzymes and proteins involved in DNA replication and briefly state their importance/function. A short concise answer will suffice. (4 pts) • Give the partner strand of this piece of DNA: • 5-ACTCATGATTAGCAG-3 (1 pt)
RNA Transcription Central Dogma
Process of Transcription has four stages: • Binding of RNA polymerase at promoter sites • Initiation of polymerization • Chain elongation • Chain termination
Transcription (RNA Synthesis) • RNA Polymerases • Template (DNA) • Activated precursors (NTP) • Divalent metal ion (Mg2+ or Mn2+) • Mechanism is similar to DNA Synthesis
Reece R. Analysis of Genes and Genomes.2004. p47. • Limitations of RNAP II: • It can’t recognize its target promoter and gene. (BLIND) • It is unable to regulate mRNA production in response to developmental and environmental signals. (INSENSITIVE)
Start of Transcription • Promoter Sites • Where RNA Polymerase can indirectly bind
Preinitiation Complex (PIC) TATA box – a DNA sequence (5’—TATAA—3’) found in the promoter region of most eukaryotic genes. Abeles F, et al. Biochemistry. 1992. p391. Transcription Factors (TF): Hampsey M. Molecular Genetics of RNAP. Microbiology and Molecular Biology Reviews. 1998. p7.