DNA STRUCTURE
DNA STRUCTURE . Dr.Heba Al-Fares Lecture 6. DNA IS THE GENETIC MATERIAL ?. The genetic material must meet several criteria Genetic material must contain complex information . The ability to store large amounts of information—instructions for all the traits and functions of an organism.

DNA STRUCTURE
E N D
Presentation Transcript
DNA STRUCTURE Dr.Heba Al-Fares Lecture 6
DNA IS THE GENETIC MATERIAL ? • The genetic material must meet several criteria • Genetic material must contain complex information. The ability to store large amounts of information—instructions for all the traits and functions of an organism. • Genetic material must contain variation. This information must have the capacity to vary, because different species and even individual members of a species differ in their genetic makeup. • Genetic material must replicate. The genetic material must have the capacity to be copied accurately. • Transmission. At each cell division, the genetic instructions must be transmitted to descendent cells with great accuracy. • Genetic material must encode phenotype. The genetic material (the genotype) must have the capacity to “code for” (determine) traits (the phenotype).
Genetic material of cells… • GENES – units of genetic material that CODES FOR A SPECIFIC TRAIT • Gene a hereditary unit that occupies a specific position within the genome or chromosome; a unit that has one or more specific effects upon the phenotype of the organism; a unit that can mutate to various allelic forms; • Phenotype the observable characteristics of a cell or an organism, such as its size and shape, its metabolic functions, and its behavior. • Genotype the genetic constitution of a cell or an organism, as distinguished from its physical and behavioral characteristics, i.e., its phenotype . • Transformation Uptake of genetic material from an external source resulting in the acquisition of new traits (phenotype is changed)
Early Studies of DNA • Scientists in the 1830s and early 1840s knew that chromosomes were passed from one generation to the next, but they did not know what chromosomes were made of. • This mystery started to unravel in 1869, when Nucleic Acids were discovered by Friedrich Mieschner as a substance contained within nuclei.
Early Studies of DNA • DNA and RNA were first described by Friedrich Mieschner in 1869. • He isolated a phosphorus-containing material from the nuclei of cells found in pus from discarded surgical bandages, and called in “nuclein”. • He later found the same material in salmon sperm. • Albrecht Kossel (1853–1927), a German biochemist, discovered that nuclein contained four organic bases—adenine (A), thymine (T), guanine (G), and cytosine (C). • A Russian-American chemist named Phoebus Levene (1869– 1940) was working with Kossel, and he identified the sugar in nuclein as deoxyribose.
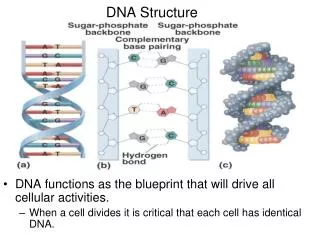
Chargaff’s Rule • Erwin Chargaff and his colleagues discovered that, within each species, there is some regularity in the ratios of the bases: the total amount of adenine is always equal to the amount of thymine (A T), and the amount of guanine is always equal to the amount of cytosine (G C) which became known as Chargaff ’s rules. • The bases form the “rung” on the DNA ladder by complementary pairing A == T C == G T == A G == C A == T T == A Sugar-phosphate backbone
Bacterial Transformation is Mediated by DNA • Griffith’s experiment was the earliest document evidence of transformation • Experiment by Frederick Griffith – 1928 • Demonstrated first evidence that genes are molecules • Two different strains of Streptococcus pneumoniae (type IIR connot revert to type IIIS) • Non-pathogenic = Avirulent = ROUGH cells (R) (type IIR) • Pathogenic = virulent = SMOOTH (S) (type IIIS) • Smooth outer covering = capsule • Capsule = slimy, polysaccharide • Encapsulated strains escape phagocytosis • The capsule alone did not cause pneumonia • Heat-killed S strain was avirulent • Ability to escape immune detection and multiply • When heat-killed S strain was mixed with living R strain the mouse dies of pneumoniae • Encapsulated strain (S) recovedred from dead mouse Now a live strain • The R strain had somehow acquired the ability to produce the polysaccharide capsule • Transformation • Ability to produce coat was an inherited trait Daughter cells also produced capsule type III trait must be replicated and transmitted. • Conclusion: genes from the dead S cells had “transformed” the live R cells into S.
Transformation 1. Mice injected with live cells of harmless strain R. 2. Mice injected with live cells of killer strain S. 3. Mice injected with heat-killed S cells. 4. Mice injected with live R cells plus heat-killed S cells. Mice live. No live R cells in their blood. Mice die. Live S cells in their blood. Mice live. No live S cells in their blood. Mice die. Live S cells in their blood.
More Transformation • Avery, MacLeod, and MacCarty carried this result further by fractionating the dead S cells. • They mixed various components (such as lipids, polysaccharides, protein, nucleic acids) of the S cells with live R cells, to determine which component caused the transformation. • They found the DNA by itself caused the transformation, and no other component had any effect. • This demonstrated that DNA was the hereditary material, but their results were not considered to be generally applicable to inheritance.
Avery, MacLeod, and MacCarty What is the transforming material? • Cell extracts treated with protein-digesting enzymes (protease) could still transform bacteria. • Cell extracts treated with RNA-digesting enzymes (Rnase) could still transform bacteria. • Cell extracts treated with DNA-digesting enzymes (DNase) lost their transforming ability. • They concluded that DNA, not protein, transforms bacteria • During the ’30s & 40’s proteins rather than DNA was thought to hold genetic information.
Hershey-Chase Experiment • Hershey and Chase in 1952, demonstrated the essential role of DNA using bacteriophage. • An important element of the Hershey-Chase experiment is that DNAcontains phosphorus but not sulfur, while protein contains sulfur but not phosphorus. • Thus it is possible to label the two types of molecule independently, with radioactive 32P and 35S. • Phage contain only DNA and protein, and no other types of molecule. • They used phage T2, infecting E. coli. • 32P-labelled DNA entered the bacterial cells when the phage infected them, and that the new generation of phage contained a significant amount of that labelled DNA. • 35S-labelled protein stayed outside the cells during an infection, and none of it ended up in the new phage. • This implies that DNA is necessary for phage replication.
Structure of DNA • The first essentially correct three-dimensional structure of the DNA molecule was proposed In 1953 James Watson and Francis Crick published a model of DNA structure. • Their work was based on X-ray crystallography data provided by Maurice Wilkins and Rosalind Franklin. • DNA consists of • two anti-parallel chains twisted into a helix. • The nitrogenous bases are paired in the center of the molecule, and • the phosphate-sugar backbones are on the outside. • Although DNA is the genetic material of all living cells, some viruses use RNA as their genetic material.
The Structure of DNA • The structure of DNA is divided to three level based on its complexity; the primary, secondary, and tertiary structures. • The primary structure of DNA refers to its nucleotide structure and how the nucleotides are joined together in linear manner to form a strand . • The secondary structure refers to DNA’s stable three-dimensional configuration, the helical structure (double helix) worked out by Watson and Crick. • DNA’s tertiary structures, which are the complex packing arrangements of double stranded DNA in chromosomes.
The Primary Structure of DNA • The primary structure of DNA consists of a string of nucleotides joined together by phosphodiester linkages. • Nucleotides DNA is typically a very long molecule and is therefore termed a macromolecule. In spite of its large size, DNA has a relatively simple structure: • It is a polymer, a chain made up of many repeating units linked together. As already mentioned, the repeating units of DNA are nucleotides, each comprising three parts: (1) a pentose sugar , (2) a phosphate, and (3) a nitrogenous base. • The sugars of nucleic acids—called pentose sugars— have five carbon atoms, numbered 1’, 2’, 3’, and 5’ • Four of the carbon atoms are joined by an oxygen atom to form a five-sided ring; the fifth (5) carbon atom projects upward from the ring. Hydrogen atoms or hydroxyl groups (OH) are attached to each carbon atom. • OH group attach to the 3’ carbon important to form the covalent linkage with each other.
Nucleotide the building blocks of DNA • A nucleotide contains either a ribose sugar (in RNA) or a deoxyribose sugar (in DNA). The atoms of the five-sided ring are assigned primed numbers. • RNA’s ribose sugar has a hydroxyl group attached to the 2-carbon atom, whereas DNA’s sugar, called deoxyribose, has a hydrogen atom at this position and contains one oxygen atom fewer overall. • This difference gives rise to the names ribonucleic acid (RNA) and deoxyribonucleic acid (DNA). • Further, the additional oxygen atom in the RNA nucleotide makes it more reactive and less chemically stable than DNA. • For this reason, DNA is better suited to serve as the long-term repository of genetic information.
The second component of a nucleotide is its nitrogenous base, which may be of two types—a purine or a pyrimidine . • Each purineconsists of a six-sided ring attached to a five-sided ring, whereas • Each pyrimidine consists of a six-sided ring only. • There are three pyrimidines • Cytosine is present in both DNA and RNA; however, • thymine is restricted to DNA, and • uracil is found only in RNA. • The three pyrimidines differ in the groups or atoms attached to the carbon atoms of the ring and in the number of double bonds in the ring. • In a nucleotide, the nitrogenous base always forms a covalent bond with the 1-carbon atom of the sugar. • sugar and a base together are referred to as a nucleoside (Adenosine , guanosine , thymidine, cytidine , uridine) If attach to deoxyribose they called deoxyguanosine.
The third component of a nucleotide is the phosphate group, which consists of a phosphorus atom bonded to four oxygen atoms. • The covalent attachment of one or more phosphate molecules to nucleoside creates nucleotide. • Phosphate groups are found in every nucleotide and frequently carry a negative charge, which makes DNA acidic. • The phosphate is always bonded to the 5-carbon atom of the sugar in a nucleotide.
Nucleotide the building blocks of DNA • The DNA nucleotides are properly known as deoxyribonucleotides or deoxyribonucleoside 5-monophosphates. • Because there are four types of bases, there are four different kinds of DNA nucleotides. • The equivalent RNA nucleotides are termed ribonucleotides or ribonucleoside 5-monophosphates. • RNA molecules sometimes contain additional rare bases, which are modified forms of the four common bases.
Nucleotide the building blocks of DNA • Polynucleotide strands DNA is made up of many nucleotides connected by ester bonds, which join the 5- phosphate group of one nucleotide to the 3-carbon atom of the next nucleotide. • These bonds, called phosphodiester linkages, are relatively strong covalent bonds; a series of nucleotides linked in this way constitutes a polynucleotide strand. • The backbone of the polynucleotide strand is composed of alternating sugars and phosphates; the bases project away from the long axis of the strand. • The negative charges of the phosphate groups are frequently neutralized by the association of positive charges on proteins, metals, or other molecules.
Nucleotide the building blocks of DNA • An important characteristic of the polynucleotide strand is its directionality or polarity. • At one end of the strand a phosphate group is attached only to the 5-carbon atom of the sugar in the nucleotide. This end of the strand is therefore referred to as the 5’ end. • The other end of the strand, referred to as the 3’end, has an OH group attached to the 3-carbon atom of the sugar. 5’- TACG-3’ • RNA nucleotides also are connected by phosphodiester linkages to form similar polynucleotide strands.
The double helix • The secondary structure of DNA refers to its three-dimensional configuration—its fundamental helical structure. • DNA’s secondary structure can assume a variety of configurations, depending on its base sequence and the conditions in which it is placed. • A fundamental characteristic of DNA’s secondary structure is that it consists of two polynucleotide strands wound around each other—it’s a double helix. • The sugar–phosphate linkages are on the outside of the helix, and the bases are stacked in the interior of the molecule. • The two polynucleotide strands run in opposite directions—they are antiparallel, which means that the 5’ end of one strand is opposite the 3’ end of the second.
DNA consists of two polynucleotide chains that are antiparallel and complementary, and RNA consists of a single nucleotide chain.
The double helix • The strands are held together by two types of molecular forces. 1- Hydrogen bonds link the bases on opposite strands. • These bonds are relatively weak compared with the covalent phosphodiester bonds that connect the sugar and phosphate groups of adjoining nucleotides. • The nature of the hydrogen bond imposes a limitation on the types of bases that can pair. • Adenine normally pairs only with thymine through two hydrogen bonds, and cytosine normally pairs only with guanine through three hydrogen bonds. • Because three hydrogen bonds form between C and G and only two hydrogen bonds form between A and T, C– G pairing is stronger than A– T pairing. • The specificity of the base pairing means that wherever there is an A on one strand, there must be a T in the corresponding position on the other strand, and wherever there is a G on one strand, a C must be on the other This keeps the width of the double helix constant. • The two polynucleotide strands of a DNA molecule are therefore not identical but are complementary.
The double helix 2- The second force that holds the two DNA strands together is the interaction between the stacked base pairs. • These stacking interactions contribute to the stability of the DNA molecule and do not require that any particular base follow another. • Thus, the base sequence of the DNA molecule is free to vary, allowing DNA to carry genetic information. • DNA normally consists of two polynucleotide strands that are antiparallel and complementary (exceptions are singlestranded DNA molecules in a few viruses). • The precise three-dimensional shape of the molecule can vary, however, depending on the conditions in which the DNA is placed and, in some cases, on the base sequence itself • Spiraling of the nucleotide strands creates major and minor grooves in the helix, features that are important for the binding of some DNA-binding proteins that regulate the expression of genetic information
B-DNA structure • The three-dimensional structure of DNA that Watson and Crick described is termed the B-DNA structure. • This structure exists when plenty of water surrounds the molecule and there is no unusual base sequence in the DNA—conditions that are likely to be present in cells. • The B-DNA structure is the most stable configuration for a random sequence of nucleotides under physiological conditions, and most evidence suggests that it is the predominate structure in the cell.
B-DNA structure • B-DNA is an alpha helix, meaning that it has a righthanded, or clockwise, spiral. • It possesses approximately 10 base pairs (bp) per 360-degree rotation of the helix; so each base pair is twisted 36 degrees relative to the adjacent bases. • The base pairs are 0.34 nanometer (nm) apart; so each complete rotation of the molecule encompasses 3.4 nm. • The diameter of the helix is 2 nm, and the bases are perpendicular to the long axis of the DNA molecule. • A space-filling model shows that B-DNA has a relatively slim and elongated structure.
B-DNA consists of an alpha helix with approximately 10 bases per turn. (a) Diagrammatic representation showing that the bases are 0.34 nanometer (nm) apart, that each rotation encompasses 3.4 nm, and that the diameter of the helix is 2 nm. (b) Space-filling model of B-DNA showing major and minor grooves. Key feature of the DNA Two strands of DNA form a right- handed double helix. The bases in opposite strands hydrogen bond according to AT/GC rule There are 10 nucleotides in each strand per complete 360⁰ turn of the helix
A and Z-DNA structure • Another secondary structure that DNA can assume is the A-DNA structure, which exists when less water is present. • Like B-DNA, A-DNA is an alpha (right-handed) helix, but it is shorter and wider than B-DNA and its bases are tilted away from the main axis of the molecule. • There is little evidence that A-DNA exists under physiological conditions. • 11 base per 360⁰ turn • A radically different secondary structure called Z-DNA forms a left-handed helix. In this form, the sugar–phosphate backbones zigzag back and forth, giving rise to the name Z-DNA (for zigzag). • 12 base per 360⁰ turn • Z-DNA structures can arise under physiological conditions when particular base sequences are present, such asstretches of alternating C and G sequences. Also high ionic strength. • Parts of some active genes form Z-DNA, suggesting that Z-DNA may play a role in regulating gene transcription.
DNA can assume several different secondary structures. These structures depend on the base sequence of the DNA and the conditions under which it is placed. Lehninger, Biochem 3/e, p.338. Fig.10-19.
Local variation in secondary structures • DNA is frequently presented as a static, rigid structure that is invariant in its secondary structure. In reality, the actual measurements vary slightly from one part of the molecule to another. • The twist between base pairs within a single molecule of B-DNA, for example, can vary from 27 degrees to as high as 42 degrees. • This local variation in DNA structure arises because of differences in local environmental conditions, such as the presence of proteins, metals, and ions that may bind to the DNA. • The base sequence also influences DNA structure locally.
Special Structures in DNA and RNA • In double-stranded DNA, the pairing of bases on opposite nucleotide strands provides stability and produces the helical secondary structure of the molecule. • Single-stranded DNA and RNA (the latter of which is almost always single stranded) lack the stabilizing influence of the paired nucleotide strands; so they exhibit no common secondary structure. • Sequences within a single strand of nucleotides may be complementary to each other and can pair by forming hydrogen bonds, producing double-stranded regions. • This internal base pairing imparts a secondary structure to a single-stranded molecule. • In fact, internal base pairing within single strands of nucleotides can result in a great variety of secondary structures. • single strands of nucleotides is a hairpin, which forms when sequences of nucleotides on the same strand are inverted complements. • The sequence 5’ TGCGAT 3’ and 5’ ATCGCA 3’ are examples of inverted complements. A hairpin consists of a region of paired bases (the stem) and sometimes includes intervening unpaired bases (the loop). RNA molecules may contain numerous hairpins, allowing them to fold up into complex structures • Another secondary structured, called a cruciform, can be made from an inverted repeat when a hairpin forms within each of the two single-stranded sequences.
Both DNA and RNA can form special secondary structures. (a) A hairpin, consisting of a region of paired bases (which forms the stem) and a region of unpaired bases between the complementary sequences (which form a loop at the end of the stem). (b) A stem with no loop. (c) Secondary structure, showing many hairpins, of an RNA component of a riboprotein, commonly referred to as the enzyme RNase P of E. coli. (d) A cruciform structure.
DNA Methylation • Methylation one of the parameter that controls transcription. • DNA methylation, in which methyl groups (–CH3) are added (methylases enzyme) to certain positions on the nucleotide bases . • In bacteria, adenine and cytosine are commonly methylated, whereas, in eukaryotes, cytosine is the most commonly methylated base. • Bacterial DNA is frequently methylated to distinguish it from foreign. • In eukaryotic DNA, cytosine bases are often methylated to form 5-methylcytosine. • In eukaryotic cells, methylation is often related to gene expression. • Methylation is most frequent on cytosine nucleotides that sit next to guanine nucleotides on the same strand: . . . GC . . . . . . CG . . . • Sequences that are methylated typically show low levels of transcription while sequences lacking methylation are actively being transcribed.