Analysis of Protein Interactions and Mutations in Yeast Using Co-IP Techniques
This study investigates the interactions among various proteins in *Saccharomyces cerevisiae* through co-immunoprecipitation (Co-IP) methods. We focus on key proteins like Sir2, Hst1, Hst2, and Sum1, along with their mutant variants. The analysis includes relative protein expression levels and interaction dynamics under different genetic backgrounds (wild-type and specific gene deletions). By exploring the protein network, we aim to enhance the understanding of regulatory mechanisms in yeast cells.

Analysis of Protein Interactions and Mutations in Yeast Using Co-IP Techniques
E N D
Presentation Transcript
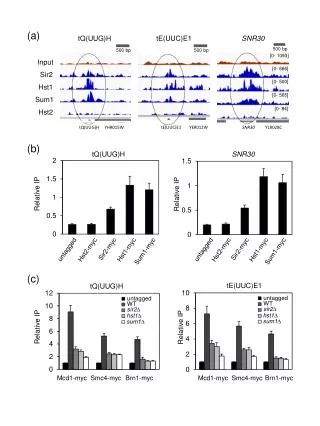
(a) tQ(UUG)H tE(UUC)E1 SNR30 500 bp 500 bp 500 bp [0- 1093] Input [0- 666] Sir2 [0- 500] Hst1 [0- 565] Sum1 [0- 84] Hst2 tQ(UUG)H YHR015W tE(UUC)E1 YER012W SNR30 YLR028C (b) tQ(UUG)H SNR30 Relative IP Relative IP Sir2-myc Sir2-myc untagged untagged Hst2-myc Hst1-myc Hst2-myc Hst1-myc Sum1-myc Sum1-myc (c) tE(UUC)E1 tQ(UUG)H untagged untagged WT WT sir2∆ sir2∆ hst1∆ hst1∆ sum1∆ sum1∆ Relative IP Relative IP Mcd1-myc Smc4-myc Brn1-myc Mcd1-myc Smc4-myc Brn1-myc