Basic Gene Expression Data Analysis--Clustering
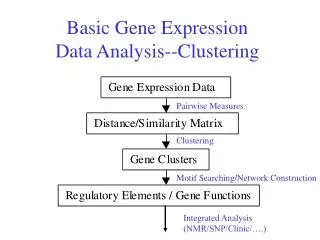
Basic Gene Expression Data Analysis--Clustering. Pairwise Measures. Clustering. Motif Searching/Network Construction. Integrated Analysis (NMR/SNP/Clinic/….). cDNA. mRNA. Treated. RT and label with fluor dyes. Control. Mix and hybridize target to microarray. Spot (DNA probe):

Basic Gene Expression Data Analysis--Clustering
E N D
Presentation Transcript
Basic Gene Expression Data Analysis--Clustering Pairwise Measures Clustering Motif Searching/Network Construction Integrated Analysis (NMR/SNP/Clinic/….)
cDNA mRNA Treated RT and label with fluor dyes Control Mix and hybridize target to microarray • Spot (DNA probe): • known cDNA or • Oligo MicroarrayExperiment
Collections of Experiments • Time course after a treatment • Different treatments • Disease cell lines • Data are represented in a matrix
Cluster Analysis • Grouping of genes with “similar” expression profiles • Grouping of disease cell lines/toxicants with “similar” effects on gene expression • Clustering algorithms • Hierarchical clustering • Self-organizing maps • K-means clustering
Gene Expression Clustering Normalized Expression Data Protein/protein complex Semantics of clusters: From co-expressed to co-regulated DNA regulatory elements
Key Terms in Cluster Analysis • Distance & Similarity measures • Hierarchical & non-hierarchical • Single/complete/average linkage • Dendrograms & ordering
Measuring Similarity of Gene Expression (x2,y2) • Euclidean (L2) distance • Manhattan (L1) distance • Lm: (|x1-x2|m+|y1-y2|m)1/m • L∞: max(|x1-x2|,|y1-y2|) • Inner product: x1x2+y1y2 • Correlation coefficient • Spearman rank correlation coefficient (x1, y1)
An Example x 3 y 4
Manhattan distance is called Hamming distance when all features are binary. Gene Expression Levels Under 17 Conditions (1-High,0-Low)
From Clustering to Correlation Expression Level Expression Level Gene A Gene B Gene B Gene A Time Time Expression Level Gene B Gene A Time
Hierarchical Clustering Given a set of N items to be clustered, and an NxN distance (or similarity) matrix, the basic process hierarchical clustering is this: 1.Start by assigning each item to its own cluster, so that if you have N items, you now have N clusters, each containing just one item. Let the distances (similarities) between the clusters equal the distances (similarities) between the items they contain. 2.Find the closest (most similar) pair of clusters and merge them into a single cluster, so that now you have one less cluster. 3.Compute distances (similarities) between the new cluster and each of the old clusters. 4.Repeat steps 2 and 3 until all items are clustered into a single cluster of size N.
Hierarchical Clustering Normalized Expression Data
Hierarchical Clustering 3 clusters? 2 clusters?
N genes N*N correlation matrix control t2 t0 t1 ... time Cluster Analysis • Correlation as measure of co-expression • Eisen et al. (1998) (PNAS, 95:14863) Experiment over time
N*N correlation matrix 1 2 3 Cluster Analysis • Scan matrix for maximum • Join genes to 1 node • Update matrix
Cluster Analysis • Result: Dendogram assemling N genes • Points of discussion • similarity based, useful for co-expression • dependent on similarity measure? • useful in preliminary scans • biological relevance of clusters?
Average distance Min distance Max distance Distance Between Two Clusters • single-link clustering (also called the connectedness or minimum method) : we consider the distance between one cluster and another cluster to be equal to the shortest distance from any member of one cluster to any member of the other cluster. If the data consist of similarities, we consider the similarity between one cluster and another cluster to be equal to the greatest similarity from any member of one cluster to any member of the other cluster. • complete-link clustering (also called the diameter or maximum method):we consider the distance between one cluster and another cluster to be equal to the longest distance from any member of one cluster to any member of the other cluster. • average-link clustering :we consider the distance between one cluster and another cluster to be equal to the average distance from any member of one cluster to any member of the other cluster. • Single-Link Method / Nearest Neighbor • Complete-Link / Furthest Neighbor • Their Centroids. • Average of all cross-cluster pairs.
Single-Link Method Euclidean Distance a a,b b a,b,c a,b,c,d c d c d d (1) (3) (2) Distance Matrix
Complete-Link Method Euclidean Distance a a,b a,b b a,b,c,d c,d c d c d (1) (3) (2) Distance Matrix
Identifying disease genes Tumor Liver Endothelial cells 1 Non-tumor Liver Proliferation Ribosomal proteins Liver-specific X. Chen & P.O. Brown et al Molecular Biology of the Cell Vol. 13, 1929-1939, June 2002
Human tumor patient and normal cells; various conditions • Cluster or Classify genes according to tumors • Cluster tumors according to genes
1) Select an initial partition of k clusters 2) Assign each object to the cluster with the closest center: 3) Compute the new centers of the clusters: 4) Repeat step 2 and 3 until no object changes cluster K-Means Clustering Algorithm
Basic Ideas : using cluster centre (means) to represent cluster Assigning data elements to the closet cluster (centre). Goal: Minimise square error (intra-class dissimilarity) : = Variations of K-Means Initialisation (select the number of clusters, initial partitions) Updating of center Hill-climbing (trying to move an object to another cluster). K-Means Clustering This method initially takes the number of components of the population equal to the final required number of clusters. In this step itself the final required number of clusters is chosen such that the points are mutually farthest apart. Next, it examines each component in the population and assigns it to one of the clusters depending on the minimum distance. The centroid's position is recalculated everytime a component is added to the cluster and this continues until all the components are grouped into the final required number of clusters.
The K-Means Clustering Method • Example
5000 2 k-means Clustering : Procedure (1) Initialization 1 Specify the number of cluster k : for example, k = 4 Expression matrix Each point is called “gene”
k-means Clustering : Procedure (2) Initialization 2 Genes are randomly assigned to one of k clusters
[(6,7) + (3,4) + …] k-means Clustering : Procedure (2) Calculate the mean of each cluster (6,7) (3,4) (3,2) (1,2)
Gene i to cluster c k-means Clustering : Procedure (4) Each gene is reassigned to the nearest cluster
Gene i to cluster c k-means Clustering : Procedure (4) Each gene is reassigned to the nearest cluster
k-means Clustering : Procedure (5) Iterate until the means are converged
6220 yeast genes 15 time points during cell cycle M/G1 phase G1 phase M phase k-means clustering : application : Result : 13 clusters of 30 clusters had statistical significance for each biological function S. Tavazoie & GM Church Nature Genetics Vol. 22, July 1999
Computation time: Hierarchical clustering O( m n2 log(n) ) K-means clustering t: number of iterations O( k t m n ) Memory requirement: Hierarchical clustering O( mn + n2 ) K-means clustering t: number of iterations O( mn + kn ) Computation Time and Memory Requirementn genes and m experiments
Issues in Cluster Analysis • A lot of clustering algorithms • A lot of distance/similarity metrics • Which clustering algorithm runs faster and uses less memory? • How many clusters after all? • Are the clusters stable? • Are the clusters meaningful?
Pattern Recognition • Clarification of decision making processes and automating them using computers supervised unsupervised • unknown number of classes • known number of classes • based on a training set • no prior knowledge • used to classify future observations • cluster analysis = one form