Plaque
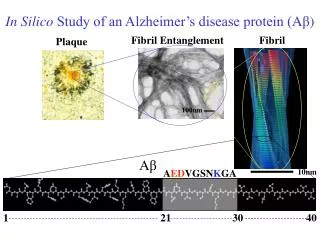
In Silico Study of an Alzheimer’s disease protein ( A β ). Fibril Entanglement. Fibril. Plaque. 100nm. A β. A ED VGSN K GA. 10nm. 1. 21. 30. 40. Experimental Background Experiments suggest Aβ(21-30) decapeptide may be the nucleating region for folding of full Aβ(1-40) peptide.

Plaque
E N D
Presentation Transcript
In Silico Study of an Alzheimer’s disease protein (Aβ) Fibril Entanglement Fibril Plaque 100nm Aβ AEDVGSNKGA 10nm 1 21 30 40
Experimental Background • Experiments suggest Aβ(21-30) decapeptide may be the nucleating region for folding of full Aβ(1-40) peptide. Aβ(21-30) Relevant to Development of AD • What is the Question? • To determine the fold of Aβ(21-30) with atomic detail and find the stabilizing interactions. • How does it help? • The fold of Aβ(21-30) may provide plausible scenarios for the initial stages of fibril formation of full Aβ(1-40). • Identification of amino acids important for folding stability may lead to strategies to • prevent fibril formation. • What did we Find? • Aβ(21-30) adopts a loop conformation with center in S26, stabilized by hydrophobic interactions between V24 and K28. • There is a value for the strength of the electrostatic interaction that optimizes the stability of the loop.
K28 K28 S26 S26 V24 V24 Experiments (our Collaborators) Nuclear Magnetic Resonance data leads to two model structures of Aβ(21-30) in solution: K28 below loop K28 above loop • Aβ(21-30) adopts a loop conformation. • V24 and K28 are close. • The two model structures differ in the orientation of K28. Which one is true? Lazo et at.,submitted to J. Mol. Biol.
Simulations(my work) • Discrete Molecular Dynamics simulations of Aβ(21-30) in a cubic box of 40 Å with periodic boundary conditions for 50ns. • kBT=0.592 Kcal/mol (room temperature). • We perform simulations for different electrostatic interaction (EI) strengths: • 0.00 < EI < 1.5 Kcal/mol (typical in the surface of proteins) • 1.50 < EI < 2.5 Kcal/mol (typical in the interior of proteins) • Hydrogen-Bond strength = 3.5 Kcal/mol (typical in the surface of proteins) • HPvalues in the range-9.3<HP<1.3 Kcal/mol (negative stands for repulsive)
K28 V24 S26 Simulations Results V-K Unpacked V V-K Packed K Solvent Accesible Surface (Å2) T.H.M.: • Hydrophobic interactions responsible for loop formation.. • Electrostatic interaction of 1.5Kcal/mol optimizes loop stability.
E22 D23 K28 Simulation Results (II) The unpacked conformations at EI=2.5Kcal/mol have strong electrostatic interactions! E22···K28 D23···K28 E22···K28 D23···K28 p=0.48 p=0.29 p=0.23 Hypotheis for Future work: We hypothesize that Aβ(21-30) undergoes partial unpacking of V24···K28 contacts and form D23···K28 electrostatic interactions upon fibril formation.
n v E22 D23 K28 Simulation Results (III) P() x 10-3 K28 S26 V24 (deg) K28 below loop plane K28 above loop plane THM: Electrostatic interaction stabilizes K28 above the loop plane. E22···K28 D23···K28
B- B+ 0.0 1 5 1.5 6 16 2.5 5 21 B- B+ 0.0 1 1 1.5 7 9 2.5 11 13 Simulation Results (IV) S K V D THM : only electrostatic interactions between E22 and K28 correlate with the orientation of K28 above the loop.
V S K D E K Conclusions • Aβ(21-30) adopts a loop conformation centered at S26, • stabilized by hydrophobic interactions between V24 and K28. • There is a particular electrostatic interaction strength that • optimize the stability of the loop conformations. • Electrostatic interactions strengths typical of the interior of proteins destabilize the • loop conformations and form strong electrostatic interactions, preferentially D23···K28. Future Work • Verify the hypothesis that Aβ(21-30) undergoes partial unfolding of V24-K28 and • formation of electrostatic interaction D23-K28 upon fibril formation with simulation • studies of many Ab(21-30).
Thank you! Collaborators Sergey V. Buldyrev# Luis Cruz* Feng Ding† Nikolay Dokholyan† Alfonso Lam Ng* Noel Lazo¶ Manuel Marques§ Shouyong Peng* Eugene Shakhnovich‡ David B. Teplow ¶ Brigita Urbanc* Sijung Yun* *Center for Polymer Studies and Dept of Physics, Boston Univ., Boston MA, USA. †Dept of Biochemistry and Biophysics, School of Medicine, Univ. of North Carolina at Chapel Hill, Chapel Hill NC, USA. # Dept of Physics, Yeshiva University, New York NY, USA. ‡Department of Chemistry and Chemical Biology, Harvard Univ., Cambridge MA, USA § Dept. of Physics and Condensed matter C IV, Univ. Autonoma Madrid, Madrid, Spain. ¶Center for Neurological Diseases, Brigham and Women’s Hospital and Dept. of Neurology, Harvard Medical School, Boston MA, USA
Ab Model D23 N27 A21 G25 G27 V24 S26 A30 E22 K28 Three bonding types describe the protein geometry: 1st neigh. 2nd neigh. 3rd neigh. (covalent) (angle) (dihedral) 1 1 1 1 2 4 2 2 4 4 3 3 3
electrostatics hydropathy Hydrogen Bond - + N O C N + + + - - - O EI Ab Model (cont.) - - - - Three types of atomic interactions: + HP
Hydropathy Interactions When two atoms i and j make a contact, they interact with a hydropathy strength. HPij=HPi+HPj , HP: free energy of transfer HP Aqueous Phase Gas Phase SAS :solvent accesible surface σ: atomic solvation parameter HPi=ΔSASi · σi
Arginine (R) Lysine (K) ΔFR=σC ·(ΣCi SASCi) +σN ·(ΣNi SASNi) ΔFK=σC ·(ΣCi SASCi) +σN ·SASNi Solve for σC and σN Atomic Solvation Parameters HPi=ΔSASi · σi
Two Representative Conformations K28 below loop plane K28 above loop plane E22-K28 interaction shown
i j i i j j Results: Loop Flexibility σ Δd • The loop is rigid only when close to the turn • E22-K28 and D23-K28 salt-bridges increase loop rigidity. • When loop forms, distances E22-K28, D23-K28 and V24-K28 corresponding to attractive • interactions decrease the most. • Flexibility of the loop strands allows K28 to flip-flop its orientation with respect to the loop