Breakout Question
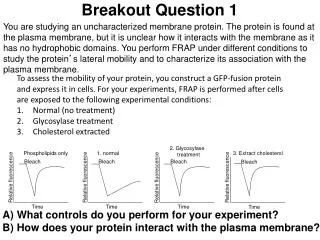
Breakout Question. Eg5 is a kinesin family member in Xenopus. To understand Eg5 function in vivo, cells are transfected with RNAi directed against this motor protein. The following images are obtained of mitotic cells. A) Describe the experiment.

Breakout Question
E N D
Presentation Transcript
Breakout Question Eg5 is a kinesin family member in Xenopus. To understand Eg5 function in vivo, cells are transfected with RNAi directed against this motor protein. The following images are obtained of mitotic cells. A) Describe the experiment. B) What do the data tell you about the function of Eg5 in cells? 1
Answer A) To determine the in vivo function of the kinesin protein Eg5 during metaphase, RNAi-mediated knockdown of Eg5 was performed and then DNA and microtubules were stained and assayed during metaphase. As a negative control, RNAi against a non-essential or non-expressed gene such as GFP is utilized. B) Without Eg5, the microtubule spindle network is disorganized and chromosomes are no longer aligned at the middle of the cell. These data suggest that Eg5 plays a role in chromosome alignment and spindle orientation. Without proper spindle formation though, chromosomes will not align, thus this is likely the primary problem. More experiments need to be performed to distinguish primary or secondary nature of this effect.
Answer • Protein B most likely represents a cyclin. According to the diagram, its presence in the cell both cycles and oscillates, with protein B dropping off in concentration after each mitotic stage, which reflects the ubiquitinylation that occurs at the end of metaphase. This dropoff corresponds from T1-T2, T3-T4, and T5-T6, thus confirming that it cycles between mitosis and interphase, and also helps modulate cell division activity, all characteristic of cyclin dynamics. • If the cells were not fertilized synchronously, it would be difficult to infer much of anything regarding the proteins because cells would be on different parts of their cycle with regards to mitosis and interphase. You would thus not be able to standardize your comparison of the cycles effectively because you would lose control over which stage they are at. Thus, this non-synchronized activity would skew results, as different cells would very likely be at different stages and protein activity might not correlate appropriately. • With the addition of an inhibitor of protein synthesis, cyclin cannot be produced. Cyclin buildup allows for entering mitosis in the cell cycle. If added to the cells at T2, the cells will not be able to enter mitosis at T3. They should stay at the 4-cell stage. If added to the cells at T3, the cells will be able to exit T3 into interphase at T4. However, the cells should arrest at T4 because they will not be able to enter mitosis in T5. Thus, the cells should stay at the 16-cell stage.
cAMP levels help regulate the amount of glycogen breakdown following epinephrine binding. Phosphodiesterase (PDE) is an enzyme that downregulates cAMP levels. PDE contains the following motif: X-R-R-X-S-F. You want to test how mutations in this motif affect cAMP levels. You transfect wild type cells with cDNA encoding either wild type or PDE with S mutated to D (this substitution mimics the phophorylated form of PDE), then you assay for levels of cAMP. The following results are found: (A) Explain wild type regulation of PDE and how the S-to-D mutation in PDE makes the cells insensitive to Adenylate Cyclase (AC) activation. (B) Additionally, you observe that the levels of activated PKA are diminished in the mutant cells compared to wild type. Explain how the PDE mutation might affect PKA activation.
Answer (A) Normally, when AC becomes active, it produces cAMP. The increased levels of cAMP induce activation of PKA. Active PKA phosphorylates multiple targets, one of which is PDE. The phosphorylation of PDE activates the enzyme, thus downregulating cAMP levels. Changing S-to-D mimics the phosphorylated form of PDE. The result of this mutation is the PDE enzyme will be perpetually active. When AC becomes active cAMP levels will increase, but the constitutively active PDE continually breaks it down. (B) PKA activity is dependent on cAMP. Without cAMP, PKA will remain inactive. Constitutively active PDE will keep cAMP levels low, thus keeping PKA in an inactive form.
Answer Sunburn acts upstream of receptor activation. The rationale for this conclusion is that activation of the Eph receptor can rescue the loss of sunburn, suggesting all downstream components of Eph RTK signaling are intact. It is most likely the ligand for the Eph RTK
You recently joined a lab that studies apoptosis with a special focus on the role of p53 in the process. Knowing the extensive utility of C. elegans genetics in the study of apoptosis, you embark on a genetic screen to identify mutants that can modify the apoptotic-resistant phenotype observed in loss-of-function p53 mutants. In wild type worms, gonad cells undergo extensive apoptosis in response to ionizing radiation (IR), but in p53-null worms apoptosis is not induced. In your screen you identify 2 new mutants (termed yin and yang) that can restore IR-induced apoptosis. To further characterize the exact placement of these components in the apoptotic pathway, you perform additional double mutant genetic analysis. The results are shown in the table below. Double mutant genotypeIR-induced apoptosis (yes/no) yin; Gain-of-function ced-9 (Bcl-2) No yin; loss-of-function egl-1 (Bim, Bid) Yes yang; gain-of-function ced-9 Yes yang; loss-of-function egl-1 Yes Based on the double mutant analysis, where in the apoptotic pathway are yin and yang?
Answer yin is upstream of ced-9 as a ced-9 gain-of-function mutant prevents apoptosis in the yin mutant worms. yin is downstream or at the same level to egl-1 as a loss-of-function egl-1 mutant is rescued by the yin mutant. yang must be downstream of egl-1 and ced-9 as IR-induced apoptosis is present regardless of the state of egl-1 or ced-9.