Optimal Sum of Pairs Multiple Sequence Alignment
100 likes | 380 Views
Optimal Sum of Pairs Multiple Sequence Alignment. David Kelley. Dynamic Programming Extension. Standard pairwise sequence alignment methods can be extended to handle k strings. But…. Runtime is O(2 k N k ) k = # of sequences N = average length of sequences Space is O(N k )

Optimal Sum of Pairs Multiple Sequence Alignment
E N D
Presentation Transcript
Optimal Sum of Pairs Multiple Sequence Alignment David Kelley
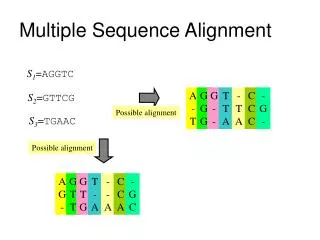
Dynamic Programming Extension • Standard pairwise sequence alignment methods can be extended to handle k strings
But… • Runtime is O(2kNk) • k = # of sequences • N = average length of sequences • Space is O(Nk) • Quickly becomes unfeasible
Enter Carillo-Lipman • Lower bound the score • Estimate distance from cell to end • Calculate sum of all pairwise distances from cell to end • If current score + estimate < lower bound • Ignore that path
MSA • Implemented in 1989 program MSA. • Used a simple progressive alignment procedure to obtain a lower bound • “generally can align 6 to 8 sequences of length 200-300 residues”
Gupta 1995 update • Re-implemented MSA more efficiently • Uses a star-tree heuristic for lower bound • Ran on Sun SparcStation 10 with 128MB of RAM • Runtimes varied (based on similarity of sequences too) • 10 Globin B proteins of ~150 a.a. took 10 min
Can we do better? • Better hardware • more RAM • multi-core processors • Better heuristics • MUSCLE, MAFFT very fast, accurate • Higher lower bound means more of the matrix can be ignored
My Project • Implement concepts from Carillo-Lipman • Use MUSCLE for lower bound • Look for opportunities to parallelize • Using openMP • Run on modern hardware
Can optimal alignment be made practical? • How much better can we do than the previous attempts? • How will maximizing sum of pairs compare to more popular alignment programs? • Compare on multiple sequence alignment database, BAliBase