Cell Phenotype Image Data Resource
Cell Phenotype Image Data Resource. Feasibility Study WP13.1. What is this about?. Cell-based assays with large-scale perturbation libraries and automated microscopy readout. Not: animals tissues pathology anatomy detailed „single-gene“ / mechanistic cell biology

Cell Phenotype Image Data Resource
E N D
Presentation Transcript
Cell Phenotype Image Data Resource Feasibility Study WP13.1
What is this about? Cell-based assays with large-scale perturbation libraries and automated microscopy readout • Not: • animals • tissues • pathology • anatomy • detailed „single-gene“ / mechanistic cell biology • electron microscopy structures • phenotypes in general
Why is it important? Powerful method for systematic and large scale association of genes with biological processes Discovery & systematic mapping of gene-gene, gene-drug interactions Genetic relationships (e.g. similarity, redundancy, epistasis) between genes involved in a phenotype may be of interest beyond the particular instance of phenotype; even across species.
Phenologs Evolutionarily conserved network of genes whose outcome is, in each organism, a certain „phenotype“ human deafness arabidopsis gravitropism Ed Marcotte
Why now? Technological breakthroughs: Genome-wide RNAi libraries Automated microscopyPowerful computers
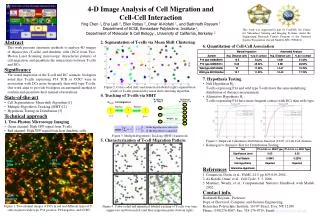
Image analysis of morphology phenotypes induced by RNAi in human cell culture G.Pau (EMBL); F. Fuchs, C. Budjan, Michael Boutros (DKFZ) Genomewide RNAi library (Dharmacon, 22k siRNA-pools) HeLa cells, incubated 48h, then fixed and stained Microscopy readout: DNA (DAPI), tubulin (Alexa), actin (TRITC) CD3EAP
siRNA perturbation phenotypes are observed by automated microscopy wt- wt- wt- BTDBD3 CEP164 CD3EAP 22839 wells DNA, tubulin, actin 4 images per well, each with 3 colours, 1344 x 1024 pixel at 12 bit
Integrated research project on cell cycle control within the 6th Framework of the EU, 8.5 M€, 2004-2008. Live cell time-lapse imaging • Genome-wide siRNA library (1-3 different siRNAs per gene) • HeLa cell line expressing H2B GFP • Seeded on siRNA spots and grown during 48h • Fluorescence time-lapse imaging (sampling rate 30min) Data • 450 chips (including replicates) • 384 spots/chip • ~ 200 000 spots • For each spot, a video sequence of 96 images • raw data: 40 TB
Related projects: A large software project between research-active laboratories at Dundee, NIA Baltimore, HMS and Madison, funded by WT, CR UK, NIH. Server-client software for visualising, managing, and annotating microscope images and metadata, and for working with experimental protocols. Aims at the “LIMS” use case. Contains a wealth of software modules, domain expertise and experience relevant to this pilot study.
Related projects: Image analysis software for biologists without training in computer vision or programming to quantitatively measure phenotypes from thousands of images automatically. Interactive exploration and analysis of data from high-throughput image-based experiments. Supervised machine learning system can be trained to recognize user-defined phenotypes, enabling automatic scoring of millions of cells. Broad Institute (A. Carpenter, T. Jones)
Workshop “Cellular Phenotype Imaging” Co-sponsored by the Wellcome Trust; 17./18. July 2008 in Hinxton. Rebecca Aarons Midori Harris Michael Howell Karol Kozak Emma Lundberg Fernand Meyer Carmel Nanthakumar Wiro Niessen JC Olivo-Marin Jeroen Raes Mihail Sarov Jason Swedlow Stefan Wiemann Thomas Baer Buzz Baum Ewan Birney Andy Blanchard Michael Boutros Alvis Brazma Stephen Bryant Anne Carpenter Jan Ellenberg Daniel Gerlich JK Heriché Wolfgang Huber Andy Lyall Erik Meijering James Mulshine Lucas Pelkmans Rainer Pepperkok Jasmine Zhou Sessions on: Data production Data analysis Biological Integration Practicalities (Standardisation, implementation)
The impact on scientific discovery Data archival for reproducibility of research scientific record avoid duplication Beyond that: „serendipidity“: further exploitation of datasets beyond what the primary authors thought of Aggregation
Aggregation: assembling the cellular phenome What is the total space of cellular phenotypes ? Can it be mapped and structured ? What are the logical relations between phenotypes ? How to infer functional modules from phenotypes? Genetic and gene-drug interactions (managing resistance and side effects) Engineering of phenotypes by combinatorial perturbations Insights beyond the actual observed phenotypes (Ed Marcotte‘s „phenologs“) Attractive prospects for scientists and funders
Two models of operation Genome sequencing Few large centers Fort Lauderdale agreement Data provision by EMBL/Ensembl/NCBI/UCSC Microarrays Distributed Journal requirement / community expectation Data provision by ArrayExpress, GEO; but also ad hoc websites
Results from this WP „White Paper“ from the July 2008 workshop: emphasis on the motivating scientific questions on how to make data contribution attractive to produces Pilot database development at EBI, with 5 different datasets & web front endBased on Ellenberg, Boutros, Huber research projectsExperience in finding common data representation
Cellular Phenotype Imaging Pilot Database Stores images and movies obtained through siRNA induced gene knockdown. Links these images and movies to their phenotypic hits, the siRNA reagents used in the knockdown and the genes these siRNA are claimed to target. Daniel Murrell 5 datasets: 182 412 movies, 191 378 images
CPI Pilot example flow See related phenotypes See associated images / movies Search for a gene of interest. See related genetic sequences See reagents targeting the sequences
Pull up the image data that links CD3EAP with a specific phenotype through reagent M-020021-00.
Conclusion Strong scientific reasons for creating such a database; great enthusiasm in the community (workshop participation!) There is currently no effort with comparable scope; there are relevant projects developing software tools. Technical questions (e.g. amount of data) but none seem insurmountable. Analogies with / lessons from ArrayExpress / marrays, NGS
Questions How generic / prototypical already are the current datasets? marrays: breakthrough was Affymetrix – verified & high quality probe sequences Current RNAi libraries still require a lot of „validation“ (rate-limiting step) ? same urgency as e.g. Genetic variation, Proteomics ?
Questions Ideal location? Hub or node? Role of other types of biological images?
Acknowledgements Daniel Murrell Greg Pau Oleg Sklyar Michael Boutros (DKFZ) Jan Ellenberg (EMBL) Jean Karime Heriché (EMBL) Thomas Walter (EMBL) Anne Carpenter (Broad Inst.) Jeroen Raes (EMBL)
Results from this WP „White Paper“ from the July 2008 workshop Pilot database installation at EBI, with 5 different datasets, web front end; experience in finding common data representation imageHTS software (R/Bioconductor): provides a complete data analysis workflow: from raw microscopy images over cell classification, screen quality assessment to heatmap / phenotype landscape graph
imageHTS R/Bioconductor package provides a complete data analysis workflow: from raw microscopy images over cell classification, screen quality assessment to heatmap / phenotype landscape graph