Recombinant DNA Technology
Recombinant DNA Technology. Recombinant DNA. Protocols that transfer genetic information (DNA) from one organism to another. Gene cloning links eukaryotic genes to small bacterial or phage DNAs and inserting these recombinant molecules into bacterial hosts.

Recombinant DNA Technology
E N D
Presentation Transcript
Recombinant DNA • Protocols that transfer genetic information (DNA) from one organism to another. • Gene cloning links eukaryotic genes to small bacterial or phage DNAs and inserting these recombinant molecules into bacterial hosts. • One can then produce large quantities of these genes in pure form.
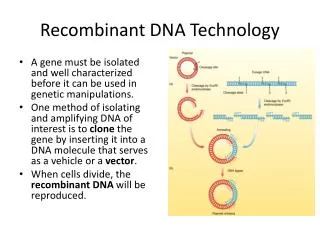
Recombinant DNA - Gene Cloning • DNA from a source organism is cleaved with restriction endonuclease and inserted into a cloning vector. • The recombinant vector is introduced into a host cell. • Recombinant cells are identified and grown.
Five Basic Steps in Gene Cloning • The first is to choose the appropriate DNA to be cloned, genomic or cDNA. • Produce a collection of DNA fragments of size suitable for inserting into appropriate vectors. • Insert DNA fragments into the vector using DNA ligase (DNA ligation.) • Introduce DNA fragments into a population of bacteria (transformation.) • Select the colonies containing desired sequence from the “library.”
Restriction Enzymes • Restriction enzymes (restriction endonucleases) cut double-stranded DNA into smaller pieces. • Bacteria use these as defense against DNA from bacteriophage. • DNA is cut between the 3′ hydroxyl group of one nucleotide and the 5′ phosphate group of the next - restriction digestion.
Restriction Enzymes • Restriction enzymes do not cut bacteria’s own DNA because the recognition sequences are modified. • Methylases add methyl groups after replication; makes sequence unrecognizable by restriction enzyme.
Restriction Enzymes • Bacterial restriction enzymes can be isolated from cells. • DNA from any organism will be cut wherever the recognition site occurs. • EcoRI (from E. coli) cuts DNA at this sequence: GAATTC
Restriction Enzymes The sequence is palindromic—it reads the same in both directions from the 5′ end. EcoRI occurs about once every four genes in prokaryotes. DNA can be chopped into small pieces containing a few genes.
Restriction Enzymes Restriction endonucleases are named using the 1st three letters of their name from the Latin name of their source microorganism Hind III • First letter is from the genus H from Haemophilus • Next two letters are the 1st two letters of the species name in from influenzae • Sometimes the strain designation is included “d” from strain Rd • If microorganism produces only 1 restriction enzyme, end the name with Roman numeral I Hind I • If more than one restriction enzyme is produced, the others are numbered sequentially II, III, IV, etc.
Restriction Enzymes • These enzymes can recognize 4-bp, 6-bp, 8-bp sequences • The frequency of cuts lessens when the recognition sequence is longer • A 6-bp cutter will yield DNA fragments averaging ~ 4000-bp or 4 kilobases (4kb) in length
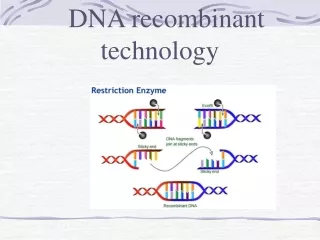
Restriction Enzymes • Many restriction endonucleases make staggered cuts in the 2 DNA strands • This leaves single-stranded overhangs, called sticky ends that can base-pair together briefly • This makes joining 2 different DNA molecules together much easier • Staggered cuts occur when the recognition sequence usually displays twofold symmetry, palindromes
Restriction Enzymes • Heteroschizomers (isochizomers)recognize the same DNA sequence but are from different organisms. • Restriction nuclease that recognize the same DNA sequence but use a different cutting site – they are also calledneoschizomers. • Restriction nuclease that produce the same nucleotide extension are called isocaudomers.
Restriction Enzymes • These enzymes cut DNA strands reproducibly in the same place, which is extremely useful in gene analysis • Isoschizomers cleave a sequence only if the cytosines of the recognition site are not methylated whereas another will cut the same sequence if these cytosines are methylated. • For example, HpaII cuts only nonmethylated CCGG sites, and MspI cuts this sequence regardless of cytosine methylation.
Gel Electrophoresis • After DNA is cut, fragments of different sizes can be separated by gel electrophoresis. • Mixture of fragments is place on a well in a porous gel. An electric field is applied across the gel. Negatively charged DNA fragments move towards positive end. • Smaller fragments move faster than larger ones.
Plasmid Cloning Vectors • Plasmids are small, circular DNA molecules that are maintained as independent extrachromosomal entities. • F plasmids carry information for their own transfer from one cell to another. • R plasmids encode resistant to antibiotics. • Cryptic plasmids have no apparent functional coding genes.
Plasmids can easily incorporate foreign DNA. • Plasmids are readily taken up by bacterial cells. • Plasmids then act as vectors, DNA carriers that move genes from one cell to another. • Each plasmid has a sequence that functions as an origin of DNA replication.
Vectors to Carry DNA Sequences A vector should have four characteristics: • Ability to replicate independently of the host cell • A recognition sequence for a restriction enzyme (cloning site) • One or more selectable/reporter genes • Small size in comparison with host’s chromosomes
Vectors to Carry DNA Sequences Plasmids have all these characteristics. • Plasmids are small, many have only one restriction site. • Genes for antibiotic resistance can be used as reporter genes. • And they have an origin of replication and can replicate independently.
Plasmid Cloning vector pBR322 • pBR322 illustrates cloning methods simply • Resistance for 2 antibiotics • Tetracycline • Ampicillin • Origin of replication between the 2 resistance genes • Only 1 site for several restriction enzymes
Cloning using pBR322 Clone a foreign DNA into the PstI site of pBR322 • Cut the vector to generate the sticky ends • Cut foreign DNA with PstI also – compatible ends • Combine vector and foreign DNA with DNA ligase to seal sticky ends • Now transform the plasmid into E. coli
Cloning using pBR322 If new DNA is inserted at that PstI restriction site, it inactivates the gene for ampicillin resistance. Plasmid then has gene for tetracyclin resistance, but not for ampicillin. This can be used to select for host cells with new DNA.
Cloning using pBR322 The cleaved plasmid DNA preparation is treated with enzyme alkaline phosphatase to remove the 5’ phosphate groups from the linearized plasmid DNA.
Transformation and selection • Traditional method involves incubating bacterial cells in concentrated calcium salt solution • The solution makes the cell membrane leaky, permeable to the plasmid DNA – competent cells. • Newer method uses high voltage to drive the DNA into the cells in process called electroporation
Transformation and selection • A natural transformation process often entails • The binding of double-stranded DNA to component of the cell wall. • Entry of the DNA into an inner periplasm. • Transmission of one strand into the cytoplasm while the other one is degraded. • If the DNA is linear molecule, integration into host chromosome. • If the introduced DNA is a plasmid, it is maintained in the cytoplasm after the second strand is synthesized.
Transformation and selection • Transformation produces bacteria with: • Religated plasmid • Religated insert • Recombinants plasmid • Identify the recombinants using the antibiotic resistance • Grow cells with tetracycline so only cells with plasmid grow, not foreign DNA only (religated insert) • Next, grow copies of the original colonies with ampicillin which kills cells with plasmid including foreign DNA (Figure 3.11)
Clone a foreign DNA into theBamHI site • Cells contain no plasmid are sensitive to both Amp and Tet. • Cells contains intact plasmids are resistant to both. • Cells contains inserted plasmids are resistant to Tet but sensitive to Amp.
Screening with replica plating • Replica plating transfers clone copies from original tetracycline plate to a plate containing ampicillin • A sterile velvet transfer tool can be used to transfer copies of the original colonies • Desired colonies are those that do NOT grow on the new ampicillin plate
pUC and β - galactosidase Newer pUC plasmids have: • Ampicillin resistance gene • Multiple cloning site inserted into the gene lacZ’ coding for the enzyme β-galactosidase • Clones with foreign DNA in the MCS disrupt the ability of the cells to make β-galactosidase • Plate on media with a β-galactosidase indicator (X-gal) and clones with intact β-galactosidase enzyme will produce blue colonies • Colorless (desirable) colonies should contain the plasmid with foreign DNA
Directional cloning • Cut a plasmid with 2 restriction enzymes from the MCS • Clone in a piece of foreign DNA with 1 sticky end recognizing each enzyme • The insert DNA is placed into the vector in only 1 orientation • Vector religation is also prevented as the two restriction sites are incompatible