Recombinant DNA Technology
Recombinant DNA Technology. Prof. Elena A. Carrasquillo Chapter 4 Molecular Biotechnology Lecture 4. Cloning in Plasmid Vector In bacteriophage vector Screening the library DNA probe Antibody probe. Major Steps in building a DNA Library.

Recombinant DNA Technology
E N D
Presentation Transcript
Recombinant DNA Technology Prof. Elena A. Carrasquillo Chapter 4 Molecular Biotechnology Lecture 4
Cloning • in Plasmid Vector • In bacteriophage vector • Screening the library • DNA probe • Antibody probe Major Steps in building a DNA Library
Genomic Library: Stores a representation of the genome (at least one copy of a gene present) • plasmid, bacteriophage, phagemid, • cosmid vectors • cDNA Library: Stores a representation of the mRNAs expressed at a certain time or stage by a microorganism or organism • plasmid, bacteriophage, phagemid, • cosmid vectors Kinds of Libraries
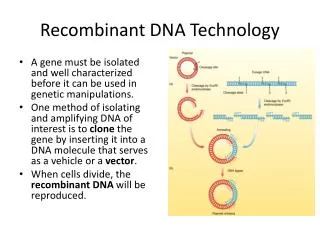
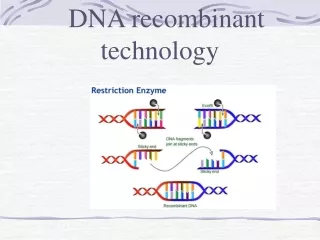
Restriction Enzyme Mechanisms: • Preparation of DNAs to be joined • (a)Staggered cut: leaves “sticky ends” How is a Library Built:
Restriction Enzyme Mechanisms: Preparation of DNAs to be joined: • (b) Blunt End How is a Library Built:
Staggered “sticky ends” Ligation of DNA cut with a Restriction Enzyme
Role of T4 DNA Ligase Ligation of DNA cut with a Restriction Enzyme
Depends on the size of DNA to be cloned • Is the protein encoded by the DNA going to be expressed in a prokariotic or eukaryotic cell? Choosing the Vector
Allows ordering of DNA fragments Restriction Enzyme Map
We can then build maps of linear and circular molecules Restriction Enzyme Map
a multiple cloning site or MCS an antibiotic resistance gene an origin of replication pBR322 is the basis of most engineered plasmids Plasmid: it’s a circular DNA molecule containing:
F plasmids-transfer information from cell to cell • R plasmids-confer antibiotic resistance • Degradative plasmids-utilization of unusual metabolites • Cryptic plasmids-No apparent function Kinds of plasmids in the wild:
Size range form less than 1 kb to more than 500 kb • Origin of replication-allows plasmid to replicate in the bacteria • Low-copy number: 1-4 per cell • High copy number: 10-100 per cell • Incompatibility groups: different kinds cannot be inside the same cell Other characteristics:
Small size (<15Kb) for optimal efficiency of transformation in bacteria • Unique restriction enzime sites for cloning • One or more selectable genetic markers to allow for differentiation of the plasmids carrying the cloned DNA vs the religated ones. Characteristics of an engineered plasmid:
Characteristics: • Interruption of b-lactamase gene gives rise to white colonies (cloned DNA) vs blue ones (empty) pUC19 another plasmid cloning vector
-Produced by site-directed mutagenesis to alter the DNA sequence but not the protein sequence of b-lactamase -This produced new restriction enzyme sites MCS: multiple cloning site of pUC19:
Creation of a DNA Library: Partial DNA digestion Purpose: To produce overlapping DNA fragments
Vary time of digestion or amount of enzyme units Progress of Reaction: agarose gel electrophoresis
Screening a library:I. Colony hybridization or Southern Blot
Method 1 • Random primers • Enzyme: Klenow Fragment + dNTPs • Non-radioactive: biotin or chemiluminescent labeled dNTPs -Radioactive:32P Preparation of DNA Probe
Method 2: 5’-end labeling • Method 3: 3’end labeling Preparation of DNA Probe
The polymerase (red) adds deoxyribonucleotides to the 3’- hydroxyl groups of the growing chains -The 5’ exonuclease (blue) removes succesive nucleotides from the 5’ phosphate ends -The 3’ exonuclease (yellow) removes succesive nucleotides from the3’ hydroxyl ends Klenow Fragment of E. coli DNA Polymerase I Enzyme Activities
Colony Hybridization Screening a Genomic Library
Procedure: Colony Hybridization:
The probe is an antibody The colonies are grown so that the recombinant protein is expressed Screening a Genomic Library: II.Colony Immunoassay
If chemiluminescense was used, light will be emitted and captured in an X-ray film Screening a Genomic Library: II.Colony Immunoassay
Defective host cells (A-) are transformed with a genomic library derived from cells that are normal with respect to that function (A+) and grown in minimal media. • Those cells harboring a plasmid that corrects the defect will grow in minimal media. Screening a Genomic Library:III.Functional Complementation
Used to obtain functional eukaryotic coding regions. • E. coli does not process introns. • First step: Isolate poly A+ mRNA with oligo (dT) cellulose Another Type of Library:cDNA Library
Second Step:Synthesis of cDNA from mRNA of specific cells cDNA Library:
Third Step:Selecting and Cloning Full length cDNA molecules cDNA Library:
For cloning inserts of 10-20 Kb • Plasmid libraries hold up to 10 kb inserts Genomic Library: Bacteriophageλ (Lambda)
Lytic Cycle:Production of progeny • Lysogenic Cycle: Integration into bacterial chromosome Genomic Library: Bacteriophageλ (Lambda)Life Cycle
Allows cloning of 45 kb DNA fragments Cosmid library
Based on P1 bacteriophage, the F plasmid and the lacZ region of pUC plasmids • It’s a low copy number plasmid • Carries 50-300kb fragments BACs: Bacterial Artificial Chromosomes
Transformation • Electroporation • Conjugation Methods of Introducing Foreign DNA
Application of an electrical field to cells Electroporation