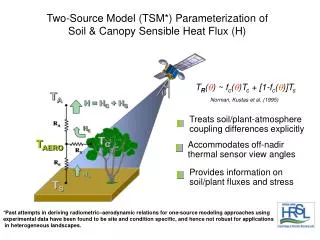
Agrawal et al, Mining sequential patterns, Data Eng., 1995
paper seminar on R. Agrawal and R. Srikant, “Mining Sequential Patterns,” Proc.11th Int’l Conf. Data Eng., pp. 3-14, Mar. 1995.

Agrawal et al, Mining sequential patterns, Data Eng., 1995
E N D
Presentation Transcript
Mining Sequential Patterns Rakesh Agrawal, Ramakrishana Srikant (1995) IBM Almaden Research Center 650 Harry Road, San Jose, CA 95120 데이터 연구실이세린 지도 교수 박종수 2014. 3. 28
Contents • Abstract • Introduction • Finding Sequential Patterns • The Sequence Phase • Performance • Conclusions and Future Work
Abstract • Introduces the problem of mining sequential patterns over a large database. • Presents 3 algorithms to solve this problem. • Shows their results of performance and scale-up experiments.
1. Introduction • 1.1 Problem Statement • The problem of mining sequential patterns is to find the maximal sequences among all sequences that have a certain user-specified minimum support. • Each such maximal sequence represents a sequential pattern.
1. Introduction • 1.2 Related Work • R. Agrawal, T. Imielinski, and A. Swami, “Mining association rules between sets of items in large databases” (1993) • T. G. Dietterich and R. S. Michalski, “Discovering patterns in sequences of events, Artificial Intelligence” (1985) • A. Califano and I. Rigoutsos, “Flash: A fast look-up algorithm for string homology” (1993) • S. Wu and U. Manber, “Fast text searching allowing errors” (1992) • M. Waterman, “Mathematical Methods for DNA Sequence Analysis” (1989) • S. Altschul, W. Gish, W. Miller, E. Myers, and D. Lipman, “A basic local alignment search tool” (1990) • M. Roytberg, “Computer Applications in the Biosciences: A search for common patterns in many sequences” (1992) • M. Vingron and P. Argos, “Computer Applications in the Biosciences: A fast and sensitive multiple sequence alignment algorithm” (1992) • J. T.-L. Wang, G.-W. Chrin, T. G. Marr, B. Shapiro, D. Shasha, and K. Zhang. “Combinatorial pattern discovery for scientific data: Some preliminary results” (1994)
1. Introduction • 1.2 Related Work • Finding of items bought together in a transaction. (Intra-transaction patterns) • AI prediction of the sequential pattern. • Finding matches for pattern in text subsequences. • Discovering similarities in a database of genetic sequences. Comparison
1. Introduction • 1.2 Related Work
1. Introduction • 1.3 Organization of the Paper • Section 2. Gives this problem decomposition. • Section 3. Examines the sequence phase in detail and presents algorithms for this phase. • Section 4. Empirically evaluate the performance of these algorithms and study their scale-up properties. • Section 5. conclusion - summary and directions for future work.
2. Finding Sequential Patterns • 2.1 The Algorithm • 1. Sort Phase • Converts the original transaction database into a database of customer sequences.
2. Finding Sequential Patterns • 2.1 The Algorithm • 2. Litemset Phase • Find the set of all litemsets L including the set of all 1-sequences. • The set of litemsets is mapped to a set of contiguous integers.
2. Finding Sequential Patterns • 2.1 The Algorithm • 3. Transformation Phase • To process repetitive determination in the following step faster, • Each transaction is replaced by the set of all litemsets contained in that transaction.
2. Finding Sequential Patterns • 2.1 The Algorithm • 4. Sequence Phase • Use the set of litemsets to find the desired sequences. • 5. Maximal Phase • Find the maximal sequences among the set of large sequences.
3. The Sequence Phase • Make multiple passes over the data to generate candidate sequences from seed set of large sequences. 25% (Support > 1.25)
3. The Sequence Phase • 2 Families of algorithms Count-all AprioriAll Count-some AprioriSome DynamicSome
3. The Sequence Phase ① • 3.1 Algorithm AprioriAll ② ③
3. The Sequence Phase • 3.1 Algorithm AprioriAll • 3.1.1 Apriori Candidate Generation p q Join
3. The Sequence Phase • 3.2 Algorithm AprioriSome
3. The Sequence Phase • 3.2 Algorithm AprioriSome • In the forward pass, we only count sequences of certain lengths. Forward phase length6 length2 length3 length4 length5 length1 Backward phase • Forward phase procedure pruning pruning …
3. The Sequence Phase • 3.2 Algorithm AprioriSome • Next() takes as parameter the length of sequences counted in the last pass and returns the length of sequences to be counted in the next pass.
3. The Sequence Phase • 3.3 Algorithm DynamicSome * Backward phase is same as AprioriSome. If step = 3, After initialization of 1, 2, 3, Generate 6, 9, 12 … Has to be initialized
3. The Sequence Phase • 3.3 Algorithm DynamicSome … AprioriAll … AproriSome … DynamicSome
3. The Sequence Phase • 3.3 Algorithm DynamicSome otf-generates (On-the-fly) generates more candidates than apriori-generate. Avoid overlapping
3 Algorithm Example • AprioriAll / AprioriSome / DynamicSome ①
3 Algorithm Example • AprioriAll ②
3 Algorithm Example • AprioriSome ②
3 Algorithm Example • AprioriSome ③
3 Algorithm Example • DynamicSome ② (step = 2)
3 Algorithm Example • DynamicSome ③
4. Performance • 4.1 Generation of Synthetic Data • Customer-sequence sizes are typically clustered around a mean and a few customers may have many transactions. • Transactionsizes are usually clustered around a mean and a few transactions have many items. • Setting: = 5,000 = 25,000N = 10,000
4. Performance • 4.2 Relative Performance • Decreased support by 1% to 0.2%.
4. Performance • 4.2 Relative Performance • Observation: • Execution time support • DynamicSome performs worse. • AprioriSome shows:
4. Performance • 4.3 Scale-up • Scale-up experiments for the AprioriSome algorithm. (AprioriSome and AprioriAll results to be very similar.)
4. Performance • 4.3 Scale-up
5. Conclusions and Future Work • Introduced a new problem of mining sequential patterns from a database of customer sales transactions. • Presented 3 algorithms for solving this problem. • AprioriSome and AprioriAll have comparable performance. • AprioriSome performs a little better for the lower values of the minimum number of customers that must support a sequential pattern. • Both scale linearly with the number of customer transactions. • Both have excellent scale-up properties with respect to the number of transactions in a customer sequence and the number of items in a transaction. • AprioriAll is preferred in some cases that need detail counts of the number of people.
5. Conclusions and Future Work In the future, • Extension of the algorithms to discover sequential patterns across item categories. • Transposition of constraints into the discovery algorithms. There could be item constraints or time constraints.