Graph Theory Aiding DNA Fragment Assembly
150 likes | 594 Views
Jonathan Kaptcianos e-mail: jkaptcianos@smcvt.edu advisor: Professor Jo Ellis-Monaghan. Graph Theory Aiding DNA Fragment Assembly. Work supported by the Vermont Genetics Network through NIH Grant Number P20 RR16462 from the INBR program of the National Center for Research Resources.

Graph Theory Aiding DNA Fragment Assembly
E N D
Presentation Transcript
Jonathan Kaptcianos e-mail: jkaptcianos@smcvt.edu advisor: Professor Jo Ellis-Monaghan Graph Theory Aiding DNA Fragment Assembly Work supported by the Vermont Genetics Network through NIH Grant Number P20 RR16462 from the INBR program of the National Center for Research Resources
DNA Sequencing: An Overview • a lab technique which looks at fragments (anywhere from 500 to 1200 nucleotides long) of DNA and determines the order of entire genome from these these individual fragments. • modern science has enabled us to determine the DNA sequences of animals and other organisms Previous approaches for fragment assembly follow the “overlap-layout consensus” algorithm • overlap: matching all possible reads and finding any overlapping • layout: finding order of reads along DNA and putting them together • consensus: deriving how sequence will appear based on layout
Problems in DNA Sequencing • there could be multiple ways to reconstruct the original strand out of the fragment pieces, or “snippets,” and only one of which is correct • the human genome has a large number of sequences that repeat an even larger number of times • if a repeating sequence is larger than the size of the viewable reads, it would make construction of the genome almost impossible Solutions: Some components in Graph Theory, specifically Eulerian Paths and de Bruijn Graphs, help us come to some possible conclusions about the problem regarding reassembled strands of DNA
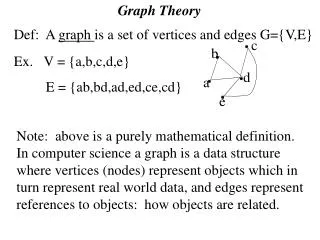
d e a a c b g f c h h b d f i j e Eulerian Circuits and Paths Eulerian Circuit – visits each edge in a graph exactly once, and ends at the same vertex in which it started. a-d-b-f-e-d-f-c-b-a is an Eulerian cycle in this particular graph Eulerian Path – visits each edge in a graph exactly once. a-b-c-d-e-f-g-c-h-f-i-j is an Eulerian trail in this particular graph
GAA ACT ATC GAC ATA TAT CTA TCG TAA CGA AAG CAT AGG GGC GGC GCA DNA Strands and de Bruijn Graphs de Bruijn Graph – a directed graph with vertices that represent sequences of symbols from an alphabet, and edges that indicate where the sequence may overlap. Example: The strand ATCGACTATAAGGCATCGAA de Bruijn graph has “snippets” of length 4, vertices of length 3, and the directed edge between two vertices represent the 4 piece snippet. S 2007
Eulerian Path Approach to DNA Fragment Assembly • abandons the previously mentioned “overlap-layout- consensus” • ultimately, converts an NP-complete Hamilton Path Problem into a simplified Eulerian Path Problem through construction of a de Bruijn graph • the number of ways to reconstruct the graph is equivalent to the number of paths which follow the respective directions and travel through all edges • the resulting problem is that there are a number of different Eulerian Paths through this graph, and we cannot tell which would resemble the original path E-M 2006
Eulerian Superpath Problem • Eulerian Superpath Problem – Given an Eulerian Graph and a collection of paths on this graph, find an Eulerian path in this graph that contains all these paths as subpaths. • The original Eulerian Path Problem is a case of the Eulerian Superpath Problem, in which every path is a single edge. Solving: • Take graph G and the system of paths P, and transform these to a new graph G1 and a new system P1. • With the goal in mind that there is a one-to-one correspondence (equivalence) between (G,P) and (G1,P1), we go on to make a series of these transformations. (G,P) → (G1,P1) → (G2,P2) →…→ (Gk,Pk) • All these transformations should lead to a system Pk in which every path is represented by one edge. • Since all transformations from beginning to end are equal, every solution of EPP in (Gk,Pk) will provide a solution to the ESPP in (G,P).
An x,y-detachment for no multiple edges • Let x = (vin,vmid) and y = (vmid,vout) be two consecutive edges in G and Px,y be all paths from P that include x,y as a subpath. • P→x is the paths from P that end on x and Py→ is the collection of paths from P that start with y. • Adding a new edge z = (vin,vout) to delete the edges x and y. • We can substitute z instead of x,y in all paths from Px,y, x in all paths from P→x, and y in all paths from Py→. • Thus, reducing an ESPP to an EPP. PTW 2001
Detachment for Multiple Edges • Let vertex vmid have multiplicity 2 and only incoming edge bex = (vin,vmid), and two outgoing edges y1 = (vmid,vout1) and y2 = (vmid,vout2) with multiplicity 1. • Since there exists a multiple edge, the Eulerian path will visit x twice, once followed by y1 and once by y2. • If an edge zis used in a detachment of x,y1 , it will shorten Px,y1to a single edge z and substitute z in all paths from Py1→ . • Equivalence will only be present if P→xis empty; if its not, there will be ambiguity about whether the last edge in a specified path Pin P→x should go to z or the remaining edge x. • This is resolved by looking at the relations between every path P and Px,y1 or Px,y2. PTW 2001
Paths and Consistency • Two paths are consistent if their union is a path and there • are no branching vertices. • Case 1:P is inconsistent with both Px,y1 and Px,y2 • In this situation, there exists no solution to the Eulerian Superpath Problem, as the data for sequencing will be inconsistent. • In the example below, the three paths possess a different way to visit edge x PTW 2001
Case 2:P is consistent with only one of Px,y1 and Px,y2 • Pis resolvable, as it can be related to one of the systems of paths. • When consistent with Px,y1, it would be assigned to the z edge created in the previous x,y1-detachment • When consistent with Px,y2, it would be assigned to edge x and no further action would be needed • The edge x is resolvable if all paths in P→x are, and therefore it is an equivalent transformation. x PTW 2001 Here, P is consistent with Px,y1
Case 3:P is consistent with both Px,y1 and Px,y2 • When this occurs on at least one path in P→x, the edge x is considered unresolvable and is postponed with the hopes of further transformations (shown below) resolving it y4,x1-detachment x2,y1-detachment z,x2-detachment PTW 2001 • Through this series of transformations, the final graph is a simplified and equivalent transformation of the first.
The x-cut • Consider the graph G with 5 edges and the 4 given paths with two edges each. • In this situation, no previous detachment discussed will allow for an equivalent transformation. • An edge x=(v,w) is removable if it is the only edge leaving v and coming into w, and if it is either the initial or final edge in every path P in the system of paths • An x-cut on this graph will turn P into a new system of paths by removing x from all paths in P→x and Px→. • As x is removed from each path, the single-edged paths y1,y2,y3,y4 that remain. • This demonstrates an equal transformation as each Eulerian Superpath in (G,P) corresponds to each in (G1,P1) PTW 2001
Some Conclusions • Scientists and researchers are able to consider large groups of edges, vertices, and paths as a significantly smaller number elements, instead of having to focus on every element in the strand of DNA. Through a series of detachments and cuts, it is possible to transform a once tangled and overwhelming graph into a simplified, equivalent and more easily resolvable graph. The Eulerian Superpath Approach on DNA Fragment Assembly doesn’t eliminate the discrepancies about the original construction of the Genome, but just makes it a little neater and easier to work with.