Frontiers Between Crystal Structure Prediction and Determination by Powder Diffractometry
760 likes | 927 Views
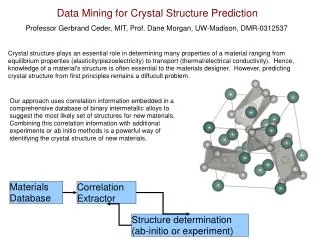
Frontiers Between Crystal Structure Prediction and Determination by Powder Diffractometry Armel Le Bail Université du Maine, Laboratoire des Oxydes et Fluorures, CNRS UMR 6010, Avenue O. Messiaen, 72085 Le Mans, France Email : alb@cristal.org. Outline

Frontiers Between Crystal Structure Prediction and Determination by Powder Diffractometry
E N D
Presentation Transcript
Frontiers Between Crystal Structure Prediction and Determination by Powder Diffractometry Armel Le Bail Université du Maine, Laboratoire des Oxydes et Fluorures, CNRS UMR 6010, Avenue O. Messiaen, 72085 Le Mans, FranceEmail : alb@cristal.org
Outline • Introduction- Prediction software and examples- Fuzzy frontiers with SDPD- More examples from the GRINSP software - Opened doors, limitations, problems- « Immediate structure solution » by search-match- Conclusion- Live demo with EVA-Bruker + PPDF-1
INTRODUCTION Personnal views about crystal structure prediction : “Exact” description before synthesis or discovery in nature. These “exact” descriptions should be used for the calculation of powder patterns included in a database for automatic identification of actual compounds not yet characterized crystallographycally.
Where are we with inorganic crystal structure prediction? If the state of the art had dramatically evolved in the past ten years, we should have huge databases of predicted compounds, and not any new crystal structure would surprise us since it would corespond already to an entry in that database. Moreover, we would have obtained in advance the physical properties and we would have preferably synthesized those interesting compounds. Of course, this is absolutely not the case.
But things are changing, maybe : Two databases of hypothetical compounds were built in 2004. One is exclusively devoted to zeolites : M.D. Foster & M.M.J. Treacy - Hypothetical Zeolites – http://www.hypotheticalzeolites.net/ The other includes zeolites as well as other predicted oxides (phosphates, sulfates, silicates, borosilicates, etc) and fluorides :the PCOD (Predicted Crystallography Open Database)http://www.crystallography.net/pcod/
Prediction software Especially recommended lectures (review papers) : 1- S.M. Woodley, in: Application of Evolutionary Computation in Chemistry, R. L. Johnston (ed), Structure and bonding series, Springer-Verlag 110 (2004) 95-132. 2- J.C. Schön & M. Jansen, Z. Krist.216 (2001) 307-325; 361-383. Software : CASTEP, program for Zeolites, GULP, G42, Spuds, AASBU, GRINSP
CASTEP Uses the density functional theory (DFT) for ab initio modeling, applying a pseudopotential plane-wave code. M.C Payne et al., Rev. Mod. Phys. 64 (1992) 1045. Example : carbon polymorphs
ZEOLITES The structures gathered in the database of hypothetical zeolites are produced from a 64-processor computer cluster grinding away non-stop, generating graphs and annealing them, the selected frameworks being then re-optimized using the General Utility Lattice Program (GULP, written by Julian Gale) using atomic potentials. M.D. Foster & M.M.J. Treacy - Hypothetical Zeolites – http://www.hypotheticalzeolites.net/
Zeolite predictions are probably too much… Less than 200 zeotypes are known Less than 10 new zeotypes are discovered every year Less than half of them are listed in that >1.000.000 database So that zeolite predictions will continue up to attain several millions more… Quantum chemistry validation of these prediction is required, not only empirical energy calculations, for elimination of a large number of models that will certainly never be confirmed.
GULP (at the Frontier ?) Appears to be able to predict crystal structures (one can find in the manual the data for the prediction of TiO2 polymorphs). Recently, a genetic algorithm was implemented in GULP in order to generate crystal framework structures from the knowledge of only the unit cell dimensions and constituent atoms (so, this is not full prediction...), the structures of the better candidates produced are relaxed by minimizing the lattice energy, which is based on the Born model of a solid. S.M. Woodley, in: Application of Evolutionary Computation in Chemistry, R. L. Johnston (ed), Structure and bonding series, Springer-Verlag 110 (2004) 95-132. GULP : J. D. Gale, J. Chem. Soc., Faraday Trans.,93 (1997) 629-637. http://gulp.curtin.edu.au/
G42 A concept of 'energy landscape' of chemical systems is used by Schön and Jansen for structure prediction with their program named G42. J.C. Schön & M. Jansen, Z. Krist.216 (2001) 307-325; 361-383.
SPuDS Dedicated especially to the prediction of perovskites. M.W. Lufaso & P.M. Woodward, Acta Cryst. B57 (2001) 725-738.
AASBU method (Automated Assembly of Secondary Building Units) Developed by Mellot-Draznieks et al., C. Mellot-Drazniek, J.M. Newsam, A.M. Gorman, C.M. Freeman & G. Férey, Angew. Chem. Int. Ed. 39 (2000) 2270-2275; C. Mellot-Drazniek, S. Girard, G. Férey, C. Schön, Z. Cancarevic, M. Jansen, Chem. Eur. J. 8 (2002) 4103-4113. Using Cerius2 and GULP in a sequence of simulated annealing plus minimization steps for the aggregation of large structural motifs. Cerius2, Version 4.2, Molecular Simulations Inc., Cambridge, UK, 2000.
Two (incredible ?) predictions • « Giant structures solved by • combined targeted chemistry and computational design."2 cubic hybrid solids structures published : • - 2004 - V ~380.000 Å3, a ~ 73Å, Fd-3m, 68 indpdt atoms (not-H) • 2005 - V ~702.000 Å3, a ~ 89Å, Fd-3m, 74 indpdt atoms (not-H) • Presented more or less as being predicted, but with indexed powder patterns and guessed content. • Are they SDPD or predictions ?The fuzzy frontier is there…
Super-tetrahedra sharing corners, building super-zeolites (MTN-analogue)
Same « prediction » process, building another MTN-analogue super-zeolite From different super-tetrahedra
Will you be able to equal or surpass these giant structure « predictions » ? YESIf the molecule, the cell and the space group are known, then the direct space methods need only 50 or 100 reflections for solving the structure, whatever the cell volume (6 DoF per molecule rotated and translated). But this is not prediction. MAYBEBy partial prediction (without cell but with known content).This is « molecular packing prediction ». NOWithout cell and without content, full total prediction at such complexity level looks impossible. Anyway, you may try to impress some Nature or Science reviewer searching for « sensational » results, by your eloquence.
Not enough full predictions If zeolites are excluded, the productions of these prediction software are a few dozen… not enough, and not available in any database. The recent (2005) prediction program GRINSP is able to extendthe investigations to larger series of inorganic compounds characterized by corner-sharing polyhedra.
GRINSP Geometrically Restrained INorganic Structure Prediction Applies the knowledge about the geometrical characteristics of a particular group of inorganic crystal structures (N-connected 3D networks with N = 3, 4, 5, 6, for one or two N values). Explores that limited and special space (exclusive corner-sharing polyhedra) by a Monte Carlo approach. The cost function is very basic, depending on weighted differences between ideal and calculated interatomic distances for first neighbours M-X, X-X and M-M for binary MaXb or ternary MaM'bXc compounds. J. Appl. Cryst. 38, 2005, 389-395. J. Solid State Chem. 179, 2006, 3159-3166.
Observed and predicted cell parameters comparison Predicted by GRINSP (Å) Observed or idealized (Å) Dense SiO2 a b c R a b c (%) Quartz 4.965 4.965 5.375 0.0009 4.912 4.912 5.404 0.9Tridymite 5.073 5.073 8.400 0.0045 5.052 5.052 8.270 0.8Cristobalite 5.024 5.024 6.796 0.0018 4.969 4.969 6.926 1.4 Zeolites ABW 9.872 5.229 8.733 0.0056 9.9 5.3 8.8 0.8EAB 13.158 13.158 15.034 0.0037 13.2 13.2 15.00.3EDI 6.919 6.919 6.407 0.0047 6.926 6.926 6.4100.1GIS 9.772 9.772 10.174 0.0027 9.8 9.8 10.20.3GME 13.609 13.609 9.931 0.0031 13.7 13.7 9.9 0.6Aluminum fluorides-AlF3 10.216 10.216 7.241 0.0159 10.184 10.184 7.174 0.5Na4Ca4Al7F33 10.876 10.876 10.876 0.0122 10.781 10.781 10.7810.9AlF3-pyrochl. 9.668 9.668 9.668 0.0047 9.749 9.749 9.749 0.8 TitanosilicatesBatisite 10.633 14.005 7.730 0.0076 10.4 13.85 8.1 2.6Pabstite 6.724 6.724 9.783 0.0052 6.7037 6.7037 9.824 0.9Penkvilskite 8.890 8.426 7.469 0.0076 8.956 8.727 7.387 1.3
Predictions produced by GRINSP Binary compounds Formulations M2X3, MX2, M2X5 et MX3 were examined. Zeolites MX2 (= 4-connected 3D nets) More than 4700 zeolites (not 1.000.000) are proposed with cell parameters < 16 Å, placed into the PCOD database :http://www.crystallography.net/pcod/ GRINSP recognizes a zeotype by comparing the coordination sequences (CS) of a model with a previously established list of CS and with the CS of the models already proposed during the current calculation).
Hypothetical zeolite PCOD1010026SG : P432, a = 14.623 Å, FD = 11.51
Other GRINSP predictions : > 3000 B2O3 polymorphs Hypothetical B2O3 - PCOD1062004.Triangles BO3 sharing corners.= 3-connected 3D nets
> 1300 V2O5 polymorphs square-based pyramids = 5-connected 3D nets
>30 AlF3 polymorphs Corner-sharing octahedra.= 6-connected 3D nets
Do these AlF3 polymorphs can really exist ? Ab initio energy calculations by WIEN2K « Full Potential (Linearized) Augmented Plane Wave code » A. Le Bail & F. Calvayrac, J. Solid State Chem. 179 (2006) 3159-3166.
Ternary compounds MaM’bXc in 3D networks of polyhedra connected by corners Either M/M’ with same coordination but different ionic radii or with different coordinations (mixed N-N’-connected 3D frameworks) These ternary compounds are not always electrically neutral.
Borosilicates PCOD2050102, Si5B2O13, R = 0.0055. SiO4tetrahedraandBO3triangles > 3000 models
Aluminoborates Example : [AlB4O9]-2, cubic, SG : Pn-3, a = 15.31 Å, R = 0.0051: AlO6octahedra andBO3triangles > 4000 models
Fluoroaluminates Known Na4Ca4Al7F33 : PCOD1000015 - [Ca4Al7F33]4-. Two-sizesoctahedra AlF6and CaF6
Results for titanosilicates TiO6octahedra andSiO4 tetrahedra > 1700 models
More than 70% of the predicted titanosilicates have the general formula [TiSinO(3+2n)]2- Numbers of compounds in ICSD version 1-4-1, 2005-2 (89369 entries) potentially fitting structurally with the [TiSinO(3+2n)]2- series of GRINSP predictions, adding either C, C2 or CD cations for electrical neutrality. n +C +C2 +CD Total GRINSP ABX5 1 300 495 464 35 1294 130 TiSiO5AB2X7 2 215 308 236 11 770 207 TiSi2O7AB3X9 3 119 60 199 5 383 215 TiSi3O9AB4X11 4 30 1 40 1 72 257 TiSi4O11AB5X13 5 9 1 1 0 11 75 TiSi5O13AB6X15 6 27 1 13 1 42 207 TiSi6O15Total 2581 1091 Not all these 2581 ICSD structures are built up from corner sharing octahedra and tetrahedra. Many isostructural compounds inside.