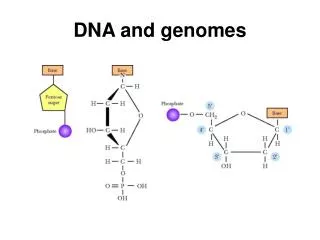
Mapping and Sequencing Genomes
Mapping and Sequencing Genomes. Sanger Sequencing. Sanger Sequencing. Sanger Sequencing-Critical Innovations. Lee Hood (1986) Radiolabelled ddNTPs Fluorescently Labelled ddNTPs eliminate radioactivity, multiplex reactions 2) Molly Craxton (1991)

Mapping and Sequencing Genomes
E N D
Presentation Transcript
Sanger Sequencing-Critical Innovations • Lee Hood (1986) • Radiolabelled ddNTPsFluorescently Labelled ddNTPs • eliminate radioactivity, multiplex reactions • 2) Molly Craxton (1991) • thermostable polymerases, cycle-sequencing • significantly reduced amount of template required, 96-well format • Michael Hunkapiller, Lee Hood, Applied Biosystems (circa 1991) • slab gelscapillary gel electrophoresis • much easier/better automation, lane tracking, high speed runs, better resolution, 3 x 96 well/24 hours 12 x 96 well/24 hours
Combined Approach Shotgun clone BAC Library/BAC end sequencing Shotgun sequence Physical Mapping BAC shotgun sequencing
Calculating Sequence Coverage Given a clone/BAC/Genome of a given size, how do I figure out how many sequencing reads to run? What coverage? How many gaps? How large are gaps?
Lander-Waterman Model Lander ES, Waterman MS (1988) Genomic mapping by fingerprinting random clones: a mathematical analysis“ Genomics 2 (3): 231- 239 • Poisson Estimate • Number of reads • Average length of a read
Poisson is a good estimate for… Helpful to think of Poisson as having to do with rates • Number of cars that pass through a toll plaza in a given hour • Number of misprints per page in a book • Number of alpha particles emitted in one second from a radioactive substance • Number of trout in a cubic meter of pond water
Poisson Distribution If for a given sequence alignment I observe, on average, 3 mis-matches every 50 bp, what is the chance of observing a 50bp window with 5 mis-matches? =3
Poisson Distribution What is the chance of observing at least one mismatch in a 50 bp window? =3
Lander–Waterman Assumptions • Sequencing reads will be randomly distributed in the genome • 2. The ability to detect an overlap between two truly overlapping reads does not vary from clone to clone
Lander–Waterman Assumptions • Sequencing reads will be randomly distributed in the genome • 2. The ability to detect an overlap between two truly overlapping reads does not vary from clone to clone
In practice… Lander-Waterman is almost always an underestimate -cloning biases in shotgun libraries -repeats -GC/AT rich regions -other low complexity regions
Mapping/Ordering BACs What is a marker? A way to uniquely locate a position in a genome What is mapping? Statistical association between markers, ordering markers in linear sequence. How do we map? “Shatter” genome and observe how often two markers travel together on the same piece of DNA What does it mean for two markers to be linked? P(M1|M2)>P(M1) What does it mean to order BACs? Create a minimal tiling path.
Marker every fifth lane BAC fingerprint gel 96 samples, 25 marker lanes 29,950 bp 1% agarose; 8 hours, 140 volts @ 14°C Marra et al., Genome Res., 7, 1072-1084 (1997) 560 bp
Whole genome map assembly Hybridize markers or identify in BAC end sequence (e-PCR).
Whole genome map assembly Edit contigs and align to map.
Verification of location by FISH • BACs were directly labeled with fluorescein conjugated d-UTP by random priming. Labeled DNA was competitively hybridized to cytogenetically normal male human metaphase chromosomes.
When is a genome finished? 1) Finishing is hard! 2) Quality values: Phred score = -10*log10P(error) Phred20=1error/100bp How much continuous phred20 sequence? 3) Gaps? 1 contig/chromosome (probably not)
EST Projects • EST=Expressed Sequence Tag • Short, single pass reads from bits of mRNA • In practice random reads from cDNA libraries • polyA primed/random primed • Sometimes libraries are tissue specific
ESTs • Downs: • Libraries are highly biased • Can be hard to know when two ESTs are derived from the same gene • (generally) high error rates Ups: • Represent the part of the genome (most) people care about • Does not require a sequenced genome • Find genes • Find SNPs • Find splice isoforms
What is the future of sequencing? • Resequencing • One human done4 billion to go • Locating polymorphisms for complex diseases • More species, more individuals • Comparative genomics • What resolution (ORFs, transcription factors, individual base pairs) determines how many genomes
Really high-throughput sequencing • High-throughput=Cheap • $50 million/per mammalian genome (now) • Reduce volumes=reduce reagent cost • Eliminate/parallelize cloning and DNA preparation • Multiplex!
The $1000 Genome http://grants.nih.gov/grants/guide/rfa-files/RFA-HG-04-003.html “…we remain very far away from being able to afford to use comprehensive genomic sequence information in individual health care.”
Key ingredients so far… • Sequencing by synthesis • Elimination/parallelization of clone production • miniaturization
Pyrosequencing I http://www.pyrosequencing.com/
Pyrosequencing II • Emulsion PCR (Dressman et al (2003) PNAS 100:8817-22) to generate templates. Eliminates library construction and DNA prep • Drop one bead/well into a “picotiter” plate (1.6 million wells) • Image reactions in a flow cell with CCD camera http://www.454.com/
Fluorescent in situ sequencing (Fisseq) • Emulsion PCR to create bead library (as in pyrosequencing) • Polymerize beads in polyacrylamide on the surface of a microscope slide (2 billion beads/slide) • Perform sequencing by synthesis reactions using fluorescently labeled dNTPs dATP-Cy3
Single Molecule Sequencing Microscope slide * * * Single DNA molecule Super-cooled CCD camera/microscope primer dNTP-Cy3 * Helicos Biosciences Corp.