Examples of pathways ...
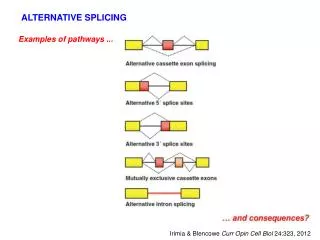
ALTERNATIVE SPLICING. Examples of pathways. Irimia & Blencowe Curr Opin Cell Biol 24:323, 2012. Types of alternative splicing vary in frequency among eukaryotes. Kim Nucl Acids Res 35: 125, 2006. Chen & Manley, 2009. Chen & Manley, 2009. How are different splicing pathways selected?.

Examples of pathways ...
E N D
Presentation Transcript
ALTERNATIVE SPLICING Examples of pathways ... Irimia & Blencowe Curr Opin Cell Biol 24:323, 2012
Types of alternative splicing vary in frequency among eukaryotes Kim Nucl Acids Res 35: 125, 2006
Chen & Manley, 2009 Chen & Manley, 2009 How are different splicing pathways selected? Splicing regulatory proteins assist or antagonize splice site recognition Fu Cell 119: 736, 2004
Positive regulation of alternative splicing - active selection of different splice site - role of enhancers (ESE or ISE) which interact with SR-type “activators” - usually < 200 nt from splice site (typically in downstream exon) often Pu-rich (or Py-rich) motifs or simple repeats Chen & Manley Nature Rev Mol Cell Biol 10:741, 2009
Negative regulation - use of normal splice site “blocked” - role of silencers (ESS or ISS) - cis-elements within exon (or intron) which interact with SR-type “repressors” eg. tissue-specific SR protein might prevent binding of U2AF to pyrimidine tract … or binding of U1 snRNP to downstream 5’splice site
Looping-out model of splicing regulation ABS & FBS = high affinity binding sites for hnRNP A/B (or hnRNP F/H) Martinez-Contreras PLos Biol 4:e21, 2006
How to discover new protein-protein interactions? Yeast two-hybrid assay Prp8p = U5 snRNP protein Prp40p = U1 snRNP protein Fused to transcription factor domain of TF In mammals: Mud2p = U2AF65 & BBP = SF1 Alberts Fig. 5-81 Weaver, Fig. 14.37 &14.38
Examples of the impact of alternative splicing on cellular functions “Alternative splicing changes protein isoforms (red half squares) by introducing new protein sequences (yellow) that are encoded by alternative exons.” Kelemen Gene 514:1, 2013
1. Sex determination in Drosophila mediated by alternative splicing - splicing pathway controls whether fly will be male or female - ratio of X chromosomes to autosomes triggers cascade of alternative splicing pathways for genes encoding: Splicing regulatory factors Sxl (sex-lethal) Tra (transformer) Transcription factors for sex-identity genes Dsx (doublesex) “Repression of tra 3’splice site involves interaction of SXL with ISS embedded in polypyrimidine tract and prevention of U2AF binding.” Matlin Nature Rev.Mol Cell Biol 6:386, 2005
sxl tra dsx Alberts Fig. 7.92
Another example: HIV retrovirus tat alternative splicing hnRNP A1 exon 3 exon 3 “Inclusion of exon 3... is determined by the nuclear ratio of specific hnRNP and SR proteins... Multimerization of hnRNP A1 from a high affinity ESS can be sterically blocked by interaction of SF2/ASF with ESE”. Show predicted results of gel mobility shift experiments. Matlin Nature Rev.Mol Cell Biol 6:386, 2005
“Action of SF2/ASF and hnRNP A1 in the selection of alternative splice sites” D = distal 5’ splice site P = proximal Interpretation of these data? Akker J Mol Endocrin 27: 123, 2002
Control of splicing pathway choice? Dose-dependent regulation - splicing efficiency dependent on relative concentration (or activity) of SR “regulators” eg. phosphorylation state - perhaps mediated by RNA pol II CTD? - cooperative or antagonistic interactions between SR-type proteins Transcriptional activators - use of different promoters for a particular gene can generate different alternatively-spliced mRNAs Models how this might be achieved? Transcription rate - if pausing of RNA pol II, opportunity for selection of different splice sites? Pawlicki & Steitz Trends Cell Biol 20:52, 2010
Effect of RNA pol II elongation rate on splicing pathway? Kinetic model Chromatin adaptor model Histone methylation status “read” by adaptor & recruits splicing factors (eg. PTB pyrimidine track binding protein) If low pol II elongation rate (or internal pausing)… chance for including alternative exon (which otherwise would be skipped over)? If high processivity… “simultaneous” presentation of both introns to splicing machinery and strong 3’ splice site outcompetes weak one? Luco et al. Cell 144:16, 2011
Integrated model for the control of alternative splicing Luco et al. Cell 144:16, 2011
How many human genes exhibit alternative splicing? Wang et al. Nature 456:470, 2008
High throughput technologies for analysis of alternative splicing Hartman & Valcarcel Curr Opin Cell Biol 21:377, 2009
EST data mining & experimental verification b) Microarray based detection a) RT-PCR based detection Modrek & Lee, Nat Genet 30:13, 2002 Wang & Cooper Nature Rev Genet. 8:749. 2007
Dscam = Drosophila axon guidance receptor - role in neuron development & synapse formation - homologue of human Down syndrome cell adhesion molecule (immunoglobulin superfamily) Red = N-terminal half of Ig2 domain Blue = N-terminal half of Ig3 domain Green = entire Ig3 domain “potentially generate more than 38,000 Dscam isoforms…. Schmucker Cell 101: 671, 2000
Therapeutic approaches utilizing splicing a) Oligonucleotide (loss of function) Antisense oligomer blocks access to splicing cis-element b) Oligonucleotide (gain of function) Bifunctional oligomers with targeting (antisense) & “effector” domains c) Trans-splicing Provide wt exon with strong 3’ ss (so that defective exon is by-passed) Wang & Cooper Nature Rev Genet. 8:749. 2007