The DNA synthesis
240 likes | 455 Views
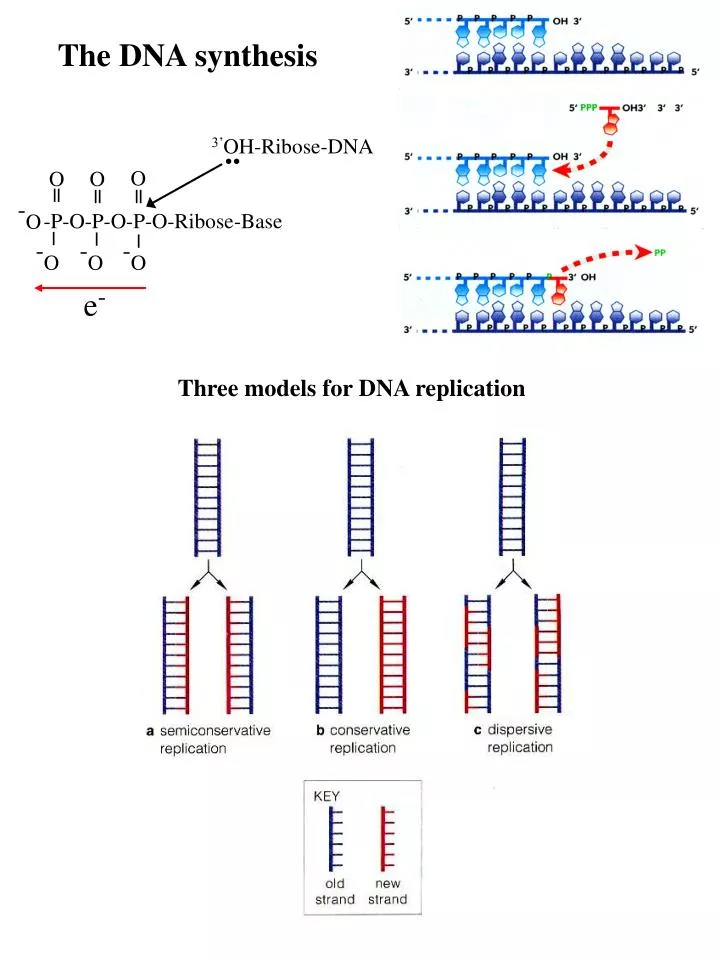
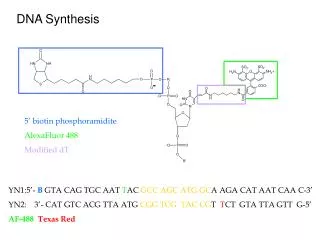
The DNA synthesis. 3’ OH- Ribose- DNA. O. O. O. ||. ||. ||. - O. -P-O-P-O-P-O- Ribose -Base. |. |. |. - O. - O. - O. e -. Three models for DNA replication. (1958). 5’-P. 3’-OH . DNA synthesis is semidiscontinuous and primed by RNA. Synthesis of Okazaki fragments (1-2 Kb).

The DNA synthesis
E N D
Presentation Transcript
The DNA synthesis 3’OH-Ribose-DNA .. O O O || || || -O -P-O-P-O-P-O-Ribose-Base | | | -O -O -O e- Three models for DNA replication
5’-P 3’-OH
DNA synthesis is semidiscontinuous and primed by RNA Synthesis of Okazaki fragments (1-2 Kb)
Synthesizing New DNA Strands An RNA primer for initiation of DNA synthesis Primer removal by RNAse H and DNA polymerase I
The holoenzyme is a complex of 900 kD (about the same mass of small ribosome unit) 1)a catalytic core(a subunit) 2)a dimerization component (t) that links two cores 3) a processivity component (b) that keeps the polymerase on the DNA 4) a clamp loader (g) that places the processivity subunits on DNA The "core enzyme" contains subunits a , e ,and . The subunit has a basic ability to synthesize DNA, the e subunit has the 3’–5’ proofreading exonuclease, and may be required for assembly. A dimer binds to the core polymerase, and provides a dimerization function that binds a second core polymerase (associated with another clamp). The holoenzyme is asymmetric, because it has only 1 complex. This complex is responsible for adding a pair of dimers to each parental strand of DNA. Each of the core complexes of the holoenzyme synthesizes one of the new strands of DNA. This corresponds to the need to synthesize a continuous leading strand (where polymerase remains associated with the template) and a discontinuous lagging strand (where polymerase repetitively dissociates and reassociates).
The structure of DNA polymerase III Catalytic core The DNA is in the classic B-form duplex up to the last 2 base pairs at the 3’ end of the primer, which are in the more open A-form. (bacterial protein which increases processivity of the catalytic subunit) The b subunit of DNA polymerase III consists of a head to tail dimer, shown in red and orange that form a ring completely surrounding a DNA duplex.
Il meccanismo d’azione della DNA polimerasi stampo A) Uno ione metallico (Mg2+ o Mn2+) riduce l’affinità del 3’-OH per il suo idrogeno favorendo la formazione di un 3’-O- necessario per l’attacco nucleofilo. B) Il secondo ione metallico coordina le cariche negative dei fosfati e e stabilizza il pirofosfato prodotto durante la formazione del legame tra innesco e nucleotide. estremità 3’OH nucleotide entrante Per la polimerazzazione dei dNTP è richiesto il corretto appaiamento delle basi e la sintesi diminuisce di un fattore 10,000 in presenza di appaiamenti errati. Pos. 3’ Pos. 3’
Come la DNA polimerasi discrimina tra desossinucleotidi trifosfato e ribonucleotidi trifosfato La DNA polimerasi non può utilizzare gli rNTP Sebbene gli rNTP siano nella cellula 10 volte più concentrati dei dNTP, essi sono incorporati ad una velocità 1,000 volte più bassa rispetto ai dNTP. Pos. 2’ Pos. 3’
L’attività 3’-5’ esonucleasica della DNA polimerasi Proofreading control (correttore di bozze ) Quando viene aggiunto un nucleotide non corretto diminuisce la velocità di polimerizzazione mentre viene stimolata l’attività esonucleasica 3’-5’. Il sistema di correzione abbassa la presenza di errori a 1 ogni 107 addizioni. DNA con appaiamenti perfetti DNA con errori DNA Pol III, triangoli Subunità θ, cerchi
The Escherichia coli DNA replication More than 500 bp/sec synthesis The loop of single-stranded template must extend for the length of at least one Okazaki fragment The model for coordinating replication by two DNA polymerases at replication fork New synthesized DNA Okazaki fragments
DNA topoisomerases provide a solution to the topological problem The substrate is ssDNA The substrate is dsDNA Type II E. coli topois. II (gyrase) and IV, eukaryotic topois. II and IV Type I E. coli topois. I and III, yeast and human topois. III, eukaryotic topois. I, archaeal reverse gyrase Reactions of topoisomerases Type I enzymes, because of their inability to make double-strands breaks, can only catalyze reactions when at least on circle bears a single-strand nick, whereas type II enzymes perform reactions with intact circles nick nick Type I Type II
Type I topoisomerases The type I topoisomerase I of E. coli (topA) relaxes highly negatively supercoiled DNA. This enzymes does not act on positively supercoiled DNA and does not need ATP to function. The formal properties of eukaryotic type I topoisomerases are similar but they can relax positive as well negative supercoils. The archaeal reverse gyrase (Solfolobolus) is able to introduce positive supercoils. Positive supercoiling could be useful to an organism growing at high temperature because it will tend to prevent DNA denaturation. A covalent bond is transiently formed between a Tyr residue of the enzyme and the 5’end of DNA target
La Topo I va incontro ad un cambiamento conformazionale La Topo I lega un tratto della doppia elica che presenta una denaturazione locale La Topo I trattiene entrambi i filamenti di DNA Sing inversion model for gyrase action Type II topoisomerases The type II topoisomerases generally relax both negative and positive supercoils. Bacterial DNA gyrase is responsible for relaxing positive supercoiling in front of replication fork. The reaction requires ATP to promote a conformational change of the enzyme. The type II topoisomerase can be used to introduce or resolve catenated duplex circles and knotted molecules. Bacterial DNA gyrase is able to introduce negative supercoils into a relaxed closed circular molecule. In the absence of ATP, the gyrase can relax negative but not positive supercoils. (-) (+) (-) (-)
Topoisomerases change the linking number of DNA in steps of either 1 or 2 Single topoisomer A group of negatively supercoiled topoisomers (A) with a given distribution of linking numbers is relaxed either by a type I or type II enzymes. The result is a group of topoisomers of increasing linking number (B). A single topoisomer form is selected (C) and subjected to relaxation by type I topoisomerases (D) and to relaxation by type II topoisomerases (E). Topoisomerases and DNA replication Topoisomerases enable the helix to be unzipped (the two strands pulled apart) without the molecule having to rotate. The replication fork is therefore able to proceed. Topoisomerasi IV DNA gyrase
The replicon Replicon is a unit of the genome in which DNA is replicated; contains an origin for initiation of replication and also have a terminus at which replication stops. The bacterial chromosome is replicated bidirectionally as a single unit from oriC. Electron micograph of -mode replication in E. coli
Demonstration that DNA replication was bidirectional in Bacillus subtilis (Gyurasits and Wake, J. Mol. Biology, 1973) 1) Bacterial spores were grown on medium containing low level of radioactive (3H) thymidine. 2) After 120 minutes of growth, high level of radioactive (3H) thymidine was supplied. 3) After 20 additional minutes DNA replication was halted. 4) DNA was exposed to X-ray film.
Methylation at the origin regulates initiation of DNA replication A bacterial (or plasmid) origin must be used to initiate replication only once per cell cycle. Initiation is associated with some change that marks the origin so that a replicated origin can be distinguished from a nonreplicated origin. The ability of a plasmid relying upon oriC to replicate in dam– E. coli depends on its state of methylation: - If the plasmid is methylated, it undergoes a single round of replication, and then the hemimethylated products accumulate in dam- cell. So a hemimethylated origin cannot be used to initiate a replication cycle. - An origin of nonmethylated DNA can function effectively both in wt and dam- strains. The SeqA mutants reduce the delay in remethylation at both oriC and dnaA, thereby accumulating an excessive number of origins. This suggests that SeqA is part of a negative regulatory circuit that prevents origins from being remethylated. SeqA binds to hemimethylated DNA more strongly than to fully methylated DNA.
DNA fully methylated. DNA replication converts GATC sites to hemimethylated state. Hemimethylated GATC sites are bound by SeqA. SeqA prevents full methylation of these sites and binding of oriC by DnaA. SeqA dissociates from GATC sites. When GATC sites become fully methylated DnaA can bind and direct a new round of replication.
Termination of DNA replication in E. coli The termination sequences function in only one orientation. This arrangement creates a "replication fork trap;" if for some reason one fork is delayed, so that the forks fail to meet at the usual central position, the more rapid fork will be trapped at the ter region to wait for the arrival of the slow fork. Termination requires the product of the tus gene, which codes for a protein (36 kDa) that recognizes the consensus sequence and prevents the replication fork from proceeding. Tus binds to the consensus sequence, where it provides a contra-helicase activity and stops DnaB from unwinding DNA. A difficulty in understanding the function of the system in vivo is that it appears to be dispensable, since mutations in the ter sites or in tus are not lethal.