Ligation Errors in DNA Computing
This study explores the reliability of the splint-aided ligation process in DNA computing, specifically focusing on the occurrence of errors due to mismatches during the ligation of DNA nodes. We examined the dependence of error rates on the number of mismatches and the combination of bases using T4 DNA ligase. Experiments revealed an 8% error rate with single mismatches, indicating that the combination of bases and competitive conditions significantly affect ligation outcomes. Our findings highlight the importance of ensuring accurate ligation for reliable DNA computing applications.

Ligation Errors in DNA Computing
E N D
Presentation Transcript
Ligation Errors in DNA Computing Y. Aoi, T. Yoshinobu, K. Tanizawa, K. Kinoshita and H. Iwasaki Biosystems 52(1999), pp.181-187 Summarized by In-Hee Lee
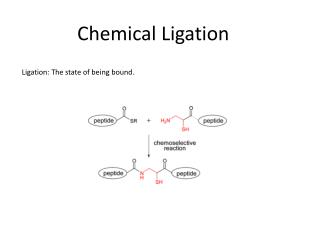
Introduction • In Adleman’s method • Generating solutions is important. • The ligation of nodes should be reliable. • Investigate the reliability of the splint-aided ligation process.
Test of Errors • Dependence on the number of mismatches • 300pmol of each oligo. • 2.8 units of T4 DNA ligase • Incubation for 20 h at 16ºC
Test of Errors 8% page, EtBr An error may occur in case of single mismatch # of mismatches
Test of Errors • Dependence on the combination of bases Fluorescein isothiocyanate (FITC)
Test of Errors • The yield of ligation 50 pmol of each oligo0.8 units of T4 DNA ligaseincubation at 16ºC for 10 m or 35 m
Test of Competing • Two sets of experiments • 1 base mismatch w/o competing • G-T or A-C • 1 base mismatch w competing • S strand and 4 R strands • Perfect match
Test of Competing • 100 pmol of each oligo • 1.4 units of T4 DNA ligase • Incubated for 30 m at 16ºC FITC EtBr staining
Test of Competing • Product of 1 is at slightly lower position • Product of 1 is not so much stained by EtBr. • Single stranded • Main product of 2 is at the same position as the product of 3 • Main product of 2 is from perfect match • In competing condition, perfect match wins the race.