Microfluidic Chromosome Sorter
Microfluidic Chromosome Sorter. Hung Li Chung ٠ Viknish Krishnan Kutty Uday Kolluri. Faculty Advisor: Dr. Helmut Strey. Project Background. Project focus: sort chromosomes accurately and cost-effectively. Why? Isolation of chromosomes has many applications: Physical gene mapping

Microfluidic Chromosome Sorter
E N D
Presentation Transcript
Microfluidic Chromosome Sorter Hung Li Chung ٠ Viknish Krishnan Kutty Uday Kolluri Faculty Advisor: Dr. Helmut Strey
Project Background • Project focus: sort chromosomes accurately and cost-effectively. • Why? Isolation of chromosomes has many applications: • Physical gene mapping • Isolation of molecular markers • Preparation of painting probes • Construction of chromosome-specific DNA libraries • Advancing research of genetic diseases • How? Sorting chromosomes using flow cytometry • Fluidic Switch Sorting • Droplet-based Sorting http://www.cshl.edu/gradschool/hatchwell_.html
Fluidic Switch Sorting Droplet-based Sorting versus • Advantages: • High throughput (5,000~10,000 cells/s) • FACS cell sorters commercially available, but only 2 FACS facilities in US sort chromosomes. • Advantages: • Inexpensive • Safe closed system design (Non-biohazardous) • Capable of handling small sample size. (100-100,000) • High yield with high purity • Self-contained Lab-on-a-Chip • Disadvantages: • Expensive ($100,000~$450,000 & technician fee) • Bio-hazardous aerosols are released by FACS sorters. • Impractical requirement of >100,000 cells to achieve high yield. • Limited purity ( 50~70%) • Disadvantages: • Speed is limited* • Not commercially available for sorting chromosomes. * Dependent on the sorting technique that is utilized.
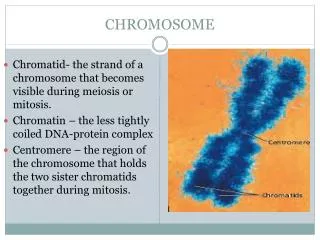
What are chromosomes? • Chromosomes are thread-like structures which are made up of long molecules of tightly-coiled DNA (hereditary information). • Chromosomes contain many genes (exons), regulatory elements (promoters) and other non-coding nucleotide sequences (introns). • In eukaryotes, uncondensed chromosomal DNA wraps around proteins (i.e. histones) to form chromatin. (Prokaryotes have a singular, circular chromosome instead of a set of chromosomes.) • In their relaxed state, chromosomal DNA molecules are used for replication, transcription and regulation. • Average chromosome width and length are 0.5 micron and 5 micron, respectively.
What are chromosomes? Metaphase chromosome Chromatid Chromatin fiber Solenoid (30 nm diamtere) Spacer DNA plus H1 histone Histone Histone DNA complex Nucleosome (11 nm diameter) DNA (2nm diameter)
Chromosome Aberration • The gain or loss of chromosomal DNA may lead to a variety of genetic disorders. Abnormalities in chromosome number or structure all called chromosome aberrations. • Deficiency, duplication, inversion, translocation, aneuploidy, and polyploidy are some forms of chromosome aberrations. http://gslc.genetics.utah.edu/units/disorders/karyotype
Karyotyping Karyotyping is the characterization of a species or organism’s chromosome complements according to their arrangement, number, size, shape, or other properties. They are useful for finding macroscopic chromosomal aberrations. http://gslc.genetics.utah.edu/units/disorders/karyotype
Means of Identification • Quantum dots • Staining • G-banding (GIEMSA) • Hoechst + Chromomycin • Spectral Karyotyping (SKY)
Image Acquisition Hoechst 33343 (BBI) 12 nm half width (HW) bandpass excitation filter at 365 nm395 nm dichroic filter418 nm longpass emission filter Chromomycin A3 (CA3) 8 nm HW badnpass excitation filter at 436 nm460 nm dichroic filter470 nm lonpass emission filter *integration time of 5 s
Image Processing Centering by calculating the center of mass in BW image Boundary Demarcation Successive Thresholding Laplacian Identification “Areal” moment of inertia
Successive Thresholding By thresholding the image successively, we can obtain the bordering for the distribution of different intensities.
Laplacian Variance of f(x,y) in some neighborhood with some radius ε is also examined High var(f(x,y)) implies existence of edge
Centering + Identification center of mass: moment of inertia: “moment of inertia” allows for easy identification of chromosome
Multivariate Sort 1. Relative intensity of BBI versus relative intensity of BBI & CA3 2. Relative intensity of BBI versus relative intensity of BBI/CA3 3. Relative intensity of BBI versus peak intensity of BBI/CA3
Sorting Consideration The peaks of fluorescence intensity is influenced by: • Quality of chromosome suspension • The narrowness of the sample stream • Intensity and frequency of laser source • Magnification (i.e. 40X vs. 63X) Detection of Abnormality The location, volume, and widthof individual distributions must be determined
Microfluidics Cell Sorting Techniques • Hydrodynamic flow control On-chip or off-chip fluidic valves capable of sorting living cells. • Optical forces Focusing radiation pressure forces from an optical beam can hold and levitate small particles (i.e. cells) in a fluidic medium without physical contact. • Electrokinetic mobilization Using a uniform, electric field to sort charged particles (or cells) in a microfluidic network. • Di-electrophoretic forces Using a non-uniform electric field to sort di-electric moieties in a microfluidic network. http://www.dafyd.me.uk/blog/docs/yoda.jpg
Chromosome extraction and isolation • Technology to lyse e-coil cells, separate chromosome from lysate, and maneuver chromosomes on a chip • Cells are lysed with osmotic shock • Chromosomes are separated from cell debris and maneuvered by dielectrophoresis
Lysing of Cells • Cells are usually sturdy and are able to withstand internal/external pressure differences • They are treated with lysozome, which softens the inner and outer membrane, and placed in highly concentrated sucrose solution • Internal osmotic pressure and the viscosity of sucrose forces the cell to deform into a sphere • De-ionized water is directed into the cell, changing sucrose concentration • Water diffuses into cells and causes it to burst due to osmotic stress • Bursting is controlled by controlling the inflow of water, thereby controlling the osmotic condition • Each cell has a different osmotic threshold for bursting
Dielectrophoresis • Movement of a polar or polarizable object in the direction of an electric gradient. • In this paper, electrodes are used to form the electric gradient. Trapping Region chromosome electrode
Protocol • After lysing, a DC electrical field 5V/cm across the chip is used to transport the cell lysate toward the Dielectrophoretic probe region. • An additional 500V/cm AC field at 500Hz is added to trap the chromosome. • As the electrical field changes, the position of the chromosome also changes.
Shows the position of the chromosome with the DC and the AC electrical field. • With the AC field turned off, the chromosome moves downwards. • With the DC field turned off, and the AC field turned on, the chromosome moves up.
Our Part We are considering the following identification and switching techniques: • Identification: multivariate sorting with hoechst and chromomycin • Di-electrokinetic movement • Use as a switch • Sorting of all 23 human chromosome pairs should be investigated • If this technique is found to be effective, it can be used to sort based on charge
The End Acknowledgments Dr. Helmut Strey, Faculty Advisor Yoda
Works Cited • Arndt-Jovin DJ Jovin TM. Multivariate chromosome analysis and complete karyotyping using dual labeling and fluorescence digital imaging microscopy. Cytometry. 1990;11(1):80-93. • Laplacian Edge Detection. http://www.owlnet.rice.edu/~elec539/Projects97/morphjrks/laplacian.html • Wang M, et al. Microfluidic sorting of mammalian cells by optical force switching. Nat Biotechnol. 2005 Jan;23(1):83-7. Epub 2004 Dec 19. • Prinz, Christelle et al. Bacterial chromosome extraction and isolation. Lab on a Chip. 2002; 2: 207-212.