Ontologies for Gene Expression
Ontologies for Gene Expression. History of ontologies in bioinformatics BioOntologies Consortium Ontologies for the biochemical networks that control gene expression. Ontologies. Clear thinking about how to structure information Clearly understand each field in a database

Ontologies for Gene Expression
E N D
Presentation Transcript
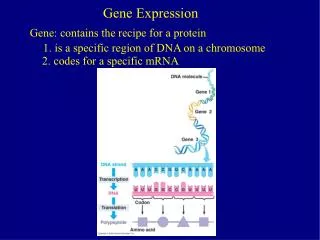
Ontologies for Gene Expression • History of ontologies in bioinformatics • BioOntologies Consortium • Ontologies for the biochemical networks that control gene expression
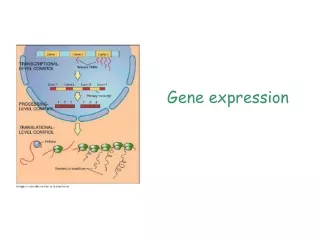
Ontologies • Clear thinking about how to structure information • Clearly understand each field in a database • Formal and informal definitions for database elements • Type of value, range of values • Product field of Gene class can be a Protein or an RNA • Ability to enforce data correctness • Ability to compute with database elements in a reliable fashion
History of Ontologies inBioinformatics • 1994 Meeting on Interoperation of Molecular Biology Databases (MIMBD-94) • BioOntologies meetings in 1997, 1998, 1999, 2000, 2001 • Ontology tutorials at ISMB conference • BioOntologies Consortium
BioOntologies Consortium • Concerned with ontology infrastructure for bioinformatics • Exchange of ontologies • Beware: All bioinformatics ontologies expressed in different ontology language • Software for constructing, interpreting, applying ontologies • http://bioontology.ingenuity.com/
BioOntologies Consortium • ISMB-2000 paper evaluating ontology exchange languages for bioinformatics
BioOntologies Consortium • ISMB-2000 paper evaluating ontology exchange languages for bioinformatics • Define criteria for evaluating existing languages • No existing languages satisfy all criteria • Desired: XML syntax, frame semantics • 1999: Karp and Chaudhri develop XOL language • 2000: OIL/DAML succeeds XOL
BioOntologies Consortium – Potential Interactions • Standards and tools • DAML/OIL • SRI’s GKB Editor ontology editor • Collaborate on ontology development • Post ontologies on BioOntologies web site
Be Precise About Ontology Uses • Data submission • Data exchange among databases • High-level database design • Mapping from ontologies to database management systems essential • Beware of flatfiles • Beware of XML
ArrayExpress • Ontology for specifying experiments • MAML import and export • SQL query access
EcoCyc Project Overview • E.coli Encyclopedia and model organism database • Tracks the evolving annotation of the E. coli genome • Over 3000 literature citations • Collaborative development via internet • Karp (SRI) -- Bioinformatics architect • Riley (MBL) -- Metabolic pathways, signal transduction • Saier (UCSD) and Paulsen (TIGR)-- Transport • Collado (UNAM)-- Regulation of gene expression • Ontology: 1000 biological classes • Database content: 16,000 instances • Over 3,300 registered users
Encoding TranscriptionalRegulation in EcoCyc -- Goals • Capture transcriptional regulatory mechanisms within a well structured ontology • Provide a training set for inference of gene networks • Interpret gene-expression datasets in the context of known regulatory mechanisms • Compute with regulatory mechanisms and pathways • Summary statistics • Pattern discovery • Complex queries • Consistency checking
Pathway Tools Extensionsfor Transcriptional Regulation • Integration of RegulonDB (Collado et al.) • Regulation ontology • Editing tools for regulatory interactions • New visualizations
EcoCyc Ontology forTranscriptional Regulation • Terminology: Transcription Unit • Definition: A set of coding regions and associated control regions that yield a single transcript • “Operons” must have more than one gene • Prokaryotic terminology • Key features of ontology • Model gross structure of transcription units, transcription factors, RNA polymerase • Model all molecular interactions as biochemical reactions • Binding of transcription factors to ligands and to DNA sites • Binding of RNA polymerase to promoter
Ontology for Transcriptional Regulation – Current Limitations • Focused on prokaryotic regulation • Mechanisms based on control of transcription initiation only, e.g., no attenuation
Ontology for RegulatoryInteractions • Common slots • Citations, Comment, Common-Name, Synonyms • Class DNA-Regions • Left-End-Position, Right-End-Position, Relative-Start-Distance • Class Transcription-Units • Components (Promoter, transcription-factor binding sites, genes, terminator) • Class Promoters • Component-Of • Promoter-Strength-Exp, Promoter-Strength-Seq • Promoter-Evidence
Ontology for RegulatoryInteractions • Class DNA-Binding-Sites • Component-Of • Regulated-Promoter, Relative-Center-Distance • Type-Of-Evidence • Classes Protein-Complexes, Polypeptides • Components / Component-Of • Class Binding-Reactions • Reactants • Activators • Inhibitors
EcoCyc Ontology forTranscriptional Regulation • One DB object defined for each biological entity and for each molecular interaction trp Int005 apoTrpR Int001 TrpR*trp site001 pro001 Int003 RpoSig70 trpL trpLEDCBA trpE trpD trpC trpB trpA
Integration of RegulonDB • RegulonDB has been loaded into EcoCyc • RegulonDB originally relational • Lisp loader tools developed for relational table dumps • Statistics: • 528 transcription units • 620 promoters • 617 DNA binding sites • 83 transcription factors
Consistency Checks onRegulonDB Data • Find transcription units containing: • Undefined components • No gene components • Genes that are not contiguous • Genes with conflicting transcription directions
Interactive Editing Tools • SRI created interactive tools for creating and modifying regulatory mechanisms • Ongoing updates to RegulonDB occur in EcoCyc
Visualization Capabilities • Transcription units • Transcription unit containing a gene: araA • Details of a transcription unit • Regulons: CRP, NARL • Pathway control • Overview: show rxns controlled by a TF (CRP, FNR), show other rxns controlled by same TF(s) (use a rxn in purine biosyn)
Characterization of the E. coliGenetic Network • 551 transcription units include 1115 (25%) genes • Controlled by 86 transcription factors • All experimentally determined
Visualization of the FullE. coli Genetic Network • Influences of transcription factors on other transcription factors • 50 of 85 TFs do not affect other TFs • Maximum network depth of 3 • Only CRP has a branching factor greater than 2 • No feedback loops other than autoregulation • Negative auto-regulation is the dominant form of feedback