Phage display
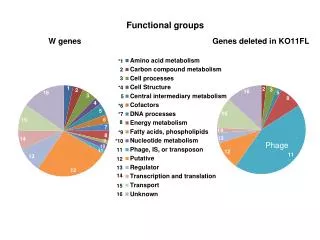
Phage display. Phage display is a term describing display of foreign (poly)peptides on the surface of phage particle. This is achieved by splicing a gene encoding such a peptide into a gene encoding a capsid structural protein. Phage Display was originally invented by George P. Smith in 1985 .

Phage display
E N D
Presentation Transcript
Phage display is a term describing display of foreign (poly)peptides on the surface of phage particle. This is achieved by splicing a gene encoding such a peptide into a gene encoding a capsid structural protein. • Phage Display was originally invented by George P. Smith in 1985
filamentous phage: M13, Ffphage Smith demonstrated that fusions to the capsid protein p3 of the non-lytic filamentous phage f1 were fairly well tolerated by cloning a fragment of the EcoRI restrictase gene in the middle section of the gene III. Progress in phage display: evolution of the technique and its applications Tomaž Bratkovič Cellular and Molecular Life Sciences, 2009 10.1007/s00018-009-0192-2
Using recombinant DNA technology, collections of billions of peptides, protein variants, gene fragment- or cDNA-encoded proteins presented on phage (so-called phage-displayed libraries) can be constructed and surveyed for specific affinity or activity. Progress in phage display: evolution of the technique and its applications Tomaž Bratkovič Cellular and Molecular Life Sciences, 2009 10.1007/s00018-009-0192-2
phagemid Phagemids combine features of plasmids (i.e., carry antibiotic resistance and enable replication of dsDNA) with features of phage vectors (i.e., allow for production and packing of ssDNA into virions).
helper phage • Phagemids are engineered to express recombinant p3 fusions under controlled conditions, but do not encode any viral structural or replication proteins. • Only upon superinfection of a bacterial host by a helper phage which contributes the missing genes can the phagemid ssDNA replication and packing take place. • Helper phage are defective in their origin of replication or packaging signal thereby ensuring preferential packing of phagemids.
helper phage transduction cDNA library phagemid construction transformation E. coli pool containing different phages screening transduction phage pool Cells with desired DNA
leucine zipper reinforced with disulphide bonds Progress in phage display: evolution of the technique and its applications Tomaž Bratkovič Cellular and Molecular Life Sciences, 2009 10.1007/s00018-009-0192-2
leucine zipper, aka leucine scissors • Each half of a leucine zipper consists of a short alpha-helix with a leucine residue at every seventh position. The standard 3.6-residues-per-turn alpha-helix structure changes slightly to become a 3.5-residues-per-turn alpha-helix.
Schematic representation of antibody fragment types displayed as fusions to p3: a single-chain variable fragment (scFv); b antigen-binding fragment (Fab) with light chain-p3 fusion (left) or heavy chain-p3 fusion (right). Progress in phage display: evolution of the technique and its applications Tomaž Bratkovič Cellular and Molecular Life Sciences, 2009 10.1007/s00018-009-0192-2
Targeted Molecular Imaging Peptide-Based Probes for Targeted Molecular Imaging Seulki Lee, Jin Xie, and Xiaoyuan Chen ,Biochemistry 2010, 49, 1364–1376
Although some success has been achieved, the use of these probes has been largely unsuccessful mainly because of their low specificity (small molecules) or limited target permeability (antibodies). • For an imaging probe to be clinically useful, it should provide a sufficient “target-to-background” ratio to maximize the “signal-to-noise” ratio or contrast in vivo. The ideal imaging compound would exhibit high binding affinity for the target, specific uptake and retention in the target, rapid clearance from nontarget tissue, adequate capillary permeability, and high stability and integrity in vivo and would be easy to prepare and safe for use. Peptide-Based Probes for Targeted Molecular Imaging Seulki Lee, Jin Xie, and Xiaoyuan Chen ,Biochemistry 2010, 49, 1364–1376
Combinatorial peptide chemistry and phage display technology, a molecular genetics approach to ligand discovery, have profoundly impacted the pool of available bioactive synthetic peptides and peptide hormones.
Combinatorial peptide chemistry Combinatorial libraries of peptide dendrimers: design, synthesis, on-bead high-throughput screening, bead decoding and characterization Noélie Maillard, Anthony Clouet, Tamis Darbre & Jean-Louis Reymond Nature Protocols 4, 132 - 142 (2009) Published online: 15 January 2009 doi:10.1038/nprot.2008.241
phage-displayed peptide libraries This method inserts the randomized oligonucleotide library DNA in-frame after a.a. 348 of capsid 10B gene. T7 lytic phage-displayed peptide libraries exhibit less sequence bias than M13 filamentous phage-displayed peptide libraries Lauren R. H. Krumpe, Andrew J. Atkinson, GaryW. Smythers, Andrea Kandel Kathryn M. Schumacher, James B. McMahon, Lee Makowski and Toshiyuki Mori Proteomics 2006, 6, 4210–4222 DOI 10.1002/pmic.200500606