Structural Templates of ATP Synthases and Uncharacterized Proteins for Computational Modeling
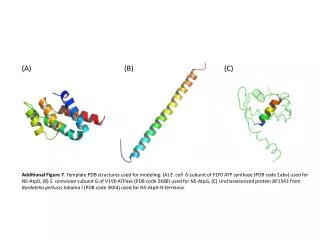
This resource details the template PDB structures employed for modeling specific ATP synthase subunits and uncharacterized proteins. It includes (A) the δ-subunit of E. coli F1F0 ATP synthase (PDB code 1ABV) utilized for NS-AtpD, (B) the subunit G of S. cerevisiae V1V0 ATPase (PDB code 2K88) used for NS-AtpG, and (C) uncharacterized protein BP1543 from Bordetella pertussis to Hama I (PDB code 3KK4), which is referenced for NS-AtpA N-terminus modeling. These structures serve as essential templates for understanding ATPase function and design.

Structural Templates of ATP Synthases and Uncharacterized Proteins for Computational Modeling
E N D
Presentation Transcript
(A) (B) (C) Additional Figure 7. Template PDB structures used for modeling. (A) E. coli δ-subunit of F1F0 ATP synthase (PDB code 1abv) used for NS-AtpD, (B) S. cerevisiaesubunit G of V1V0 ATPase (PDB code 2K88) used for NS-AtpG, (C) Uncharacterized protein BP1543 from Bordetellapertusistohama I (PDB code 3KK4) used for NS-AtpA-N terminus.