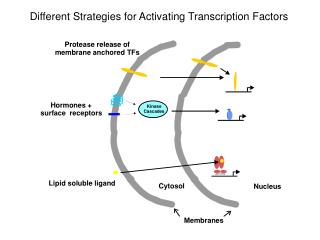
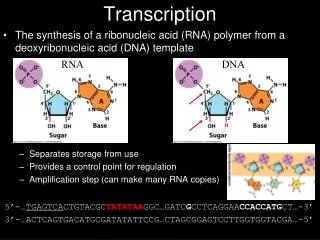
Activating Transcription
320 likes | 532 Views
Activating Transcription. Activation of gene structure (Chromatin structure) Initiation and processing of transcription Transport to cytoplasm Translation of mRNA. transcription, splicing of mRNA, mRNA stability, translation efficiency,.

Activating Transcription
E N D
Presentation Transcript
Activation of gene structure(Chromatin structure) Initiation and processing of transcription Transport to cytoplasm Translation of mRNA transcription, splicing of mRNA, mRNA stability, translation efficiency,
There Are Several Types (Classes)of Transcription Factors Basal factors (TFs) and RNA polymerase bind to promoter and TATAA box. Activators are proteins that recognize specific short DNA sequences inducing the efficiency of the promoters. Co-activators are proteins required for a more efficient transcription. They do not bind DNA. Regulators of chromatin structure How do co-activators and regulators work in transcription? Figure 25.2
Independent Domains Bind DNA and Activate Transcription • DNA-binding activity and transcription-activation are carried by independent domains of an activator. • The role of the DNA-binding domain is to bring the transcription-activation domain into the vicinity of the promoter. Figure 25.3 ---DNA-binding and Activating-binding domains are independent---(TF)
The Two Hybrid Assay Detects Protein–Protein Interactions • The two hybrid assay works by requiring an interaction between two proteins: • one has a DNA-binding domain • the other has a transcription-activation domain Figure 25.6
Activators Interact with the Basal Apparatus • The principle that governs the function of all activators is: • A DNA-binding domain determines specificity for the target promoter or enhancer. • The DNA-binding domain is responsible for localizing a transcription-activating domain in the proximity of the basal apparatus. • An activator that works directly has: • a DNA-binding domain • an Activating domain
The two-hybrid system brings the idea of the co-activators Activators may use a co-activator-two proteins- Activators-one protein two domains As long as protein complex binds both a specific DNA site and contacts the basal complex, the subunits can be encoded by separate genes.
An activator that does not have an activating domain may work by binding a co-activator that has an activating domain. Figure 25.7
How does the activator/co-activator stimulate transcription? 1- Induce some conformational change in the Transcriptional Complex 2-Increase the binding of the RNA polymerase to the promoter 3-Increase the efficiency of the Binding/Positioning of the RNA polymerase to the promoter How does this increase in the efficiency of the Binding/Positioning of the RNA polymerase to the promoter stimulate transcription? • Several factors in the basal apparatus are targets with which activators or co-activators interact. Figure 25.8
1) 2) RNA polymerase exists as a holoenzyme (i.e. the enzyme is already associated with TFs) Binding of the Holoenzyme and Mediator Figure 25.9 TFs may manipulate the structure of chromatin
Some Promoter-Binding Proteins Are Repressors Repressor • Repression is usually achieved by affecting chromatin structure. • There are repressors that act by binding to specific promoters. --affecting chromatin structure --binding proteins to DNA sequence ---Expression of Histone 2B. Testis tissues: Oct-1, CAAT and TATA are associated with TFs Embryonic tissues: another proteins (repressors), not TFs, are associated with CAAT boxes (CDP-CAAT-displacement Protein) Figure 25.10
25.7 Response Elements Are Recognized by Activators are specific DNA sequences located in both promoter /enhancer regions, which identify genes under common regulation Metallothionein (MT) gene Figure 25.11 TPA (phorbol esters) Proteins bind to these specific regulatory elements. General principle is that any one of several elements can independently activate the gene. What does this general principle mean?
Each response element is recognized by a specific activator. • A promoter may have many response elements. • Elements may in turn activate transcription independently or in certain combinations.
There Are Many Types of DNA-Binding Domains • Activators are classified according to the type of DNA-binding specificdomain. • Members of the same group have sequence variations of a specific motif that confer specificity for individual target sites.
Modes of regulation vs.DNA-binding domain Transcription factors are activated in different ways Regulation DNA-binding Most regulatory schemes involve a modification of a protein that changes its conformation or location. Activators may have the following domain/motifs: Steroid receptor Zinc finger Helix-turn-helix Helix-loop-helix Leucine zipper Sub-cellular Localization?
A Zinc Finger Motif Is a DNA-Binding Domain • A zinc finger is a loop of ∼23 amino acids. • It protrudes from a zinc-binding site formed by His and Cys amino acids. • A zinc finger protein usually has multiple zinc fingers. -(Cys, His, Phe, Leu -Zn 2+ complex) -Cys-X2-4-Cys-X3-Phe-X5-Leu-X2-His-X3-His -Organized as a single tandem repeats Figure 25.13
The C-terminal part of each finger forms an α-helix. • The helix binds one turn of the major groove of DNA. • Some zinc finger proteins bind RNA instead of, or as well as, DNA. • beta-sheets (?) • TFIIIA and elF2beta Figure 25.14
Steroid Receptors Are Activators • Steroid receptors are examples of ligand-responsive activators. • They are activated by binding a steroid (or other related molecules). • DNA-binding and ligand-binding domains. Figure 25.16 Receptor- binding TFs-binding / DNA-binding
Steroid Receptors Have Zinc Fingers • The DNA binding domain of a steroid receptor is a type of zinc finger that has Cys but not His residues. • Glucocorticoid and estrogen receptors each have two zinc fingers. • The first determines the DNA target sequence. • Steroid receptors bind to DNA as dimers. Figure 25.17 Steroid receptors(zinc finger) have several independent domains.Cys2/Cys2
Steroid receptors fall into two groups: Glucocorticoid (GR), mineralocorticoid (MR), androgen (AR) and progesterone (PR) receptors all form homodimers that recognize the consensus sequence TGTTCT. The estrogen receptor (ER) is a homodimer that recognizes TGACCT. Thyroid (T3R), vitamin D (VDR), retinoic acid (RAR) and 9-cis-retinoic acid (RXR) form heterodimers and recognize the sequence TGACCT. The other part of the heterodimer is RXR.
Binding to the Response ElementIs Activated by Ligand-Binding • Binding of ligand to the C-terminal domain increases the affinity of the DNA-binding domain for its specific target site in DNA. Figure 25.19
A steroid response element consists of two short half sites that may be • palindromic or direct repeat. • A receptor recognizes its response element by the orientation and spacing • of the half sites Figure 25.20 Figure 25.21 Steroid Receptors Recognize ResponseElements by a Combinatorial Code
The sequence of the half site is recognized by the first zinc finger. • The second zinc finger is responsible for dimerization, which determines the distance between the subunits.
without ligand With ligand
Homeodomains Bind Related Targets in DNA • The homeodomain (Helix-turn-Helix, HTH) is a DNA-binding domain of 60 amino acids that has three α-helices. Figure 25.24
The C-terminal α-helix-3 is 17 amino acids and binds in the major groove of DNA. • The N-terminal arm of the homeodomain projects into the minor groove of DNA. • Proteins containing homeodomains may be either activators or repressors of transcription. Figure 25.25
Helix-Loop-HelixProteins Interact byCombinatorial Association • Helix-loop-helix (HLH) proteins have a domain of 40 to 50 amino acids. • The motif comprises two amphipathic α-helices of 15 to 16 residues separated by a loop. • The helices are responsible for dimer formation. • The basic regions is responsible for binding to DNA Figure 25.26 HLH proteins have a protein association domain and a DNA binding domain. Dimerization and basic domains are necessary for DNA binding.
Leucine Zippers Are Involved in Dimer Formation • The leucine zipper is an amphipathic helix that dimerizes. • The zipper is adjacent to a basic region that binds DNA.
Dimerization forms the bZIP motif: • The two basic regions symmetrically bind inverted repeats in DNA. Leucine rich sequence Dimers Amphipathic alpha-helix Activator Protein 1 (AP1), a transcription factor has two major subunits, Jun and Fos. Figure 25.28
RECEPTOR Activation ? AP1