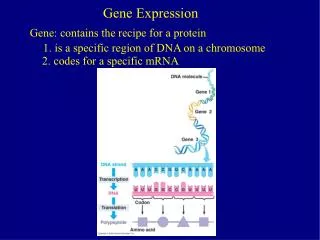
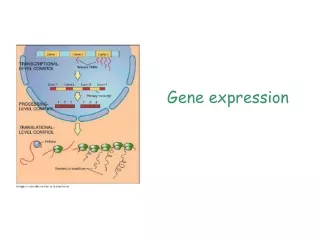
Gene Expression
Gene Expression. Christina Azodi, Lexi Bloom, Martine Stewart & Hope Yu Middlebury College : Bioinformatics and Genomics : May 3 rd 2011. POPs Microarrays . Part I: Software Comparisons. Bioconductor BRB Array NIA Array. 1. Bioconductor. Reads Affymetrix data from CEL files

Gene Expression
E N D
Presentation Transcript
Gene Expression Christina Azodi, Lexi Bloom, Martine Stewart & Hope Yu Middlebury College : Bioinformatics and Genomics : May 3rd 2011 POPs Microarrays
Part I: Software Comparisons • Bioconductor • BRB Array • NIA Array
1. Bioconductor • Reads Affymetrix data from CEL files • Upload data and normalize the intensity values • Very powerful tool- R packages exist to run 100s of different types of analysis • Our main difficulty: beginner programmers!
2. BRB Array • Uses Excel as the front end • The analytic and visualization tools are developed in R, C, Fortran and Java • Visual Basic for Applications integrates the components and hides the complexity • Normalize and summarize with GC-RMA
3. NIA Array • Initial Motivation • Trying another approach • No programs to download • Ability to analyze data stored in .txt files - The Hochstenbach set • Ability to run Principal Component Analysis
3. NIA Array: Basic Use • Obtain microarray data from NCBI GEO • Save in tab-delimited format • Upload the data matrix • Upload the probe matrix • Choose parameters • Fill in known information • Run
3. NIA Array: Basic Use • Obtain microarray data from NCBI GEO • Save in tab-delimited format • Upload the data matrix • Upload the probe matrix • Choose parameters • Fill in known information • Run
3. NIA Array: Basic Use • Obtain microarray data from NCBI GEO • Save in tab-delimited format • Upload the data matrix • Upload the probe matrix • Choose parameters • Fill in known information • Run
3. NIA Array: Challenges • Running 2-color arrays • Data available from NCBI GEO is already normalized to subtract out control levels • NIA Array requires non-normalized data (ie: hybridization values for both control and experimental) to be able to run 2-color arrays • Consequence: we were unable to retest the Hochstenbach gene set (which is 2-color) • Requires specific manipulation of input data format • Including specific knowledge of array type and experimental design—can be difficult to find
3. NIA Array: Perl to manipulate .txt • Goal: strip out the word “sample” from the text file (many times—tedious by hand) • Used • open (INPUT, $file) and open (OUTPUT, $fixedfile) to read the input file and write to the output file • $line=~ s/sample 1/ /g to replace “sample 1” with a blank space • This program could be manipulated to be useful for relabeling other data sets
3. NIA Array: Capabilities • Provides output of significantly up- and down-regulated genes • Scatter plots • Principal Component Analysis
3. NIA Array: Principal Component Analysis • Concept • Reduce the dimensionality of the data without compromising variation • Makes data easier to visualize graphically • Reduces output to a more manageable size
Part II: A Study of Two POPs • PCB & Dioxin • Disclaimers • Our Expression Results
Persistent Organic Pollutants (POPs) • Organic chemical substances • Survive in the environment for long periods of time • Travel throughout the environment • Bioaccumulate • Toxic to humans and many ecosystems • Specific effects: • Cancer • Allergies • Damage to the nervous systems • Reproductive disorders • Damage to the immune system http://www.arcticportal.org/features/2009/persistent-organic-pollutants-a-great-environmental-and-human-risk-in-the-arctic
POPs of Interest • Polychlorinated biphenyls (PCBs): • Dioxin (TCDD): https://sci10bestq3bm.wikispaces.com/PCB-silver+class+%2709 http://www.clu-in.org/contaminantfocus/default.focus/sec/Dioxins/cat/Overview/
Hochstenbach et al., 2010 • Whole-genome gene expression using microarray techniques • Human Peripheral Blood Mononuclear Cells (PBMCs) • Identified 48 genes that distinguish between immunotoxic and nonimmunotoxic chemicals.
Criteria Our criteria: • Fold change in gene expression >2 in at least two sample sets • Fold change in gene expression >8 in any of the sample sets • P<0.005 as determined by class comparison *venn diagrams made at http://www.pangloss.com/seidel/Protocols/venn.cgi Hochstenbach et al.criteria: • Fold change in gene expression >1.5 or <0.67 • Fold change in gene expression >1.5 or <0.67 in three of five donors • P<0.001 in a t-test of experimental vs. control
Tools used in BRB Array Scatterplots: • In ArrayTools, generated scatterplots with fold change limits at 2 and again at 8 • Generated a list of both up- and down-regulated genes
Tools used in BRB Array Finding significant differences in expression: • In ArrayTools, select class comparisons between groups of arrays • Generates a table of genes whose expressions are significantly different across sample groups as determined by the p-value limit (we used p<0.005) • Generates a table with log-transformed gene expressions for significant genes across all sample groups • Generates a heatmap to illustrate the differences between control and experimental conditions for all significant genes
Disclaimers • Limited to studies available through NCBI GEO • Arrays were done by different labs • Variation in concentration and exposure time • Dioxin (TCDD) • PCBs (77, 126, 153 and Arclor) • Limited in our knowledge of the system
Results: BRB Array • FN1 was differentially regulated in 4 of our studies- two PCB & 2 dioxin • ATF3 was differentially expressed in Hoch, H Hep w/ PCB and in Martine's study • Frequently differentially expressed in our studies, but not included in Hochstenbach's top 38: CYP1A1, CYP1a2, CYP1B1, Aldh3a1, Aldh1a7, Ugt1a6 • Lots of similarities between the female Rats treated with Arclor and TCDD
Results: BRB Array Results: BRB Array • Fibronectin1 (FN1) • High MWT glycoprotein • Located in plasma and at the cell surface (ECM) • Functions in cell adhesion and migration processes • 450 KDa dimer • Composed of repeating structural motifs: FN-I,II,III • FN-III10 mediates cell adhesion via an integrin-binding motif http://www.ks.uiuc.edu/Research/fibronectin/index.old.html
Immune Response: Role of integrins in NK cells cytotoxicity • NK cells are lymphocytes that destroy infected cells • Integrins are adhesion receptors that function in NK cell migration and adhesion of NK to target cells • ICAM1, ICAM2, ICAM3, FN1, and VCAM1 are all common ligands for integrins • Other genes to highlight: • ATF3&5 were upregulated • SOS1&2 were upregulated • ICAM1&3 were upregulated • IL-8 was downregulated
Hypotheses… • Transcription of FN1 may be affected by POPs and is thus downregulated so that the immune system cannot destroy infected cells • ICAM1 is most likely upregulated during the natural immune response in preparation to clear the body of cells infected with POPs • IL-8 is upregulated naturally during the inflammatory response, but found that it was downregulated in our gene set • ATF3 may be involved in the inhibition of some genes in the pathway
Conclusions • Microarray analysis software can be incredibly powerful, however can only get out how much you can put in • Limited in: • Programming knowledge • Biological system background • Possibilities for analysis are endless