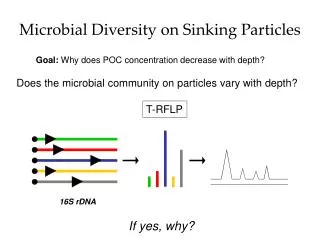
Microbial Diversity in a Dye Treating SBR
310 likes | 549 Views
Microbial Diversity in a Dye Treating SBR. by Dr. Naeem ud din Islamia College Peshawar. Biotreatment Different modes. B: isolated organisms. A: using mixed culture. C: isolated enzymes. Dyes are hard to degrade, and often result in harmful intermediates.

Microbial Diversity in a Dye Treating SBR
E N D
Presentation Transcript
Microbial Diversityin a Dye Treating SBR by Dr. Naeemuddin Islamia College Peshawar
Biotreatment Different modes B: isolatedorganisms A: using mixed culture C: isolated enzymes Dyes are hard to degrade, and often result in harmful intermediates
A Specially Designed Airlift BR from a previous Experiment for SND achieving was used The Nitrogen Removing Process was well established in that Reactor 93 % of Ammonia and COD at an HRT of 12 hrs.
MG dye- • textile industry, biological stain and antifungal. • phytotoxic, a respiratory poison, and teratogen
This SBR was subjected to gradually increasing dye concentration Optimization was achieved at a dye concentration of 25 mg/l and increased HRT of 36 hrs In this experiment we used the activated sludge as a renewable biological resource to adsorb the usual environmental concentrations of the MG dye.
In that optimized state The Color and COD removal was 80 % ammonia removal declined to 70 %. Biomass, 4 + 0.7 to 6 + 0.5 gm/l, SVI was in the range of 30 to 65 ml/gm
λmax 618 nm UV-Vis spectrophotometric scan of the biodecolorization of malachite green.
A B c OUR Correlation between ammonia, biomass, dye concentration and OUR
Knowledge about the microbial community in a dye treating reactor would be useful in association with operational conditions, to eliminate the pollutants efficiently. likely to cause the domination of certain groups of bacteria This aspect inspired our interest to know the microbial community evolved under the selective pressure of the Dye in the SBR.
Microbial community structure in the Dye Treating SBR Sludge • 16S rRNA gene Library • Phylogenetic Analysis
PCR-amplification, clone library construction and sequencing Bacterial universal primers 27F (3′-AGAGTTTGATCATGGCTCAG-5′) and 1492R (3′-TACGGYTACCTTGTTACGACTT-5′) were used for amplification.
Phylogenetic analysis The obtained sequences were edited and aligned using the BioEdit software and CLUSTAL_W program (Thompson, 199724). The sequences were compared to the known GenBank sequences using Basic Local Alignment Search Tool (BLAST). Phylogenetic trees were constructed by neighbor-joining method with the MEGA package . Identical sequences were recognized by phylogenetic tree analysis.
Phylo-genetic analysis If the sequences similarity was more than 97 %, they were considered as identical and used for further phylogenetic analysis as an operational taxonomic unit (OTU).
Culture-Independent Phylogenetic tree of the clones from the dye treating SBR,
Phylogenetic distribution profile of microbial community (Culture Independent) in the SBR.
Culture-Dependent Method • Nineteen isolates were selected from the SBR and their 16S rRNA genes were sequenced, and compared with similar sequences of the reference organisms BLAST search. Figure 6 shows the phylogenetic tree based on the culture dependent isolates identified with sequences of the NCBI BLAST. • Some of the clones identified with the well-known biodegraders, the notable being Dokdonellakoreensis, Rhodobactor, ShingomonasandParacoccus species.
Phylogenetic tree of isolates from the dye treating SBR The isolates Identified with a- & g-Proteobacteria
Phylogenetic distribution, as illustrated by isolates in the SBR involved in the biotreatment MG. All these groups well represented in the polluted environments
Table 2 shows the phylogenic affiliation and abundance of the clones. The sequences identifying with 5 divisions of Proteoabacteriai.eά-, β-, γ- §-proteobacteria and Verrucomicrobia groups were obtained. The β-, and γ-proteobacteria were in high abundance, valuing 24 % and 45 % of the total clones. The other small groups, consisting of ά-,§-proteobacteriaandVerrumicrobia groups, were 4 %, 9 % , and 2 % respectively. A moderate amount of clones, about 9 %, ranked with the uncultured bacterial strains with sequenced data in the NCBI. • The similarity of six culture independent clones(HT-69, HT-47, HT-51, HT-66, HT-38, HT-7, HT-72), to the known sequences in the GenBank was lower than 95%. Due to difficulty in translating 16S rRNA gene sequence similarity values into nomenclature, it is assumed that similarity values to the known sequences below 95% may be regarded as evidence of the discovery of novel species(3). Thus there is ample possibility of unidentified bacteria in the SBR used in the present study.
inferences • SBR with good SVI, effectively removed MG, COD and nitrogen up to 25 mg/L dye, above which a strong inhibition of these processes was observed. • The autotrophic nitrifying bacteria were not detected at high dye concentration, acting as bio-indicators for the MG toxicity. The ammonia removal pathway was, however, present, an indication of the microbialredundancy.
inferences • Majority of the sequences identified with the β- and γ-Proteobacteria. pollutant degrading bacteria, like rhodobacterales, sphingomonadaleswere in plenty. • The first time that MG treated in a nitrifying BR, with its inhibitory effects, and microbial community monitored. • Both culture-dependent and Culture independent methods must be used to have a true picuture of microbial diversity.
References • Aksu, Z.(2005). Application of biosorption for the removal of organic pollutants: a review. Process. biochem. 40, 997–1026. • Altschul, S. F.; Gish, M. W.; Myers, W.; Lipman, D.J.(1990). Basic local alignment search tool. J. mol. biol. 215, 403-410. • Amann, R.I.; Ludwig W.; SchleiferK.H. (1995).Phylogenetic identification and in situ detection of individual microbial cells without cultivation. FEMSmicrobiol. rev. 59, 143-169. • Azmi, W.; Kumar, R.; Sani & Banerjee, U.C. (1998). Biodegradation of triphenylmethane dyes. Enzyme. microb. technol. 22, 185-191. • Cabral, G. I.; Penha, S.; Matos, M.; Santos, A. R; Franco, F.; & Pinheiro, H.M. (2005). Evaluation of an integrated anaerobic /aerobic SBR system for the treatment of wool dyeing effluents. Biodegradation. 16, 81-89 • Cha, C.J.; Daniel R.D.; Carl, E.C. (2001). Biotransformation of malachite green by the fungus Cunninghamellaelegans. Appl. Environ. Microbiol. 67, 4358-4360 • Chen, K.C.; Wu, J.Y.; LiouD.J.; Hwang, S.C.J. (2003). Decolorization of the textile dyes by newly isolated bacterial strains. J. biotechnol.101, 57–68. • Daneshvar, N.; Ayazloo, M.; Khataee. A.R.; Pourhassan, M. (2007) Biological decolorization of dye solution containing Malachite Green by microalgae Cosmarium sp. Bioresource Technology. 98, 1176-1182. • Govoreanu, R.; Seghers, D.; Nopens, I.; Clercq, B. D., Saveyn, H.; Capalozza, C. (2003). Linking floc structure and settling properties to activated sludge populations dynamics in an SBR..Wat. Sci. Tech. 47, 9-18. • Hitz H.R.; Huber, W.; Rud, R.H.(1978). The adsorption of dyes by activated sludge, J. Soc. Dyers. Colour.94: 71–76. • Hong, J.; Otaki, M. (2003). Effects of photocatalyis on biological decolorization reactor and biological activity of isolated photosynthetic bacteria. J. Biosci. Bioeng. 96, 298-303. • Jiang, H.L.; Tay, J.H.; Tay, S.T.L.(2004). Changes in structure, activity and metabolism of aerobic granules as a microbial response to high phenol loading. Appl. Microbiol.Biotechnol. 63, 602–608. • Kandelbauer, A.; Guebitz G. M.(2005). Bioremediation for the Decolorization of Textile Dyes — A Review. Environmental Chemistry, Springer Heidelberg
Kipopoulou, A.M.; Zouloulis, A.; Samara, C.; Kouimtzis, Th. (2004). The fate of lindane in the conventional activated sludge treatment process, Chemosphere. 55, 81–91. Kumar, S.; Tamura, K.; and Nei, M. (2004). MEGA3: Integrated software for Molecular Evolutionary Genetics Analysis and sequence alignment. Briefings in Bioinformatics. 5, 150-163. Shi-Quan, N.; Jun, F.; Ying, J.; Yuichi, I.; Tohru, U.; Satoshi, M. (2006). Analysis of bacterial community structure in the natural Circulation system wastewater bioreactor by using a 16S rRNA gene clone liberary. Microbiol. Immunol, 50, 937-950. Ong, S-A.; Toorisaka E.; Hirata, M.; Hano, T. (2005). Treatment of azo dye Orange II in aerobic and anaerobic-SBR systems. Process Biochem. 40, 2907-2914 Rai, H.S.; Singh S.; CheemaP.P.S.; BansalT.K.; BanerjeeU.C. (2007). Decolorization of triphenylmethane dye-bath effluent in an integrated two-stage anaerobic reactor. J. Environ. Manage. 83, 290–297 Saitou, N.; Nei, M. (1987). The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406-425. Sani, R.K.; Banerjee, U. C. (1999). Decolorization of triphenylmethane dyes and textile dye-stuff effluent by Kurthia sp. . Enzyme. microb. technol. 24, 433-437. American Public Health Association.(1998). Standard Methods for the Examination of Water and Wastewater. 19th ed. Washington, DC, USA. Thompson, J.D.; Gibson, T.J.; Plewniak,F.; Jeanmougin, F.; Higgins, D.G. (1997). The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876-82. Xu, P.; Qian X. M.; Wang, Y. X.; Xu. Y.B. (2004). Modeling for waste water treatment by Rhodopseudomonaspalustris Y6 immobilized on fiber in a columnar bioreactor. ApplMicrobiolBiotechnol. 44, 676-682. Zilouei, H.; Soares, A.; Murto, M. (2006). Influence of temperature on process efficiency and microbial community response during the biological removal of chlorophenols in a packed-bed bioreactor. ApplMicrobiolBiotechnol. 72, 591-599.